Structural Biochemistry/Volume 4
Translational science is a type of scientific research that has its foundations on helping and improving people’s lives. This term is used mostly in clinical science where it refers to things that improve people’s health such as advancements in medical technology or drug development.
Examples of Application
[edit | edit source]For a long time, pathologists have noticed the fact that cholesterol was present in unhealthy arteries. In the 1960s, epidemiological studies illustrated the correlation between serum cholesterol and coronary heart disease. In the 1980s, inhibitors of HMG-CoA reductase (statins) became available to the market. These drugs were created using the biochemical knowledge of the pathways for cholesterol synthesis and transport. Subsequent clinical trials were performed to collect safety and efficacy data about the drug. After the safety and effectiveness of the drugs were confirmed, physicians and the public were educated about the drugs, and the drugs became widely used. All of this contributed to a reduction of death caused by coronary heart disease. Considering that heart disease is the biggest cause of death in the world, this example demonstrates how scientific knowledge can be used to create drugs that can improve human health or reduce human mortality. Other examples of how scientific knowledge has affected the clinical environment are “inhibitors of angiotensin-converting enzyme, inhibitors of oncogenic tyrosine kinases, and insulin derivatives with more favorable pharmacokinetic profiles”.
Glutamine [1] as a Therapeutic Target in Cancer
Another example of translational science in action is the discovery that certain cancers show a considerably high rate of glutamine metabolism. In addition, glutamine has been shown to be an integral part of metabolic functions and protein functions in cancer cells. Therefore, by designing drugs that can reduce glutamine uptake in cancer cells can potentially provide new cancer therapeutics. In fact, there are several types of drugs that have been developed that have been shown to suppress glutamine uptake. L-γ-glutamyl-p-nitroanilide (GPNA) is an example of a designed drug that serves as a SLC1A5 inhibitor that can inhibit the uptake of glutamine in glutamine-dependent cancer cells and thereby inhibit the activation of the mammalian target of rapamycin complex (mTORC1) which regulates cell growth and protein synthesis. However, more research is being done to develop a drug that specifically targets and inhibits cancer cells without damaging normal cells. In addition, translational science has led to the development of an FDA approved drug, phenylbutyrate, [Buphenyl (Ucyclyd Pharma) or Ammonaps (Swedish Orphan] that lead to a major reduction of glutamine levels in blood plasma. Also, L-asparaginase [Elspar (Merck & Co.Inc)] has also been shown to decrease glutamine levels, but it is extremely toxic in adults. The fact that cancer is a major disease that has plaqued our society in recent years, this is a good example that illustrates the importance of how translational science has led to the development of new therapeutics to help patients win the battle against cancer. It also illustrates that scientific research is constantly conducted to better drug development and a necessary aspect of science to better out lives.
Medication to Arthritis Pain
Also, through advancement of biochemical knowledge and its complementary technology, a new way to fight arthritis pain is formulated. The pain resulting from arthritis is commonly treated with aspirin, Advil, and ibuprofen. However, most of these medications lead to gastrointestinal organ damages. Up to 100,000 hospitalization and 16,500 deaths is in fact lead by side effects from those non-steroidal anti-inflammatory drugs (NSAID). Fortunately, studying NSAID at its molecular level led to identification of the solution to this major problem. The scientist discovered that NSAIDs inhibits two enzymes called “cyclooxygenases,” or COX-1 and COX-2. Even though COX-1 and COX-2 have similar functions, COX-2 is formed as a reaction to an injury and infection. This leads to inflammation and immune response, which is the reason why NSAID blockade of COX-2 relives pain and inflammation from arthritis. On the other hand, COX-1 enzyme is responsible for the production of prostaglandins, which plays role in protecting the stomach linings from the acids. As NSAID inhibits COX-1, it may lead to a major problem, ulcers. With this knowledge, biochemist scientists are able to create a type of medication that not only reduces pain and inflammation, but also removes the side effects caused by the drug. Today, this drug is called Celebrex and it is a great example of how the advancement of biochemistry and drug development technology can open up new paths to better treatments for millions of people suffering with illness.
Imprinting
Imprinting is a process that is independent from Mendelian inheritance that takes place naturally in the animal cells, which is an example of epigenetics. In more cases, two copies of genes work the same way but in some mammalian genes, either the father’s or the mother’s copy is amplified rather than both being turned on. Imprinting does not occur selectively based on gender, but occur because the genes are imprinted (chemically marked) during creation of eggs and sperm. For instance, imprinted gene of insulin like growth factor 2 (Igf2) plays a role in growth in mammalian fetus. Considering Igf2, only the father’s copy is expressed while the mother’s copy remains silent and not expressed in the life of its offspring. Interestingly, this selective silencing of the imprinted genes seems to take place in all of the mammals except platypus, echidna, and marsupials.
The questions that scientist wondered regarding this was why would evolution tolerate this kind of process that risks an organism’s survival since only one of two of the gene is expressed? The answer to this is that the mother and the father have different interest and this resulted from a competition. For example, the father’s main interest for the offspring is for it becomes big and fast because it will increase the survivability, which will in turn give greater chance of passing on the genes to the next generation. On the other hand, the mother desires strong offspring like the father, but due to limited physical resources for pregnancy duration, it is wiser to divide the resources among the offspring instead of just one. Today, more than 200 imprinted genes in the mammals are identified and some of these imprinting genes regulate embryonic growth and allocation of resources. Furthermore, mutation in these genes leads to fatal growth disorders. There are scientists now trying to understand how Igf2 and other imprinted genes stay silent in the life of cells because it can be manipulated for treatment to various mutational diseases.
Medications in Nature
There are many substances that can be used as medications and drugs already existing in the nature. For example, in 1980s, Michael Zasloff was working with frogs in the lab at the National Institutes of Health in Bethesda. There was an interesting component to the skin of the frog that allowed prevention of infection even with surgical wounds. This observation led to isolation of a peptide called Magainin that was produced by the frog as the response to the injury. Furthermore, peptide made from frog skin possessed micro-bacteria killing properties and there are hundreds of other type of molecules called alkaloid on the amphibian skin. After careful study, the scientist realized that the compound responsible for the painkilling ability is called epibatidine. Epibatidine is however too toxic to humans for pain relieving medication. Knowing the chemical structure, the goal of making similar effect drug isn’t too far.
The Sea and Cancer Treatments
The ocean is a vast resource for anti cancer treatment and its success stems from the diversity it embodies. The earth is made up of around 70% of bodies of water and there are thousands of species living in the ocean, making it the most diverse marine ecosystem. Out of the 36 known phyla, the ocean contains 34. There are many drugs that have been founded based on natural substances and the ocean is a large provider of anti cancer treatments. Marine microorganisms are extremely important and can be difficult to utilize based on where they live. Many microbes live in very specific environments and it can be difficult to find an adequate supply.
One of the maritime organisms used in cancer treatment is the ascidian Diazona. The Diazona is a source for the peptide Diazonamide A. Diazonamide A has been known to be a growth inhibitor in the cells. Compared to others like Vinblastine and Paclitaxel, Diazonamide A has the best result in inhibiting cell growth after a 24 hour exposure to Human Ovarian Carcinoma. After testing down as a joint project in the UCSD Medical center, Diazonamide A proves to be a viable anti cancer treatment. The peptide has potency with in vitro cytotoxicity because it inhibits mitotic cell division. According to in vitro date, the reason that Diazonamide A is a good inhibitor of cell growth is its attack on the cell’s tubulin assembly. Structurally, the peptide is not similar to any known drug candidate but it is still considered an “active lead compound”. However, the lack of availability of the peptide ultimately renders it unviable due to lack of practicality. Without a ready source, it cannot advance to the preclinical stage and is not a practical option for drug making and distribution. However, in 2007, a synthesis for the peptide has been available and thus finally allowing it to go further in development. An example of how peptide synthesis is an important step in drug development.

Another organism that can be used for anti-cancer treatment is the Soft-Coral Eleutherobia. Similar to the hard coral, but they are soft bodied and have no defenses against the environment. Its extract, the organic cytotoxin Eleutherobin, shows potency in cytotoxicity of 10nl/mL. Similar to the Diazonamide A, Eleutherobin is also a mitotic inhibitor. It stops the HCT116 colon carcinoma cell line by stabilizing the microtubules. When treated soluble tubulin with Eleutherobin, the microtubules are stabilized in a similar fashion to Taxol. In certain circumstances, Eleutherobin competes with the Taxol for target sites. Eleutherobin is, therefore, almost identical to Taxol and it is also a leading prospect for cancer research. However, similar to the Diazonamide A, it is not a practical lead to follow as there is no supply. There have been synthesis but none could yield practical results within reasonable costs. Because of those difficulties, it has not been advanced to preclinical stage.

A fungal strain extracted from the surface of the Halimeda. Like the two previous examples, this strain is also a potent cancer treatment. It works against HCT116 human colon carcinoma. Unlike Eleutherobin, this is not similar to Taxol but it is potent against cell lines resistant to Taxol. Halimide as a natural product, can be converted into the synthetic product NPI-2358. NPI-2358 differs from Halimide with a t-butyl group. The NPI-2358 has shown more progress than either Diazonamide A or Eleutherobin in its clinical stages. It has shown great in vivo results and since 2006, has been in phase 1 of clinical trials. In in vivo experiments with breast adenocarcinoma, the NPI-2358 has shown significant necrosis in tumor. The experiment includes a the breast adenocarninoma to be grafted onto the dorsal skin of a mouse with a viewing window. After 15 days of treatment, the group treated with the NPI-2358 shows a reduction in the tumor size in comparison to the control group. Upon closer inspection, the NPI-2358 seems to target tumor vasculature.
An organism that has been founded through the sampling methods is the Salinispora. They are special in that they require salt for growth and in cultivating the bacteria, grains of salt need to be added directly. They have unique colorings and 16s rDNA sequence. More than 2500 rDNA strains from the Salinispora have been studied. Their geographical location is along the earth’s equator, they have been discovered in Hawaii, Guam, Sea of Cortez, Bahamas, Virgin Islands, Red Sea, and Palau. The extract from the bacteria has shown cytotoxicity against HCT-116 colon carcinoma. To prepare, the Salinispora is cultured in shake flask for 15 days with XAD absorbent resin, the resin is then filtered with methanol. The cytotoxic effect of the Salinispora is very effective, it has a broad but very selective of cancer line cells. Above 2 microM, the bacteria is effective against several cancer types. The bacteria proves to be a wonderful inhibitor of cancer mechanism. Salinosporamide A is very effecting in inhibiting proteasome in vivo. It shows an average IC50 of less ca. 5nM against the 60 call line panel. It is also active against several other types of cancers. Currently as of 2006, it is in phase 1 of human trial and shows great promise as a future drug against some cancers.
Similar to the Salinispora, another marine stain of the Marinospora also requires salt. They are discovered along the Sea of Cortez by deep ocean sampling. Like the Salinispora they also have unique 16S rDNA sequences. They are also morphologically identical to the Streptomyces. Their extracts show good potency against drug-resistant pathogenic bacteria and cytotoxicity against certain cancer cell lines. The fractionation of the strain also leads to a new macrolide class. Marinomycin A from the strain is shown to be very potent against melanoma. It has been advanced to the hollow fiber assay.
The ocean is a diverse place and the many organism that it houses has great medicinal effects. Many of the populations were previously unknown but with strategies of deep sea sampling, in two years, 15 new genera of bacteria from 6 families have been cultured. More amazingly, of the 15 genera cultured, 11 have shown two be good inhibitors of cell growth and can be possible anticancer ingredients. That simply shows the wealth that the ocean has to offer in battling against dangerous illness.
Stem cell
There are cell in the human body that is completely generic and it has the ability to express extremely broad array of genes. Simply put, stem cells are able to becoming all kinds of cells in the body with an unlimited potential. Specifically speaking, this cell exists for few days after conception and is called embryonic stem cell. Once these embryonic stem cells differentiate, there are cells called adult stem that too have similar ability as the embryonic stem cell. These cells are located throughout the body, mostly in bone marrow, brain, muscle, skin, and liver. Then tissues are damaged by injury, disease, or age, it can be replaced by these stem cells. Adult stem cells however are dormant and remain undifferentiated until the body signals for its need. Adult stem cells have the capacity of self-renewal but different from the embryonic stem cell in a sense that it exists in small numbers and aren’t too flexible in differentiating. Adult stem cells plays role in therapies that treats lymphoma and leukemia. Scientists are able to isolate an individual’s stem cells from blood and grow them in the laboratory. After high dosage chemotherapy, the scientist can use the harvested stem cells to transplant and inject into replacing the cells destroyed by the chemotherapy. James A. Thomson of the University of Wisconsin was the first one to isolate stem cells from embryo into growing them in the lab. Stem cell research opens possibilities to treating Parkinson’s disease, heart disease, etc. diseases that involve irreplaceable cells.
AIDS Treatment
To understand strategies to combat HIV-1 infections, a study of its biology must be conducted. HIV-1 was found to interact with the host cells by means of their glycoproteins, gp120 and gp41. The receptors CD4, CCR5, and CXCR4 recognize these envelope proteins and together, they lead to the fusion of the virus and the host cell. In the beginning, how HIV-1 was treated was by preventing the protein from maturing and stop the RNA to replicate into the DNA. However, since both of these happen after the host genome as already been infected- a much more attractive strategy is to stop the virus from fusing with the cell. This line of research leads to the discovery of many HIV entry inhibitors.

Since the glycoproteins gp120 and gp 41 play an important role in viral infection, their structures were studied. The gp120 interacts with the CD4 by undergoing a conformational transformation. This transformation exposes the gp120 to the receptor proteins CCR5 o CXCR4. gp41 also has a major conformational transformation that changes from a prefusion complex with the gp120 into a structure that is able to place the viral and host membranes side by side. Entry inhibitions that can prevent that step from happening can prevent the cell from being infected. The gp41 has a six-helix bundle (6-HB) made from N-and C- HR regions. The N-HR forms a core and the C-HR packs tigtly against it. The formation of stable crystals from these peptides aids in the search of finding a peptide inhibitor for the six-helix bundle. An HIV-1 entry inhibitor is the T-20, which is a homolog to the helical C-HR region of the gp41.
Experiments in the absence of high resolution structure of gp41 shows that the T-20 interacts with the N-HR helical region and acts as an entry inhibitor. When high resolutions of the gp41 structures were available, the T-20 was shown to form a heterocomplex that inhibits the formation of the 6-HB hairpin is required of the viral and host genome fusion. This result shows that either the C-HR or the N-HR could act as an entry inhibitor. The N-HR peptides are trihelical and they have deep hydrophobic pockets, these pockets have a complementary pocket-binding domain (PBD) present on the C-HR peptides. This stable helical structure shows that a point for the success of the T-20 is its ability to have a helical structure as well. In general, to improve inhibitions, w can increase the helicity of the C-peptides, increase the T-20 interactions with the N-peptide trihelices, or to make N-peptides that form soluble, stable triple helix core. A good relationship between the tendency to form a stable helix with inhibition activity was seen from many entry inhibitors.
Unlike the T-20, the C-34 has a PBD sequence that can interact with the hydrophobic pockets of the N-HR core. The C-34 is therefore effective against strains of HIV that are resistant against T-20. The C-34 works by preventing the formation of the 6-HB. On the other hand, T-20 works by interacting with the lipids that interrupts the membrane fusion pore from forming. If both characteristics of both the C-34 and the T-20 were incorporated into a single inhibitor, the result would be very potent.
Recent studies have shown that fatty acid and cholesterol may be used to act as peptide fusion inhibitors for gp41. During T-20 binding to liposomes the LBD domain plays a role in the fusion process. Fatty acids were shown to have similar binding characteristics which make them possible candidates to act as fusion inhibitors along the binding locus. Researchers speculate that the reason for this is because (C-16)-DP combats HIV-1 by increasing inhibition around the viral membrane. Similarly cholesterol helps by targeting C34 to lipid rafts which also increase inhibition activity around the membrane. These kinds of applications are extremely useful when combined with drug treatments that have limited local concentration.
N-peptides are another viable solution for inhibition because they show similar properties to that of N-HR on gp41. They contain the 5-helix design which inhibits HIV-1 fusion at nM concentrations. Studies show that stability of the triple helical core may be correlated to the effectiveness of HIV treatment. It has been shown that disulfide stabilizes the trimeric coiled-coil core which will increase inhibition properties. When combined with N-HR and C-HR inhibitors this virus has an increasingly difficult time surviving without mutating further. The downside to N-peptide treatment is the higher molecular weight which may lead to immunogenicity if not used through injections.
A final possible fusion inhibitor was found using D-amino acids. These D-amino acids mimic binding to the trimeric core. Additionally these amino acids are highly resistant to protease degradation which makes them more effective than T-20 which cannot be absorbed by paracellular passages in the intestine. This leads to the possibility of using D-amino acids in topical treatments which can be easily applied at a less expensive cost.
Protein and Drugs
Each individual human body has variations in the genetic makeup that leads to difference in general health. There are environmental and lifestyle factors involved, but the response that every individual have on medications is due to a variant gene called cytochrome P450 protein. This protein is in charge of processing any kind of drugs the body intakes. Due to uniqueness of the individual’s genes, the encodings for the cytochrome P450 is very exotic. This information was discovered in the 50s as certain patients had different side effects to anesthetic drug, which was fatal. Through experiments, the scientist realized that genetic variation can cause a dangerous side effect because cytochrome P450 protein wasn’t able to break down the medication in the normal way. Even medicines like Tylenol can sometimes give no relief to the body because of the genetic variation. Fortunately, with greater knowledge of this, pharmacogenetic scientists can now develop drugs that are customized based on individual’s genes.
Obstacles and Potential Solutions
[edit | edit source]To mediate between laboratory science and clinical science is not an easy task. It requires a vast amount of different types of complex and specialized knowledge, and this brings up a lot of problems and obstacles. One research team is not even nearly enough to create a bridge between basic and clinical science. One proposal is to create research teams that specialize in different steps to interconnect basic science into the clinical environment. It also seems very practical to train individuals that can mediate between these different steps so that if a research team does not exist at one step, these mediators or translators can try to find assistance from teams at adjacent steps of the process. These proposed solutions on how to implement translational science depend on the cooperation of various types of scientists. The idea is to allow translational scientists to have easy access to the wide array of intricate and specialized knowledge needed to bridge the gap between scientific research and clinical and medical advancements.
Over the years, an increasing emphasis have been placed Translational Science. National fundings and policies have greatly facilitated the growth of the field creating opportunities for the advancement of applied clinical research informatics (CRI) and translational bioinformatics (TBI). Examples includes The National Cancer Institute's caBIG program which engineered a variety of service oriented data-sharing, data-managing, and knowledge management systems, and the CTSA which aim focuses on informatics training, database design/hosting, and execution of complex data analysis. The issue however, is that such programs, and the fundings that accompany it are geared towards solving immediate problems while neglecting to focus on foundational CRI and TBI research that is crucial to the growth of biomedical informatics subdisciplines that ensures future innovations. Furthermore, the funds are usually allocated to service oriented research that provides the resolutions to immediate problems while the policies restrict the resources of data networks to specific universities or centers. Several resolutions have been proposed in response to the issue at hand:
- Rigorous campaign advocacy to ensure that foundational CRI and TBI knowledge and practice is both recognized and supported as a core objective of translational change which will engage informaticians as equal partners in planning an execution as opposed to mere service providers.
- Community effort to refine and promote national scale agenda that focuses on challenges and opportunities facing CRI and TBI allowing them to do more than just react to new fundings and policies.
- Creation of a forum to ensure that establishment of policies and fundings affecting CRI and TBI are open to researched of the field and not limited to the few institutions, investigators, or lenders.
National Institutes of Health (NIH) and Clinical and Translational Science Awards (CTSA)
[edit | edit source]The National Center for Research Resources (NCRR) of the National Institutes of Health (NIH) has created the CTSA for the purpose of encouraging translational science and research. The NCRR has used the CTSA to give funding to infrastructure for translational science in areas such as “biostatistics, translational technologies, study design, community engagement, biomedical informatics, education, ethics and regulatory knowledge, and clinical research units”. The CTSA has six major goals with regards to translational science. The first goal is to train individuals in the field across the entire translational spectrum. This involves giving training to MD to allow them to act as clinical investigators, but it also involves teaching PhDs the fundamentals about the medical world. The expected result from this training would be that PhDs would know when they have come across something that would be of medical importance and so that MDs could understand what it is that the PhDs are trying to say to them. Secondly is the goal of trying to simply the translational process. This means trying to speed up the translational process as much as possible without losing regards to safety. This would include making expertise available to clinical researchers by the way of institutional review boards, FDA regulations and applications for investigating new drugs. The third goal of the CTSA is to take advantage of advances in informatics, imaging, and data analysis by applying these advances directly to research that clinical investigators are doing. By taking advantage of these resources, it is more likely that the investigators will come up with a meaningful study. The fourth goal is to find a way to encourage and protect the careers of translational researchers. An example of a path that propagates this career is MD/PhD programs. These types of programs could bridge the so-called translational divide by education people in aspects of both the medical and scientific fields. The success of these programs depends largely on tuition assistance provided, making it so that the graduates of these programs are not burdened with large loans to pay off. The fifth goal is to provide team mentoring, as well as support to junior clinical scientists. This goal can be achieved in part through programs like the K12 awards. The final goal of the CTSA is to catalogue all research resources in order to make these resources available to everyone possible that could need them.
References
[edit | edit source]- Payne, Phillip, Embi, Peter, Niland, Joyce "Foundational biomedical informatics research in the clinical and translational science era: a call to action." JAMIA, August 2010 vol. 17 no. 6 615-616. Web. <http://jamia.bmj.com/content/17/6/615.full.html#ref-list-1>.
- McClain, Donald. "Bridging the gap between basic and clinical investigation." Trends Biochem Sci. 2010 Apr;35(4):187-8. Epub 2010 Feb 19.
- "The Structures of Life." National Institutes of General Medical Sciences. U.S. Department of Health and Human Services, July 2007. Web. <http://www.nigms.nih.gov>.
- Z. Cruz-Monserrate, H. Vervoort, R. Bai, D. Newman, S. Howell, G. Los, J. Mullaney, M. Williams, G. Pettit, W. Fenical and E. Hamel "Diazonamide A and a Synthetic Structural Analog: Disruptive Effects on Mitosis and Cellular Microtubules and Analysis of Their Interactions with Tubulin." Molecular Pharmacology, June 2003 vol. 63 no. 6 1273-1280. Web. <http://molpharm.aspetjournals.org/content/63/6/1273.full#cited-by>.
- B. Long, J. Carboni, A. Wasserman, L. Cornell, A. Casazza, P. Jensen, T. Lindel, W. Fenical, C. Fairchild "Eleutherobin, a Novel Cytotoxic Agent That Induces Tubulin Polymerization, Is similar to Paclitaxel (Taxol)" Cancer Research, March 15, 1998. Web. <http://cancerres.aacrjournals.org/content/58/6/1111.full.pdf>.
- W. Fenical "The Growing Role of the Ocean in the Treatment of Cancer" Lecture. April 2010
- "The New Genetics." National Institutes of General Medical Sciences. U.S. Department of Health and Human Services, October 2006. Web. <http://www.nigms.nih.gov>.
- "The Chemistry of Health." National Institutes of General Medical Sciences. U.S. Department of Health and Human Services, August 2006. Web. <http://www.nigms.nih.gov>.
- F. Naider, J. Anglister"Peptides in the Treatment of AIDS" Curr Opin Struct Biol. <http://www.ncbi.nlm.nih.gov/pmc/articles/PMC2763535/>
- "Inside the Cell." National Institutes of General Medical Sciences. U.S. Department of Health and Human Services, July 2007. Web. <http://www.nigms.nih.gov>.
INTRODUCTION
[edit | edit source]Mellitus diabetes is categorized as a group of chronic (lifelong) diseases in which high levels of sugar (glucose) exist in human blood. It is the 9th leading cause of death in the world, killing more than 1.2 million people each year. Diabetes is categorized in many groups: type 1, type 2, gestational, prediabetes and a few more. However, the two main groups are type 1 and type 2, type 2 being more common. Different symptoms correlate to different types of diabetes, but generally, diabetes exhibit similar symptoms such as the following:
Type 1 Symptoms include:
- Frequent urination
- Unusual thirst
- Extreme hunger
- Unusual weight loss
- Extreme fatigue and irritability
Type 2 Symptoms include:
- Any of the type 1 symptoms
- Frequent infections
- Blurred vision
- Cuts/bruises that are slow to heal
- Tingling/numbness in the hands/feet
- Recurring skin, gum, or bladder infections
Many complications arise and are caused by diabetes. People may experience problems in their vision which potentially leads to blindness, numbness in their feet and hands, especially in the legs, hypertension (high blood pressure), their mental health, hearing loss, and others that are gender related. Although there is no cure for type 1 diabetes, maintaining an idea body weight with an active lifestyle and a healthy diet can prevent and sustain type 2 diabetes.
TYPE 1 DIABETES
[edit | edit source]In type 1 diabetes, the pancreas fails to produce little or no insulin. Insulin is a hormone that is produced by beta cells located in the pancreas. It functions as a transporter of sugar (glucose) into cells throughout the body. Glucose travels through the hemoglobin, an enzyme found in blood, and is stored away and later used for energy by the body’s organs. The failure to produce insulin results with a lack of an appropriate amount needed for the human body to function at a normal pace. Without enough insulin, sugar accumulates in the bloodstream, thus raising the blood’s sugar level – this event is called hypertension.
There is no exact cure for type 1 diabetes and the exact cause is still unknown. Researchers believe it is an autoimmune disorder, a condition in which the immune system mistakenly damages healthy tissues. In this case, the pancreas would have been attacked, preventing its function to produce insulin. Type 1 diabetes shows hereditary correlation, meaning this disease can be passed down through families. Also, the adolescent group is the most often diagnosed group of people.
TYPE 2 DIABETES
[edit | edit source]In type 2 diabetes, fat, liver, and muscle cells become insulin resistant. Those cells do not respond to insulin correctly and fails to obtain the sugar (glucose) that is being transported. Because glucose is crucial for the cells’ functions, the pancreas would sustain the equilibrium. That means, if the cells do not intake the glucose, the pancreas would automatically create more insulin to make sure the cells have enough. The remaining sugar in the bloodstream would accumulate, resulting with hyperglycemia.
Maintaining a healthy diet is essential in preventing type 2 diabetes. Low activity level, poor diet, and excess body weight increases the risk of type 2 diabetes because increased fat levels slows down the ability to properly use insulin.
Treatment
[edit | edit source]Although there is no known cure for diabetes, it can be managed with certain precautions.
Type 1 & 2 Diabetes Management
[edit | edit source]Those with type 1 & 2 diabetes, who want to maintain a healthy lifestyle, exercising regularly and maintaining a healthy weight, eating healthy foods, and most importantly monitoring their blood sugar levels.
Diabetics should try to maintain their blood sugar levels between these readings: Daytime glucose levels: between 80 and 120 mg/dL (4.4 to 6.7 mmol/L) Nighttime glucose levels: between 100 and 140 mg/dL (5.6 to 7.8 mmol/L)
People with type 1 diabetes need insulin to survive, therefore, they typically inject themselves with insulin using either a fine needle and syringe, an insulin pen, or an insulin pump.
TESTING
[edit | edit source]To determine if one is diagnosed with diabetes, a few tests must be done.
- Fasting Blood Glucose (Type 1, 2) - Blood test must be higher than 126 mg/dL twice
- Random (nonfasting) Blood Glucose (Type 1) - Blood test is higher than 200 mg/dL - Must be confirmed with fasting test
- Oral Glucose Tolerance Test (Type 1, 2) - Blood level is higher than 200 mg/dL after 2 hours
- Hemoglobin A1c (Type 1, 2) - Normal: <5.7% - Pre-diabetes: between 5.7%-6.4% - Diabetes: >6.5%
- Ketone (Type 1) - Done with urine or blood sample (If sugar level is > 240 mg/dL, when ill (ex. pneumonia, stroke, etc.), when nauseated, vomiting, when pregnant)
STATISTICS
[edit | edit source]Out of 25.8 million people in the United States, 8.3% of all children and adults are diagnosed with diabetes. It is the leading cause of nontraumatic lower-limb amputations, kidney failure, and blindness in the United States and contributes greatly to heart disease and stroke.
CLINICAL RESEARCH
[edit | edit source]Currently, thousands of laboratories are focusing on the study of diabetes: type 1, type 2, its relationship with heart, kidney diseases, obesity, and many more.
One specific research was done by Karolina I. Woroniecka and company in the Albert Einstein College of Medicine of Yeshiva University. Their topic mainly focuses on the relationship between diabetes and kidney failure, known as diabetic kidney disease (DKD), which is the prominent cause of kidney failures in the United States. The research topic is called “Transcriptome Analysis of Human Diabetic Kidney,” and was published in September 2011. Its objective was to provide a collection of gene-expression changes in human diabetic kidney biopsy samples after being treated. Gene-expression is defined as the translation of information from a gene into a messenger RNA and then to a protein. Transcriptome analysis is often used to obtain insight into disease pathogenesis, molecular classification, and the identification of biomarkers, indicates the presence of some sort of phenomenon, used for future studies and treatments. This study was able to catalog gene expression regulation, identify genes and pathways that may either play a role in DKD or serve as biomarkers.
44 dissected human kidney samples were used in this experiment, portioned out according to their racial status and glomerular filtration rate (25-35 mL/min). A glomeruli is a cluster of capillaries around the end of a kidney tubule. Their method included a series of statistical equations to identify expressed transcripts found in both the control and the diseased samples. Also, algorithms helped the study by defining the regulated pathways.
The human kidneys were obtained from donors and leftover portions of kidney biopsies. The samples were manually microdissected and only the samples without any degradation were further used through the amplification of the RNA. Before any treatment, the raw samples were normalized using the RMA16 algorithm. Its purpose is to obtain a stabilized set of data and reduce any inconsistencies in their patterns. This is where the Benjamin-Hochberg testing was used at a p value < 0.05. After, the oPOSSUM software determines the overrepresented transcription factor binding sites (TFBSs) within a catalog of coexpressed genes and is then compared to a control set. The differentially expressed transcripts that comply with the statistical conditions undergo analysis that uses a ratio to determine the top canonical pathways – the Fischer exact test is used at p value < 0.05. Immunostaining is a major component in the visualization and final step of the procedure. This procedure requires the use of a specific antibody to detect a specific protein in a sample. The following primary antibodies were used: C3, CLIC5, and podocin. The Vectastain ABC Elite kit was used for the secondary antibodies to bind to the proteins and then, 3,3”diaminobenzidine was applied for visualizations. Immunostaining is typically scored on a scale of 0-4, correlating to the amount of activity on that specific protein.
Results from this experiment identified 1,700 differently expressed probesets in DKD glomeruli and 1,831 probesets in diabetic tubuli (seminiferous tubules); probeset is a collection of more than two probes and is designed to measure a single molecular species. There were 330 probesets that were commonly expressed in both compartments. Pathway analysis emphasized the regulation of many genes that factored into the signaling in DKD glomeruli. Some molecules included Cdc42, integrin, integrin-linked kinase, and others. Strong enhancements for the inflammation-related pathways were shown in the tubulointerstitial compartment. Lastly, the canonical signaling pathway was regulated in both the DKD glomeruli and tubuli, which are associated with increased glomerulosclerosis.
With ongoing research about diabetes-linked diseases, results that are obtained contribute to the overall understanding of the biochemical processes and issues. Dr. Karolina I. Woroniecka and company are one of the many research teams throughout the world that dedicate their jobs to saving or improving people’s well-being. This study is one of the many that contribute to the complications of diabetic-related kidney diseases. However there are many more studies that relate to diabetes, such as obesity and heart attack/failure.
Hiroaki Masazuki and company conducted a project relating obesity and diabetes, “Glucose Metabolism and Insulin Sensitivity in Transgenic Mice Overexpressing Leptin with Lethal Yellow Agouti Mutation.” This article was published in August 1999 from the Department of medicine and Clinical Sciences, Kyoto University Graduate School of Medicine at Kyoto, Japan. The objective of this research project was to determine the usefulness of leptin for the treatment of obesity-related diabetes. Leptin is an adipocyte-derived blood-borne satiety factor that increases glucose metabolism by decreasing food intake and increasing energy expenditure. Two different types of mice were crossed and examined at weeks 6 and 12 during the experiment. The first type was a transgenic skinny mice overexpressing leptin breed, with allele Tg/+, and the second is a lethal yellow KKAy mice, commonly used as models for obesity-diabetes syndrome, with allele Ay/+. The F1 animals’ metabolic phenotypes were examined, noting everything from body weight to their sensitivity of insulin and concentrations of leptin. This study was able to demonstrate the potential usefulness of leptin along with a long-term caloric restriction for the treatment of obesity-related diabetes. It demonstrated that hyperleptinemia can delay the onset of impaired glucose metabolism and hasten the recovery from diabetes during caloric food restriction in the crossed F1 bred mice, Tg/+ and Ay/+. Hyperleptinemia is defined as increased serum leptin level.
Although leptin may have been found to be potentially useful in treating diabetes, the fact that a caloric food restriction is required suggests that leptin can stimulate glucose metabolism independent of body weight. Other studies have demonstrated that leptin stimulates glucose metabolism in normal-weight nondiabetic mice and also improves impaired glucose metabolism in over-weight diabetic mice with leptin deficiency. Masazuki and company have created transgenic mice models overexpressing leptin (allele Tg/+) that exhibit insulin sensitivity and increased glucose tolerance. A liver-specific promoter controls the overexpression of leptin and insulin sensitivity results with the activation of signaling in the skeletal muscle and liver. In this study, Masazuki and company genetically crossed the transgenic mice and lethal yellow obese mice. The resulting 4 genotypes are: Tg/+: Ay/+, Tg/+, Ay/+, and wild-type +/+. When at week 6, all the mice were at normal body weight and at week 12, the mice with the Ay/+ allele clearly developed obesity. At 9 weeks, +/+, Ay/+, and Tg/+: Ay/+ were placed on a 3 week food restriction diet and analyzed at week 12.
The research design and methods include: measurements of body weight and cumulative food intake, plasma leptin, glucose, and insulin concentrations, glucose and insulin tolerance tests, and caloric food restriction experiments and later statistical analysis were done. Body weights were measured daily since the mice were 4 weeks old and food intake was measured daily over a 2-week period. Blood was sampled from retro-orbital sinus of mice at 9:00AM. Plasma leptin concentrations were determined using radioimmunoassay (RIA) for mouse leptin. Insulin and plasma glucose concentrations were determined by the glucose oxidase method with a reflectance glucometer. The glucose tolerance tests (GTT) were done after an 8 hour fast and injections of 1.0 mg/g glucose. The insulin tolerance tests (ITT) were done after a 2 hour fast and injection of 0.5 mU/g insulin. The blood was then drawn from the mice tail veins at periodic times after injection at 15, 30, 60, and 90 minutes and blood was drawn from before the injections to measure comparable results. The food restriction experiment was based off of the cumulative food intake at week 12. The mice were then provided with 60% of the amount of food consumed. The exact same tests were measured: plasma leptin, glucose, and insulin concentrations were also determined; GTT and ITT were also done. At the end, all these data were analyzed and expressed at ±SE.
The results identified a large difference in body weights with the four genotypes. At week 4, all the mice showed no significant difference in body weight. At week 6 of age, Tg/+ mice gained approximately 20-30% less weight than the control +/+ mice and indicated a sign of developing adiposity compared to +/+, Tg/+: Ay/+, and Ay/+ mice. At this time, the control, Tg/+: Ay/+, and Ay/+ mice showed no drastic difference in body weights. However, by week 12 of age, the mice with the Ay/+ allele developed obese. As for plasma leptin concentrations, 6 week old Tg/+ mice were approximately 12 times those of the control +/+ mice, at week 12, they were 9 times higher. The concentrations in Ay/+ and +/+ mice were roughly equivalent. The concentrations of Tg/+:Ay/+ mice were 8 times higher than those of the +/+ mice and at week 12, they were higher than the Ay/+. At week 12, the body weight of Tg/+ was ~23% less than the control’s. The food intake of Tg/+ reduced significantly after 6 and 12 weeks of age compared to the control litter. The food intake of Ay/+ mice increased by 50% compared to the control litter. The food intake of Tg/+: Ay/+ mice, compared to the +/+ mice, were roughly the same. The food intake of Ay/+ and Tg/+: Ay/+ were approximately the same. At week 6, the plasma glucose concentrations among all 4 genotypes were the approximately the same. At week 12, the glucose levels of Tg/+ and +/+ mice were the same. However, the glucose level of Ay/+ and Tg/+: Ay/+ elevated significantly compared to the control but compared to each other, they were the same. As for plasma insulin concentration levels, the Tg/+ mice greatly decreased compared to the control at week 6. The plasma insulin concentrations in Tg/+: Ay/+ mice were higher than the control. At this point, the Ay/+ mice demonstrated marked hyperinsulinemia compared to the rest of the genotypes. GTTs and ITTs showed that the plasma glucose elevation is significant in Tg/+ compared to the control. 30 minutes after the injection, the glucose concentrations increased greatly in Ay/+ mice compared to the control.
The genotypes’ glucose metabolisms were examined after their food restriction. 60% of their total food intake were given to these mice and after 2 weeks, the body weights of Tg/+ were 17% less and +/+ were 12% less compared to before and the Ay/+, Tg/+: Ay/+ body weights also decreased. The plasma leptin concentrations in Tg/+ mice were higher than those in +/+ mice and Tg/+:Ay/+ were higher than those of Ay/+. The leptin concentrations between Tg/+ compared to Tg/+:Ay/+ and those of +/+ and Ay/+ were approximately the same. After 3 weeks of food restriction, the plasma glucose concentrations among +/+, Ay/+, and Tg/+:Ay/+ were similar. The plasma insulin concentrations, however, in Ay/+ mice were higher than those of +/+ and Tg/+:Ay/+ mice.
The results indicated that glucose tolerance and insulin sensitivity are increased in Tg/+:Ay/+ mice and plasma leptin concentrations in Tg/+:Ay/+ are higher than regular Ay/+ mice. These indicate that overproduction of leptin can prolong the start of impaired glucose metabolism in Tg/+: Ay/+ mice and endogenous leptin cannot in Ay/+ mice. Leptin can apply its anti-diabetic effect in normal weight animals at week 6. At week 12, Tg/+:Ay/+ mice developed resistance to the anti-diabetic action of leptin. In this study, glucose metabolism is somewhat improved in Ay/+ after a long term body weight reduction due to the 3 week food restriction while the metabolism is improved in Tg/+:Ay/+ compared to Ay/+ and control which suggests that hyperleptinemia enhances glucose level when body weight is stable. Persistent hyperleptinemia delays the beginning of impaired glucose metabolism and quickens the recovery from diabetes in Ay/+ mice in combination with food restriction.
Emilie Vander Haar and her team in the University of Minnesota Minneapolis studied “Insulin signaling to mTOR mediated by the Akt/PKB substrate PRAS40.” In this study, they were able to identify PRAS40 as a crucial regulator of insulin sensitivity of the Akt-mTOR metabolic pathway which can potentially help target the treatment of cancers, insulin resistance, and hamartona syndromes. Insulin activates the protein kinases Akt, also known as PKB, and mammalian target of rapamycin (mTOR) which stimulates protein synthesis and cell growth. This study was able to identify PRAS40 as a unique mTOR binding partner and is induced under conditions that inhibit mTOR signaling. Akt phosphorylates PRAS40, which is crucial for insulin to stimulate mTOR. These findings contribute to the clinical studies of type 2 diabetes insulin related pathways.
mTOR is a kinase-related protein that is a key mediator of insulin. Inhibition of mTOR in mammals proves to reduce insulin resistance and extend lifespan. mTORC1 is a nutrient and insulin regulated complex that is formed from mTOR when it interacts with raptor and a G-protein. This complex is involved in the cytoskeleton regulation and Akt phosphorylation; however the interactions and associated proteins in response to insulin have not been identified. In order to do so, Haar and her team used a mass spectroscopy method. An mTOR antibody prepared mTOR immunoprecipitates from T-cells and the proteins that were bound to the regulator were eluted from the precipitates. The mixtures of proteins were trypsinized and the mass spectra were obtained. The highest P scores obtained from the derived peptides illustrated that mass spectroscopy isolated mTOR-binding proteins. Three sequences were obtained and contributed to the finding that Sin1 is crucial in the formation of the mTOR interaction. The PRAS40 peptide sequence was also identified. However, in order to confirm the hypothesis that mTOR binds with PRAS40, T-cells’ precipitates, which carries PRAS40, were analyzed using western blotting. Compared to the control, PRAS40 was found to bind only with mTOR and nothing else. It was shown that PRAS40 binds specifically in the mTOR carboxy-terminal kinase domain. Certain conditions inhibit mTOR signaling increases affinity, binding abilities, of the PRAS40 mTOR interaction. These conditions include depriving leucine or glucose from the media solution, treatment with the glycolytic inhibitor and mitochondrial metabolic inhibitors. The increase in affinity leads to disrupting the raptor-mTOR interaction, which results with destabilizing the PRAS40-mTOR interactions. This tightened bond between the proteins under nutrient deprivation conditions proposes a hypothesis that states PRAS40 has a negative role in regulating mTOR.
In order to further understand the consequence of PRAS40 in mTOR signaling, the regulator was downregulated in 3T3-L1 and HepG2 cells. The phosphorylation process of Akt at Ser 473 and S6K1 (a mTOR substrate) at Thr 389 was studied. PRAS40 silencing led to a significant decrease in Akt phosphorylation in the cell lines which resulted with negative effects on the Akt components. PRAS40 silencing also led to increased levels of S6K1 phosphorylation, and suggests that the PRAS40-knockout mTOR complex is still active in S6k1 phosphorylation. The mechanism was studied and results indicated that PRAS40 silenced cells and resulting activated state of mTOR may contribute to Akt inactivation – a feedback inhibition. That was the first part of PRAS40 analysis, PRAS40 silencing. In the second part, PRAS40 was overexpressed. Increasing the levels of PRAS40 in cells resulted with decreased S6K1 phosphorylation. These results prove that PRAS40 inhibition of mTOR regulation is likely to require mTOR and raptor binding.
After determining the inhibitory function of PRAS40 in mTOR signaling, Haar and her team studied the role PRAS40 plays in the regulation of mTOR. PRAS40 knockdown in mice and human cells weakened the ability of insulin to stimulate phosphorylation. PRAS40 silencing reduces the levels of phosphorylation in both cell types. In order to further study the response of mTOR to insulin, sample cells were treated with insulin. The data collected proposes that PRAS40 silencing detaches mTOR from Akt signals – PRAS40 plays a crucial role in regulating Akt signaling to mTOR. It also demonstrates that Akt phosphorylation of PRAS40 is crucial for mTOR activation through the use of insulin.
The next matter that Haar and her team touched upon was the study that nutrient starvation has dominant effects on PRAS40-mTOR interaction. PRAS40 was hardly released from mTOR when the conditions are deprived of leucine. Also, the amount of 14-3-3, an interaction induced on PRAS40 phosphorylation, bound to mTOR and PRAS40 was significantly reduced under deprived leucine conditions – the interaction was prevented under non-nutrient conditions. 14-3-3 interactions with mTOR and PRAS40 were also prevented under leucine-deprived conditions. In all, these results prove that PRAS40 is a key mediator of Akt signals to mTOR and a negative effector of mTOR signaling. PRAS40 is a crucial regulator of in insulin sensitivity of mTOR signaling, an important role in insulin resistance.
Some methods of this experiment included the use of antibodies in western blotting, plasmid constructions and mutagenesis, the identification of mTOR-interacting proteins, cell culture and transfection, coimmunoprecipitation, chemical crosslinking, and lentiviral preparation, viral infection, and stable cell line generation. Human PRAS40 cDNAs were provided and mouse PRAS40 cDNA samples underwent PCR amplification and then subcloned into mammalian expression vector. All these cloned samples were confirmed by sequencing. PRAS40 Thr 246 was replaced by amino acids: alanine, glutamate, and aspartate. This is done through a site-directed mutagenesis kit. The way that mTOR immunoprecipitates is through the use of an mTOR antibody on cells cultured in 10% fetal bovine. The cell samples were lysed in a buffer and then incubated with 20ul of protein G resin and 4ug of mTOR antibody. The mTOR precipitates were washed with lysis buffer and the binding proteins were eluted by incubation. The mTOR binding proteins were diluted with digestion buffer and then incubated overnight with trypsin. These samples underwent analysis by mass spectrometry. Data would only be considered accurate when the P score is greater or equal to 0.95. For chemical crosslinking experiments, T cells were treated with dithiobis and then harvested and lysed in a buffer. The precipitates were then analyzed using the SDS-PAGE method. In order to measure the cell-size, T cells were infected with lentiviruses, and then selected in the presence of zeocin. The cell samples were trysinized the following day and diluted 10 times. The ViCell cell-size analyzer analyzed the size of 1.0 mL of diluted cell culture sample.
Overall, the study of PRAS40 in regulating mTOR insulin signaling can potentially lead to potential targets for the treatment of different diseases relating to type 2 diabetes, cancers, and insulin resistance. The results indicate that the Akt/PKB substrate, PRAS40, provide negative effects on the signaling of mTOR. The binding suppresses mTOR activation and insulin-receptor substrate-1 (IRS-1 and Akt, therefore, uncoupling the response of mTOR to Akt signals. PRAS40’s interaction with mTOR is induced under certain environmental conditions such as nutrient, leucine and serum deprivation. In general, this project was able to identify that PRAS40 is a crucial mTOR binding partner that intervenes Akt signaling to mTOR.
Endoplasmic Reticulum Stress Stimuli and Beta-Cell Death In Type 2 Diabetes
[edit | edit source]Obesity is related with insulin resistance, however Type 2 Diabetes, a complex known for increased levels of blood glucose due to insulin resistance in the muscle and liver tissue as well as impaired insulin secretion from pancreatic beta-cells, solely cultivates in genetically predisposed and insulin resistant subjects with the beginning of beta-cell dysfunction. As research progresses there is clear data that demonstrates that beta-cell failure and death are due to unresolvable endoplasmic reticulum stress, bringing chronic and strong activation of inositol-requiring protein 1. Endoplasmic reticulum stress can start and generate the characteristics of Beta-cell failure and death observed in Type 2 Diabetes.
Glucose Transport Deficiency in Type 2 Diabetes
[edit | edit source]GLUT4 glucose transporters migrate to the cell surface in response to insulin signaling, thereby upgrading glucose levels in muscle and fat cells. This is accomplished by stimulating vesicle transport of glucose to where it is needed. Adult onset diabetes is often the result of gradual increase in insulin tolerance in individuals who overeat. This desensitization to the effects of insulin interferes with metabolism because vesicles containing GLUT4 are not able to efficiently fuse with the cell membrane therefore glucose uptake into cells is inhibited. By understanding this pathway, researchers may eventually find a therapeutic workaround to treat those suffering from Type 2 diabetes. Presumably, this could be accomplished by synthesizing molecules that mimic the function of GLUT4 and its auxiliaries to resolve the trafficking problem. Alternatively, the insulin pathway could be targeted.
RESOURCES
[edit | edit source]1. http://www.diabetes.org/diabetes-basics/diabetes-statistics/
2. http://www.ncbi.nlm.nih.gov/pubmedhealth/PMH0001350/
3. http://www.ncbi.nlm.nih.gov/pubmedhealth/PMH0001356/
4. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC2682681/
5. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3161334/
6. http://diabetes.niddk.nih.gov/dm/pubs/statistics/
7. http://diabetes.diabetesjournals.org/content/60/9/2354.full
8. http://www.mayoclinic.com/health/type-1-diabetes/DS00329/DSECTION=treatments%2Dand%2Ddrugs
9. http://www.mayoclinic.com/health/type-2-diabetes/DS00585/DSECTION=treatments%2Dand%2Ddrugs
10. http://diabetes.diabetesjournals.org/content/48/9/1822.short
11. http://www.ncbi.nlm.nih.gov/pubmed/22482906
12. http://www.nature.com/ncb/journal/v9/n3/pdf/ncb1547.pdf
References
[edit | edit source]1. Endoplasmic reticulum stress and type 2 diabetes. Back SH, Kaufman RJ. Annu Rev Biochem. 2012;81:767-93. Epub 2012 Mar 23. Review. PMID: 22443930 [PubMed - indexed for MEDLINE]
Overview
[edit | edit source]Alzheimer's is a form of dementia, a decline in mental ability that affects everyday life. This disease attacks the brain and causes problems associated with memory, thinking, and behavior. As time progresses, the symptoms usually get worse. It is usually assumed that Alzheimer's is a result of aging; however this is not the case. Aging simply increases the risk factor of obtaining this disease. As of now, there is no cure for Alzheimer's; treatments usually are only able to slow the disease from progressing.
In association with Alzheimers disease are peptide proteins known as β-Amyloid. They are found in the brains of humans diagnosed with Alzheimers Disease. Researchers have been trying to study these peptides so that they can work to find a cure (or treatment) that would help the patients. To understand so, it requires understanding the structure of the proteins. Because of the complex structures, there has been limited compilations, but there has been progress. [1]
Symptoms
[edit | edit source]Memory – Memory loss associated with Alzheimer’s disease persists and only gets worse. Some symptoms may include:
- Repeating the same sentence over and over
- Forgetting conversations, past events, or future appointments
- Misplacing of possessions
- Forgetting names of family members and relationships
- Difficulty understanding surrounds, may not know when or where
Speaking, Writing, Thinking, and Reasoning
- Initially having trouble finding the right words during a conversation
- Eventually lead to the loss of speaking and writing abilities
- May eventually having trouble understanding conversation or written text
- Poor judgment and slow response
Changes in personality and behavior
Brain degradation of Alzheimer’s may affect how people feel. People with Alzheimer's may experience:
- Depression
- Anxiety
- Social withdrawal
- Mood swings
- Distrust in others
- Increased stubbornness
- Irritability and aggressiveness
- Changes in sleeping habits
As the disease progresses, the symptoms could only increase in severity to more severe memory loss, confusion about events in regards to time and space, and disorientation. To try and understand the correlation between the symptoms and the disease, there must be an understanding of the structures of the proteins that have commonly been found in all the patients- β-amyloid (Aβ) peptides. Many people who are affected with Alzheimer's require continuous attention and care since they are unable to perform even basic daily activities.
Test For Alzheimer's
[edit | edit source]A diagnosis of Alzheimer’s disease may include a complete physical and neurological exams, CT (computed tomography), and MRI(magnetic resonance imaging). Biopsy of the brain and identifying evidence of any of the following: Neurofibrillary tangles, Neuritic plaques, and Senile plaques. Neurofibrilary tangles are twisted filaments of proteins within nerve cells that clog up the cell, inhibiting neurotransmitters and the function of the nerve cell. Neuritic plaques are abnormal clusters of invalid or dying nerve cells. Senile plaques are areas of waste products around proteins that were produced by dying nerve cells. All of these may be caused by or inducing Alzheimer’s disease.
Amyloid Fibrils
[edit | edit source]
β-amyloid (Aβ) peptides segregate into different domains. Included are amyloid fibrils, protofibrils, and oligomers. These have been under study in hopes to understand Alzheimers Desease. Unfortunately, understanding their whole structure has been difficult. The B-ayloid (AB) peptide forms naturally in the human body within in the brain as a protein precursor of a proteolytic fragment. It is specifically the AB amyloid fibriles that form the core of a dense plaque within the brain leading to alzheimers disease.
Amyloid fibrils are known as fibrillar polypeptide in collection with an intermolecular cross-β structure. X-ray diffraction showed that the B-strands hydrogen bond with each other and orient in a parallel manner along an axis. The B-amyloid (AB) peptide is amphiphilic having a hydrophobic C-terminus lasting for about 37-42 residues, and a hydrophilic N-terminus. These structures twisted as crossovers, and estimated to have a length of about 1 micrometer. Unique about the AB fibrils are their polymorphism. This refers to their ability to conform to different arrangements. These arrangements include the fibrils different in the number of protofilaments, differing in their orientation, and differing in their substructure. These differences is relevant for humans because it could contribute to folding , reactions, and eventually to the level of alzheimers that the human has. [1]
Aside from the polymorphism, there is further diversity among the amyloid peptides due to structural deformations. This includes different bends and twists. These deformations allow study of nanoscale mechanical properties of the fibrils.
Structure of β-amyloid
[edit | edit source]The β-amyloid peptide is a natural forming proteolytic peptide found in the human brain. It is intrinsically unstructured, meaning that it lacks a stable tertiary structure. Many β-amyloid peptides have disordered and unfolded structures that can only be observed using NMR (nuclear magnetic resonance). The peptide is amphipathic, possessing a hydrophilic N-terminus and a hydrophobic C-terminus. The C-terminus can bind up to 36-43 amino residues, which creates the overall structure of the peptide chain. A great number of β-amyloid isoforms differ by one amino residue; many are closely related to Alzheimer’s. β-amyloid undergoes many complex fibrillation pathways, creating intermediate structure such as oligomers, amyloid derived diffusible ligands, globulomers, paranuclei, and protofibrils. When any of these intermediates decide to plaque the walls of cerebral blood vessels, Alzehimer’s disease may be underdevelopment.
The cross-β sheet structure of a β Amyloid
[edit | edit source]Many different amyloid-like polypeptides show a common cross β-sheet structure. These β-sheets are perpendicularly attached microcrystal backbone through non-covalent and hydrogen bonds, and they have parallel conformations between sheets. Recent studies have shown crystallographic evidence of these microcrystal called steric zippers, which are present in many amyloid fibrils. Steric zipper is a structure of a pair of two cross-β sheets with side chains that resemble a zipper. There are dry and wet interfaces of the cross β-sheet conformation. The wet interface is covered by water molecules, which create a greater distance between two adjacent sheets. The dry interface does not contain water so the distance between two adjacent sheets is much closer. While the polar side-chains of the wet interface is stabilized by hydrogen bond interaction, the side-chains of the dry interface are integrated by adjacent side-chains by stacking the previously mentioned steric zippers. Different β-amyloids with different lengths of residues favor either the parallel or antiparallel form. A β-amyloid can also have segmented parallel and antiparallel structures. For example, residues 1-25 would have one conformation. Residues 26-43 would have the other. From These β-sheet structures assign many distinguishable properties to β-amyloid. For example, β-amyloids have a high affinity to specific dyes such as Congo red and Thioflavin T. These dyes can help mark and track the activity of β-amyloids inside the brain.
Models of Amyloids
[edit | edit source]There have been two forms of Aβ peptides that have been under study: Aβ(1-40) and Aβ (1-42). The numbers 40 and 42 refer to their respective amount of residues.
The Aβ(1-40) is proposed to be more pathogenic than the Aβ(1-42) form. When experimented with the model Drosophilia melanogaster, the Aβ(1-42) showed to be toxic and result in a shorter life-span. Aβ fibrils are a big factor leading to the alzheimers disease. It’s been hypothesized that it is toxic and eventually kills the cells that come in contact by penetrating their membranes. It’s suggested that the activity of these peptides are intracellular rather than extracellular. AB amyloid fibrils are complex units segregate into different populations. To try and understand Alzheimers Disease would mean having to understand the population of the Aβ amyloid fibrils. Doing so will allow researchers and scientist work for the disease treatment
It is often difficult to isolate the Aβ amyloid peptides, thus the amount of information obtained from them is very limited. The full-length structure of the AB amyloid fibrils have yet to be uncovered, even with use of X-ray crystallography. Many other forms of measurements have been used to study the Aβ- amyloids. These include infrared spectroscopy, NMR, mass spectrometry, electron paramagnetic resonance. Unfortunately, the data received are rather indirect. The most direct way would be to use solid state NMR and electros cryomicroscopy (cryo-EM). These allow the distinction of Aβ amyloid fibrils at near-atomic resolution. They give chemical shifts and even the bond angles. From here it allows the researchers to ID the residues and their sheet structure.
There have been many models of the Aβ peptides proposed. But because it must be considered that in different conditions different fibrils can conform, it’s critical to have much caution. The general model of an Aβ fibril is a U-shaped peptide told, refered as a β-arc. [2]
A Molecular Link Between the Active Component of Marijuana and Alzheimer's Disease Pathology
[edit | edit source]Recent studies have shown that the active component of marijuana, Δ9-tetrahydrocannabinol (THC), inhibits AChE-induced β-amyloids aggregation in the pathology of Alzheimer’s disease. Some studies have demonstrated the ability of THC to provide neuroprotection against the toxicity of β-amyloid peptide. One of the causes of Alzheimer’s disease is the deposition of β-amyloid in portions of the brain that are important for memory and cognition. This deposition and formation of a plaque in the brain is caused by enzyme acetylcholinesterase (AChE). AchE is an enzyme that degrades acetylcholine, which in turn increase the amount of neurotransmitters released into the synaptic cleft. It also functions as an allosteric effector that accelerates the formation of amyloid fibrils in the brain. In vitro studies have demonstrated the inhibition of AChE has decreased β-amyloid deposition in the brain, and THC is a very good inhibitor.
THC binding to AChE using AutoDock revealed that THC has a high binding affinity to AChE. Not only do they bind well, interactions were observed between THC and the carbonyl backbone of AChE, residues of Phe123 and Ser125. Furthermore, the ability of THC to inhibit AChE catalytic activities were tested using steady-state kinetic. The results have shown that THC inhibits AChE at a Ki of 10.2 uM. This number is relatively competitive with the current drugs in the market that treat Alzheimer’s disease. While THC shows competitive inhibition relative to the substrate, this does not necessitate a direct interaction between THC and the AChE active site. In fact, enzymes can bind to the PAS allosteric site on AChE while still blocks the entry into the active site of AChE, preventing it from depositing plaque. This why THC serves as an uncompetitive inhibitor of AChE substrate.
References
[edit | edit source]"What is Alzheimer's?." Alzheimer's and Dementia. Alzheimer's Association, 2012. Web. 20 Nov. 2012. <http://www.alz.org/alzheimers_disease_what_is_alzheimers.asp>. Fandrich, Marcus, Schmidt, Matthias, and Nikolaus Griforieff: Trends Biochem Sci. Author Manuscrpt: Recent Progress in Understanding Alzheimer's B-amyloid Structures. 2011 June; 36(6) 338-345.
Sipe, J.D. Amyloidosis. A. Rev. Biochem. 61, 947−76 (1992).
Glenner, G.G. Amyloid deposits and amyloidosis: The -fibrilloses (first of two parts). New. Engl. J. Med. 302, 1283−1292 (1980). | PubMed | ISI | ChemPort |
- ↑ a b Fandrich, Marcus, Matthias Schmidt, Mikolaus Grigorieff (February 2011). "Recent Profress in Understanding Alzheimer's B-amyloid Structures". Trends Biochemistry. 36 (6): 338–45. doi:10.1002/ana.410380312. PMID 7668828.
{{cite journal}}: CS1 maint: multiple names: authors list (link) Invalid<ref>tag; name "pmid7668828" defined multiple times with different content - ↑ Roher AE, Lowenson JD, Clarke S, Woods AS, Cotter RJ, Gowing E, Ball MJ (November 1993). "beta-Amyloid-(1-42) is a major component of cerebrovascular amyloid deposits: implications for the pathology of Alzheimer disease". Proc. Natl. Acad. Sci. U.S.A. 90 (22): 10836–40. Bibcode:1993PNAS...9010836R. doi:10.1073/pnas.90.22.10836. PMC 47873. PMID 8248178.
{{cite journal}}: CS1 maint: multiple names: authors list (link)
Overview
[edit | edit source]

Despite the many pharmacological advancements achieved in the past few decades through structural biochemistry, prescribed medications work in fewer than fifty percent of the patients who take them. The reason for this is that, while mostly similar, everyone’s genome is slightly different and responds differently to the same medications. The underlying cause is that there exist variants in the genes that make Cytochrome P450. Cytochrome P450, an example of which is pictured at right, refers to a large and diverse family of enzymes that process the drugs that we take. Therefore, there are as many different responses to the drugs that we take as there are variants in the gene.
If one knows a patients entire genome, however, it would be relatively easy to predict the types of medications that would work and which ones would be least effective. This is the idea behind personalized medicine, a medical model that uses information from a patient’s genome and proteome to optimize his or her medical care. Personalized medicine is the ultimate goal on the medicinal stairway model, pictured at right. The lowest step is the use of blockbuster drugs; the more advanced step is the stratified medicine level; and the top step is personalized medicine, the most specific and accurate of the three techniques to patient care.
In the previous fifty years, the primary medical model has been that of the “blockbuster drugs,” or medicines that work for a majority of the generic population. Specifically, a blockbuster drug refers to a drug that generates more than $1 billion of revenue for the patent owner each year. Some examples of past blockbuster drugs are Lipitor, Celebrex, and Nexium. Leading scientists in the field of biochemistry acknowledge that we are slowly leaving the blockbuster drug era and are in the midst of moving to the second level, stratified medicine.
Stratified medicine refers to managing a patient group that has shared biological characteristics, such as the presence or absence of a gene mutation. Molecular diagnostic testing is used to confirm these similarities; then, the most optimal treatment is selected in hopes of achieving the best possible result for the group. An example of stratified medicine in practice is grouping patients with breast cancer who have estrogen receptor positivity or HER2 over-expression, and who can be treated according to these characteristics with an anti estrogens or a HER2 inhibitor.
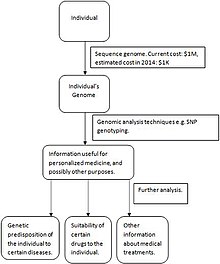
Lastly, personalized medicine is the desired goal we have yet to fully reach. Proteomic profiling, metabolomic analysis, and genetic testing of each individual are required to optimize preventative and therapeutic care. The diagram pictured on the right summarizes the steps needed for personalized medicine to be successful. First, an individual’s genome is sequenced. As technological sequencing techniques advance, the cost of sequencing a genome will decrease, making the personalized medicine model more accessible and more economical. Then, genomic analysis techniques such as SNP genotyping and microarrays are used to gather information about which medicines will work best (for example, regarding an individual’s genetic predisposition toward certain diseases and how long a certain drug will be effective). A current example of the progress made toward personalized medicine is the measurement of erbB2 and EGFR proteins in breast, lung and colorectal cancer patients are taken before selecting proper treatments. As the personalized medicine field advances, molecular information elucidated from tissues will be combined with a patient’s medical and family history, data from imaging, and a multitude of laboratory tests to develop more effective treatments for a wider variety of conditions.
Because everyone has a unique set of genome, advantages of having personalized medicine through pharmacogenetic approaches include:
1. Increase effectiveness of the drug For example, using the right medicine and dosage in order to allow it absorb more easily by a patient’s body
2. Minimize side effects
References
[edit | edit source]PricewaterhouseCoopers’ Health Research Institute,(2009). [The new science of personalized medicine] http://www.pwc.com/personalizedmedicine
Shastry BS (2006). "Pharmacogenetics and the concept of individualized medicine". Pharmacogenomics J. 6 (1): 16–21.
Pharmaceutical Market Trends, 2010-2014, from Urch Publishing
Jørgensen JT, Winther H. The New Era of Personalized Medicine: 10 years later. Per Med 2009; 6: 423-428. A model organism is an indispensable tool used for medical research. Scientists use organisms to investigate questions about living systems that cannot be studied in any other way. These models allow scientists to compare creatures that are different in structures, but share similarities in body chemistry. Even organisms that do not have a structural body, such as yeast and mold, can be useful in providing incites to how tissues and organs work in the human body. This is because enzymes used in metabolism and the processing of nutrients are similar in all living things. Other reasons model organisms are useful are that they are simple, inexpensive, and easy to work with.
Examples of model organisms:
Escherichia Coli: Bacterium
[edit | edit source]There are good and bad bacteria. The one form of bacterium one is usually familiar with is the E. coli that is associated with tainted hamburger meat. However, there also exist "non-disease-causing" strains of E. coli. in the intestinal tracts of humans and animals. These bacteria are the main source in providing vitamins K and B-complex. They also help in the digestive system and provide protection against harmful bacteria. Differentiating between harmful and helpful strains of E. coli can help distinguish the genetic differences between bacteria in humans and bacteria that cause poisoning.
Dictyostelium Discoideum (Dicty): Amoeba
[edit | edit source]Amoeba is microscopic cell which is 100,000 times smaller than a grain of sand. This organism has between 8,000 and 10,000 genes and many of them are similar to those in humans and other animals. Dicty cells usually grow independently. However, with limited food resources, these cells can pile on top on each other to form a multicelled structure of up to 100,000 cells. When migrating, this slug-like organism will leave behind a trail of slime. They can disperse spores that are capable of generating new amoeba.
Neurospora Crassa: Bread Mold
[edit | edit source]This type of model organism is used world wide in genetic research. Researchers like to use bread mold because it is easy to grow and can answer questions about how species adapt to different environments. Neurospora is also useful in the studying of sleep cycles and rhythms of life.
Saccharomyces Cerevisiae: Yeast
[edit | edit source]Yeast is commonly used in research, but it is also an important part of life outside the laboratory. It is a fungus and has eukaryote properties. Researchers prefer yeast because it is fast to grow, cheap and safe to use, and easy to work with. Yeast can be used as a host for mammalian genes, and it allows scientists to study how they function inside the host. Discoveries of the antibiotic penicillin or the protein called sirtuin that interferes with aging resulted from observing fungus.
Arabidopsis Thaliana: Mustard Plant
[edit | edit source]Arabidopsis is a flowering plant that is related to cabbage and mustard. Researchers often use this plant to study plant growth because it has very similar genes with other flowering plants and little encoded-protein DNA, which makes it easy to study genes. Arabidopsis has eukaryotic cells, and the plant can mature quickly in six weeks. Cell communication in plants operates much like human cells do, and this makes it easier to study genetics.
Caenorhabditis Elegans: Roundworm
[edit | edit source]Roundworms are as tiny as the head of a pin, and they live in dirt. In the laboratory, they live in petri dishes and feed on bacteria. This C. elegans creature has 959 cells and one third of the cells form the nervous system. Researchers like to use roundworm because it is transparent, which allows a clear view of what goes on in the body. C. elegans has more than 19,000 genes compare to a human of about 25,000 genes. C. elegans is the first animal genome to be decoded, and the major of the genes is similar to that of humans and other organisms.
Drosophila Melanogaster: Fruit Fly
[edit | edit source]This type of fruit fly is most commonly used in research. Fruit flies in the laboratory are often exposed to harmful chemicals and radiation that can change their DNA sequences. Researchers allow flies to mate and then study their offspring for mutations. The mutant flies help researchers to study detective genes. Fruit flies can reproduce quickly, which makes it easy to create mutant flies, and this enables researchers to study how genes function. By relating some of the defects found in fruit flies with those in humans, researchers may discover the defective genes as well.
Danio Rerio: Zebrafish
[edit | edit source]Zebrafish habitat in slow streams, rice paddies, and the Ganges River in East India and Burma. They are also found in pet stores. Researches prefer zebrafish because their eggs and embryos are transparent, which provide a plain view of the development process. It takes only 2 to 4 days for zebrafish cells to divide and form the fish's body parts: eyes, heart, liver, and etc. This research enables scientists to study birth defects, the proper development of the heart and blood.
Mus Musculus: Mouse
[edit | edit source]Mice are mammals like humans, and we share 85 percent of our genes. Because mice are very similar to people, they are used to study diseases in humans. Scientists can create "knockout" mice, missing gene mice, and study how the mice function.
Rattus Norvegicus: Rat
[edit | edit source]Rat was the first animal to be used in scientific research. For the most part, laboratory rats are used to test drugs and most of what we know about cancer started with rat research. Rats are mammals and are bigger than most model organisms, which makes it easy for scientists to perform experiments on the rat brain. Scientists have learned about substance abuse and addiction, learning and memory, and neurological diseases through rats. They are also useful in the studying of asthma, lung injury, and arthritis disease.
Pan Troglodytes: Chimpanzee
[edit | edit source]The fact that chimpanzees share 99 percent of the genes with humans makes them very unique for studying human genome. Because they are immune to malaria or AIDS, ongoing medical research is trying to discover the reason at genomic level.
Lambda Phage: Virus
[edit | edit source]A lambda phage is a bacterial virus, or bacteriophage that infects a bacterial species such as escherichia coli. This virus has a temperate lifestyle meaning that it may well reside within the genome of its host until its lyses out of its host, also known as lysogeny. The lambda phage consists of constituent parts that all take part in its integration in the host genome. The phage contains a capsid head, a tail, and tail fibers for which is used to latch on to its host. The importance of this model organism to scientists is its ability to incorporate itself into its hosts genome allowing to integrate its genomic DNA into its host. This is especially helpful to scientists for genomic work.
Chlamydomonas reinhardtii: Green Algae
[edit | edit source]This model organism is a unicellular green algae that is primarily used to study the mechanisms of photosynthesis, regulation of metabolism, cell to cell recognition, adhesion, response to nutrient deprivation and flagella motility. It proves to have a great significance to scientists because it can grow in media lacking organic carbon and chemical energy sources. In addition this model organisms is associated with fluorescence and can grow in the dark when supplied with other unicellular green algae as a source of hydrogen.
Reference
[edit | edit source]The New Genetics. National Institute of General Medical Sciences. Revised October 2006. http://publications.nigms.nih.gov/thenewgenetics.
Introduction
[edit | edit source]Clinical and basic research are two different worlds, but there needs to be a bridge connecting those (which creates translational research) to be able to open more doors with research opportunities by utilizing both types of research. There are difficulties of translational research, thus there needs to be a new generation of scientists who are willing to be educated and participate in translational research efforts, and new programs are being created to facilitate these efforts.
From the laboratory to the patient's bedside
[edit | edit source]There are many examples about how basic science has led to a discovery or a new drug even when it's initial motive was to run experiments only for the scientist's own interest. For example, pathologists were observing cholesterol in atheromatous plaques and studies in the '60s shows that serum cholesterol and coronary artery disease have an association with one another. Based on the knowledge of biochemical pathways of cholesterol synthesis and cholesterol transport, inhibitors of HMG-CoA reductase were created. Physicians and the public were educated on this drug and also clinical trials were implemented to prove the drug was effective and safe. In the end, the drugs were adopted into the public and reduced the mortality rates of coronary disease.
Clinical and Translational Science Awards (CTSA) Consortium
[edit | edit source]CTSA was founded to help support and fund various translational research opportunities. There are six objectives: 1. There needs to be training of individuals across the translational spectrum; MDs need to be educated to become translation investigators and PhDs need to be educated i medical pathophysiology so if a basic finding has a direct translatability, they will know how to approach this finding. 2. There needs to be simplification of the translational process to become very efficient and accelerate the process, but while still keeping the research subject safe. 3. There needs to be the best types of technology in order to provide the best research for translational investigation. 4. There needs to be programs to nurture the careers of translational researchers. Thus, there are MD/PhD programs to bridge the divide of basic and clinical research and the success of these programs will allow tuition support so trainees are overwhelmed with huge loans. 5. There needs to be team mentoring for junior clinical scientists with programs such as the K12 awards. 6. There needs to be cataloguing research resources so tools will be available to a wide set of users.
Potential future challenges
[edit | edit source]One of the challenges translational research faces is that translational research is only successful if many people or teams are working together to create a certain outcome. Thus, it is hard to figure out who will be considered the first author for the published article, who gets the grant, the promotion, and more.
Another is the problem of "pipeline" findings. Translational research hones down on a certain topic or research in order to find a drug or discovery, but basic research, the topics are very broad. Also, many discoveries or drugs were found out by accident by basic researchers. How will basic research turn out if more money and grants are being handed to translational efforts?
Lastly, it is unknown how pharmaceutical companies would react to the boom of rich data and information because of translational efforts. It is implied that pharmaceutical companies will then mass produce drugs (produce millions) instead of producing thousands.
Reference
[edit | edit source]McClain, Donald A. "Bridging the gap between basic and clinical investigation" Trends in Biochemical Sciences 35.4 (2010) 187-188. Academic Search Complete. Web. 21 Nov. 2012.
What is it?
It is the study of tissues of plants and animals. Histologist usually use tissues, which are groups of specialized cells and promote staining onto these tissues after they are sectioned. There are typically two types of tissue section preparations. Usually one can freeze the tissue and then section them into 30-60 microns, or a “paraffin-embedded” method which is where hot wax is poured over the tissue of interest and molds into a block. The block is then sectioned using a microtome and the tissues are usually much thinner.
History
Marcello Malpighi a contemporary of Robert Hooke, scientist who discovered the cell was known to be the first histologist. He was able to use archaic microscopes from the 17th century and study the structure and component of tissues. However, another scientist named Marie Francois Bichat, who came after Malpighi introduced the word tissue and put the framework of many cells working collectively together for a specific function. Some consider Bichat to be the founder of present day Animal Histology. In modern day, histologist usually only collect four main types of tissues, which include the connective, epithelial, muscle and nerve tissues. It was not until after Bichat's death did the word histology began to circulte. Histology derives from two Greek words, Histo, meaning tissue, and logo meaning study. However, some consider Rudolph von Kolliker the actual father of Histology primiarly because of his work on creating an actual textbook and emphasizing tissues, this book is called "Handbuch der Gewebelehre".
Tissue fixation
Is the method by which scientist stop enzymatic activity and preserve the tissue in its most natural state. The fixatives, chemicals used in tissue fixation are usually toxic and will disinfect bacteria and parasites for a period of time. However, if upon fixating the tissue, overexposure to certain fixatives may lead to masking of proteins by masking epitopes. Usually, tissues are fixed by methods of immersion or perfused fixation. In order to do so formaldehyde is used in water. Paraformaldehyde is also available to be used as well.
Freezing
Is necessary in order to preserve the integrity of the tissue component and site of study. However, because much of tissue has water within in, another solvent must be used to drive water out as to not damage the tissue upon freezing. This can be done by using sucrose as solvent/media. Much like osmosis, as concentration of sucrose outside of the tissue increases water will rush out of the tissue to equilibrate the solution it is in. This process is called cryoprotection.
Sectioning slides
The method of sectioning varies from lab to lab, some labs use the paraffin technique while others use the freezing method.
Frozen Slide Method
Before samples are prepared they must be placed in sucrose for a relative day or two to drive the water out by osmosis. Upon dissection of the tissue of interest a section of the tissue must then be made and placed upon a container. Tissue freezing medium is then applied to the container and the sample is placed with it. Freezing under dry ice is then applied and then another layer of tissue freezing medium is placed so that it covers entire container. After letting the medium freeze for about half an hour, a square cut is made. The frozen cut is then placed on a mounting disc and tissue freezing medium is applied once again in order to mount the sample onto the disc. Once this is done and the disc is then mounted onto the microtome or cryostat sectioning can then begin. Upon mounting the disc on the cryostat, a blade is then applied to cut the sections. One must first however align the sample so that the cut is applied in a planar fashion. Cuts can be applied in various ways depending on how the user wants to obtain their sections.
References:
Tuszynski Lab - UCSD
http://erocha.freehosting.net/histologystuff.htm
| This page or section is an undeveloped draft or outline. You can help to develop the work, or you can ask for assistance in the project room. |
What types of Stains are there?
There are a variety of different stains out there to be used for variety of tasks. Each staining method and reagent are similar to that of antibody/antigen effects and are used to study the tissues of organisms. Each specific stain will allow the researcher to distinguish what is happening overall within the tissue but does not provide in-depth analysis of the individual interactions within the cells of the tissue. H and E staining
H and E staining is an acidophilic, basocphilic staining method. This means that there is a component of an acid and a component that contains a base within this staining method. Basophilic, meaning something that likes bases. Therefore, acids would react to this such as nucleic acids. Acidophlic meaning proteins that are in the cytoplasm would react to acidic dyes because they like acids or negatively charged molecules. In this case H, which stands for hematoxylin is a basic dye, this means that it is positively charged and stains acidic structures such as nucleic acids. Hence, following this Eosin would be acidic and would then stain any structures that are basic. Eosin stains the bases with a pink color and hematoxylin stains the acids with a purple color.
Definition
[edit | edit source]Neuroscience is a science that describes the study of the nervous system. It involves studying its anatomy, chemistry, physiology, development, and functioning. It is an interdisciplinary science that involves psychology, mathematics, physics, chemistry, engineering, computer science, philosophy and medicine. It should be noted that there is a difference between neuroscience and neurobiology. Neurobiology specifically refers to the biology of the nervous system whereas neuroscience refers to the entire science (chemistry, physics, etc.) of the nervous system. Neuroscience has become a enormously popular field in the last few years; for example, the Society for Neuroscience currently has about 30,000-40,000 members, maybe more.
Alzheimer's Disease
[edit | edit source]Alzheimer’s disease is a form of dementia. It is a degenerative, terminal disease that is currently incurable. Death of neurons in the hippocampus of the brain causes memory impairment in those with Alzheimer’s disease. As the neurons degenerate, neurofibrillary tangles develop. Neurofibrillary tangles are intracellular accumulations of hyperphosphorylataed tau. There is now evidence that a truncated p73 isoform, ΔNp73, protect the neurons against tau hyperphosphorylation and tangle formation. It prevents tangle formation by inhibiting c-Jun N-terminal kinase (JNK). Furthermore, ΔNp73 protects neurons against cell death by antagonizing p53. P53 mediates programmed cell-death in a process called apoptosis. It kills neurons by inducing the expression of pro-apoptotic proteins. P53 over expression results in tau hyperphosphorylation in cells. The new discovery that ΔNP73 protects neurons from tau hyperphosphorylation as well as tangle formation provides insight for potential therapeutic roles of ΔNp73 inducers. Drugs can be developed to enhance the neuroprotective actions of ΔNp73.
Alzheimer's Disease (AD) seems to be initiated by the dysfunctional activities of two proteinases (γ- and β-secretase) which generate a series of aggregation-prone peptides called Aβ from their substrate, amyloid precursor protein (APP). The amount of Aβ peptides that accumulates is believed to be the main factor that induces neuronal dysfunction and death.
Parkinson's Disease
[edit | edit source]Parkinson's disease is a motor disorder caused by the death of neurons in the midbrain area. These neurons are typically capable of releasing dopamine, a biogenic amide neurotransmitter, at synapses located in the basal nuclei. When the neurons are destroyed, nerve cells are no longer able to send signals that allow for communication between the substantia nigra and corpus striatum in the brain. Lack of these signals affect the control of muscle movement. The symptoms that result include tremors in the limb or muscle, slowed movement and poor balance. Currently, little is known about the cause for the destruction of neurons and why Parkinson's disease occurs. However, molecular studies have been able to link genetics to rare cases that occur in young adults. Consequently, this has led to the belief that this disease may be inherited although there is still much controversy surrounding this discussion. Scientists are also determining whether or not defects in genes required for mitochondrial function has also been linked to and early onset of this disease.
Similar to Alzheimer's, Parkinson's disease is more common as people get older. For adults at age 65, around 1% are at risk for Parkinson's while at age 85, this rises to 5%. There are nearly 1 million people in the US today that suffer from Parkinson's disease. There is currently no cure for Parkinson's disease. However, there are methods used to maintain the symptoms that include brain surgery, drugs that have the ability to be converted into dopamine and cross the blood brain barrieras well as deep brain stimulation. In laboratories, scientists have also experimented using rats with a similar induced condition. They place dopamine secreting neurons in the midbrain or basal nuclei that help control motor functions. Whether or not this technique would work on humans is still being researched.
Sources
[edit | edit source][1] Carlson, Neil R. Physiology of Behavior. Boston: Pearson Education, Inc., 2007.
[2] Glass, Jon. "What Causes Parkinson's? Age, Genetics, Environment, and Other Factors." WebMD. WebMD, n.d. Web. 27 Oct. 2012. <http://www.webmd.com/parkinsons-disease/parkinsons-causes>.
[3] Reece, Jane B. Campbell Biology, 2011
Reference
[edit | edit source]1. Molony A, et al. “Alzheimer’s disease: insights from Drosophila melanogaster models.” Trends Biochem Sci. 2010 Apr;35(4):228-35. Epub 2009 Dec 25. The neuron is a cell that transmits information via electrical and chemical signaling throughout the nervous system. There are many different neurons of various sizes and function.
Components
[edit | edit source]
All the brain and nervous system are based on communication among nerve cells, known as neurons. Each neuron is like any other cell in the body. Each neuron is surrounded by a membrane and filled with liquid and has a nucleus containing its genetic material. Neurons are specialized to receive and transmit information. All of the neurons gather information either from other cells of the body or from the environment. They transmit information to other neurons and/or other kinds of cells. A typical neuron has an enlarged area, which is the cell body. The cell body contains the nucleus. Neurons have branches or nerve fibers. The branches on which information is received are known as dendrites. The dendrites are a branched structure that receives signals from other cells. Each neuron has a longer tail-like structure, or axon. The axon transmits information to other cells. Axons can be branched at the tips. The axons of many kinds of neurons are surrounded by a fatty, segmented covering called the myelin sheath. The covering acts as a kind of insulation and improves the ability of axons to carry nervous system signals rapidly. A single neuron may be capable of receiving messages simultaneously on its dendrites and cell body from several thousand different cells. Most neurons usually have a soma, dendrites, axons, and a terminal button. The soma is the cell body of the neuron, and it contains the nucleus and is responsible for many of the processes of the cell.
Classification
[edit | edit source]Neurons are classified based on (1) the number of extensions from the cell body (soma) and (2) the neuron's function:
Sensory neurons receive sensory signals from sensory organs. These signals are then sent to the central nervous system via short axons. These neurons are also called Pseudo-unipolar neurons due to the short extension that divides them into two branches. One of these two branches functions as an axon, while the other functions as a dendrite.
Motor neurons are neurons that control motor movements of the body. They take commands from the cortex and send the signal to the spinal cord, or to the spinal cord to the muscles. Motor neurons and interneurons make up the family of multipolar neurons that possess a single axon and many dendrites.
Interneurons also called, associated neurons, are neurons that interconnect various neurons within the brain or spinal chord. The majority of these neurons are seen in the brain, connected densely. These neurons relay information and conduct signals between neurons. Interneurons may also be called bipolar neurons due to their two main extensions, a single dendrite and an axon.
Interneurons can be either efferent neurons, which carry signals away from the brain, or afferent neurons, which carry signals to the brain.
Neuron Classification
[edit | edit source]Neurons can be organized in two ways: 1) based on anatomy and 2) based on function. Neurons can be Pseudounipolar, bipolar, anaxonic or multipolar. Pseuounipolar neurons have a single axon and their soma are off to the side. Bipolar neurons have axons extending off both sides of the soma. Anaxonic neurons have no obvious axon but have a soma and dendrites. Multipolar neurons do not have long axons, but have extremely branched dendrites. Pseudounipolar and bipolar neurons are sensory (or afferent) neurons. Anaxonic and multipolar neurons are interneurons within the Central Nervous System. Multipolar neurons also function as efferent neurons.
Neuron Degradation
[edit | edit source]Neuron replacement is limited in the brain, though neuron phagoptosis does occur. Viable neurons are phagocytosized by lipopolysaccharide (LPS). Microalgia is responsible for the eating of viable neurons, but it is also responsible for eating apoptopic neurons which may be beneficial because it may reduce debris and inflammation. Inflammation in the brain can cause microalgia to eat viable neurons, however this can be blocked by blocking phagooptic signaling. Microalgia also kills developing neurons in the protein in the cerebellum and hippocampus.
Synapse
[edit | edit source]The synapse is a junction between the terminal button of an axon of one neuron and the dendrite of another neuron. In this junction, one neuron sends information to another neuron via electrical or chemical signaling. The process for sending information is called action potential where electrical impulses are sent down an axon of a neuron.
Also, the synapse is a small gap, or commonly referred to as a connection, between two cells that allows for the first cell (the presynaptic cell) to communicate with the second cell (the postsynaptic cell) through a chemical signal. These chemical signals are called neurotransmitters, and once they are released by the presynaptic cell, they act on the postsynaptic cell through specialized protein molecules called neurotransmitter receptors.
A synapse is a connection which allows for the transmission of nerve impulses. Synapses can be found at the points where nerve cells meet other nerve cells, and where nerve cells interface with glandular and muscular cells. In all cases, this connection allows for the one-way movement of data. The human body contains trillions of synapses, and at any given time, huge numbers of these connections are active.
Axonal Transport
[edit | edit source]Axons lack ribosomes and an endoplasmic reticulum and because of this, the cell body must synthesize proteins and send them through the axon via axonal transport. There are two main types of axonal transport: slow and fast. Slow axonal transport is used for moving proteins through the axon that are not used up quickly by the cell, such as enzymes and cytoskeletal proteins. Fast axonal transport is used for moving proteins down the axon that are needed much more quickly in the cell, such as organelles.
Microtubules and Their Role in Axonal Transport
Microtubules provide a crucial role in fast axonal transport systems that supply synaptic vesicles with vital chemical messengers by providing the long cells with highways for material to be transported through. Two families of proteins, Dynein and Kinesin, are in charge of vesicle transportation through microtubules. With Dynein being charge of retrograde transport and Kinesin being in charge of ante-retrograde transportation, their combined proportions provides the axon with variability in transport velocity as well as the potential for intentional halts in vesicle transport.
Information Processing
[edit | edit source]Information processing is the brain's process of interpreting the receiving information and knowledge. There are three stages in information processing: sensory input, integration, and motor input. There are also different types of neurons in the information processing. They are sensory neurons, interneurons, motor neurons, and neurons coming out of the brain. Sensory neurons transmit information from sensors like the ears that detect stimuli like sound. Interneurons are neurons that make up most of the neurons in the brain. Motor neurons are the ones transmit signals to muscle cells so that they can contract. Lastly, neurons that come out of the brain are nerves that instigate the reaction or motor output. In addition, there are two main nervous systems that help to interpret the information. They are the central and peripheral nervous systems. Central nervous system consist of the brain and the nerve cord where the neurons that are in charge of integration are here. And the peripheral nervous system consists of neurons that receive sensory input and result in the motor output. [edited:"There are three stages in information processing: sensory input, integration, and motor input." - correct is "and motor -output-"]
Chemical Signaling
[edit | edit source]Chemical signaling is the physical chemical interchange that takes place in the synaptic cleft. Vesicles containing neurotransmitters are released by an incoming axon and received by receptors on opposing ends to induce a response on the recipient neuron. Chemical signaling via molecules secreted from the cells and moving through the extracellular space. Signaling molecules may also remain on cell surfaces, influencing other cells. Chemical signaling can involve small molecules (ligands) or large molecules (cell-surface signaling proteins). This signaling can be received either on the surface of cells by receptor proteins or within the interior of cells but also by receptor proteins. An example can be within-cell reception of signals is of steroid hormones. Signals also can be intentionally provided, such as is the case of hormones, or instead can be present for reasons that are not specifically for the purpose of providing a signal. The example can be carbon dioxide levels in blood.

Glial Cells
[edit | edit source]Glial cells are not neurons. They significantly outnumber neurons and are therefore vital to the role of the nervous system. It was previously thought that Glial cells merely aided with physical support within the nervous system. However, Glial cells actually electrically communicate with neurons and provide important biochemical support to them. Common types of Glial cells include 1. oligodendrocytes 2. astrocytes 3. microglia and 4. ependymal cells, all of which are found in the Central Nervous System. Glial cells found within the Peripheral Nervous System include 5. Schwann Cells and 6. satellite glial cells.
1. Oligodendrocytes
- Oligodendrocytes are one of the types of the neuroglial cells that is mainly responsible for myelinating central axons in central nervous system. Myelination refers to the act of oligodendrocytes wrapping around the axon with myelin sheath that is made of lipid and protein. Myelination of the oligodendrocytes have crucial effects on the transmission of neural signals by increasing the speed at which action potentials are conducted along axons. This action allows the neural signal to travel long distance with short energy and time.

2. Astrocytes
- Astrocytes constitute 20-50% of the volume in most brain areas, especially in the central nervous system. It is mainly responsible for the physical and metabolic support of the brain. It has many other functions including generating numerous proteins such as N-CAM, laminin, fibronectin, growth factors as well as cytokines, which is responsible for signaling proteins involved in the immune system.
3. Microglia
- Microglia is one type of the neuroglial cells that is mainly responsible for acting as macrophages. Microglia takes up about 5-20% of the mammal brains that act as mediators of immune response. Microglia cells constantly move around within the central nervous systems analyzing for damaged neurons, plaques, and infectious agents.
4. Ependymal cells
- Epedymal cells aid in separating the fluid components of the Central Nervous system by creating an epithial layer and are also a source of neural stem cells. Epedymal cells in the ventricular system of the brain form capillaries that form chroid plexus in each ventricle of the each hemisphere of the brain. Chroid Plexus then produces cerebrospinal fluid (CSF). 60-80% of CSF comes from chroid plexus and rest from extrachrodial sources.
5.Schwann cells
- Schwann cells' functions are very similar to oligodendrocytes. They myelinate neurons within the peripheral nervous systems instead of the neurons in the central nervous system. The main difference is that schwann cells are about 100 micrometres long that only covers the portion of the axons individually whereas one oligodendrocytes can mylinate multiple axons by stretching out their dendrites.
6. Satellite glial cells
- Satellite glial cells are a type of glial cells that cover the exterior side of neurons in the peripheral nervous system. Satellite glial cells' functions are similar to astrocytes in the central nervous system. Although there is still ongoing research to discover the specific functions and mechanisms, it is so far discovered that the satellite glial cells supply nutrients to the peripheral neurons as well as regulating neurotransmitter by uptaking and inactivating the neurotransmitters.
Cerebrospinal fluid (CSF)
[edit | edit source]Cerebrospinal fluid(CSF) is bodily fluid that circulates around the nervous system and throughout body that is produced in choroid plexus of each ventricle system of the brain. It is commonly used for diagnostic information about the normal and pathological states of the nervous system.
Functions of CSF:
1. It provides buoyancy and support to brain and chord that protects against rapid movements and trauma.
2. The fluid delivers nutrition for both neurons in both CNS and PNS and for other glial cells.
3. It functions like lymphatic system and removes wastes out from the nervous system.
4. It controls homeostasis of the ionic composition of the local microenviroment of the cells of the nervous system.
5. It acts as a transport system for releasing factors, hormones, neurotransmitters, metabolites.
6. The fluid controls H+ and CO2 concentrations (pH levels) in the CSF that may affect both pulmonary ventilation and cerebral blood flow.
7. The fluid is essential in medical fields in which it provides diagnostic information about the nervous system.
References
[edit | edit source]Gorazd B. Stokin and Lawrence S.B. Goldstein, "Axonal Transport and Alzheimer's Disease". Annual Review of Biochemistry Vol. 75: 607-627 (Volume publication date July 2006) Print
Silverthorn, D. (2012) Human Physiology: An Integrated Approach, 6th edition. Prentice Hall.
Purves, Dale, Principles of Cognitive Neuroscience, Sinauer Associates, Inc., 2008.
http://commons.wikimedia.org/wiki/File:Astrocytes.jpg
Dubuc, Bruno. <http://thebrain.mcgill.ca/flash/a/a_01/a_01_cl/a_01_cl_ana/a_01_cl_ana.html>
- ↑ Campbell, Neil and Reece, Jane. (2007). Biology 8th Edition.Benjamin-Cummings Publishing Co. ISBN 978-0321543257
The action potential is the brief electrical impulse that is responsible for the propagation of information down the axon of a neuron. Some important concepts involved with the action potential are the membrane potential, resting potential, threshold. The membrane potential is the difference in electrical potential inside and outside the cell. The resting potential is the membrane potential of a neuron when the neuron is at rest and not receiving any excitatory or inhibitory signals. In many neurons, the resting potential is approximately –70 mV. The threshold is the membrane potential that has to be reached to produce an action potential. An action potential is an all or nothing phenomenon. It either reaches the threshold and produces an action potential or it doesn’t.
Process
[edit | edit source]
The first step in the action potential is for the neuron’s membrane potential to reach the threshold. This change in the membrane potential can be caused by a variety of factors including excitatory stimulation by another cell or neuron. An increase in the membrane potential is also called a depolarization. After the threshold is reached, voltage-gated sodium channels open, and sodium ions enter the cell, increasing the resting potential. As the voltage continues to increase, the voltage-gated potassium channels open as well, and potassium ions leave the cell. However, the membrane potential continues to increase as the sodium ions are still entering the cell. When the membrane potential reaches its peak, the action potential is attained, and the sodium channels become refractory and no more sodium ions enter the cell. Potassium ions continue to leave the cell, and the membrane potential slowly decreases towards the resting potential. This decrease in the resting potential is called a hyperpolarization. At the resting potential, the potassium channels close and the sodium channels “reset” so that they can open again if the threshold is reached. Since all the potassium channels don’t close as soon as the resting potential is reached, extra potassium ions leave the cell, decreasing the membrane potential past the resting potential. The membrane actually undergoes an afterhyperpolarization, a drop in the membrane potential past the resting potential. Eventually, the membrane potential returns to the resting potential as the potassium ions diffuse back into the cell as the cell membrane is quite permeable to potassium ions. Sodium-potassium transporters pump sodium ions out of the cell and pump potassium ions into the cell.
Mechanism (Refer to diagram)
The resting potential is set by the sodium potassium pump which pumps 3 Na+ and 2K+ in using the energy of 1 ATP. This energy is required because the ions are being pumped against their concentration gradient. Resting potential lies between -60 to -80 mV. At resting potential, no signals are being sent across the membrane. Depolarization is the second step in this mechanism and occurs when the cell becomes less negative because Na+ voltage gated channels have opened and Na+ ions are now entering the cell causing a reduction in the magnitude of the membrane potential. Once the membrane potential reaches -55 mV, it has reached threshold which will trigger an action potential. The third step is called the rising phase where more Na+ voltage gated channels open which causes the cell to become less negative. It brings a positive charge to the cell which further enhances depolarization. More sodium channels open, resulting in a positive feedback loop. The following step is called the falling phase because eventually, Na+ voltage gated channels get inactivated and block additional sodium from entering. Mean while, the K+ voltage gated channels start to open and hyperpolarization takes place. Hyperpolarization causes an increase in the magnitude of the membrane potential resulting in making the cell more negatively charged. The last part of this mechanism is called the undershoot because the Na+ voltage gated channels close completely while K+ channels also begin to slowly close. And this returns the membrane potential back to rest just as in the first step of the mechanism.
Na+/K+ Pump Mechanism The ion pumps, with bound to ATP, binds 3 intracellular Na+ ions. The ATP is hydrolyzed, leading to phosphorylation of the pump and subsequent release of ADP. There is a conformational change in the pump takes place exposing the Na+ ions to the outside. The phosphorylated form of the pump has a low affinity for Na+ ions, so they are released. The pump then binds 2 extracellular K+ ions, which causes the dephosphorylation of the pump, reverting it to its previous conformational state, transporting the K+ ions into the cell. The dephosphorylated form of the pump has a higher affinity for Na+ ions than K+ ions, so the two bound K+ ions are released. ATP binds, and the process starts again.
Conduction of Action Potential
As an action potential travels down an axon, it is regenerating the depolarization. The action potential is transferred to neighboring regions, conducting action potential throughout the entire axon. While an action potential is being conducted in a particular region of the axon and is undergoing falling phase, the area behind it is hyperpolarizing and undergoing the falling phase. This area is known as the repolarized zone caused by the outflow of potassium ions. As a result, the inactivated Na+ channels behind the area of depolarization prevent any action potentials from traveling backwards, making them uni-directional.
Properties
1. Action potential is always depolarizing.
2. Action potential's amplitude is independent of stimulus pathway.
3. Action potential has all or none response. If the frequency reaches or passes the threshold, action potential occurs. If it falls under the threshold, no action potential occurs.
4. Amplitude does not decay with distance. It is indeed transferred equally, without any loss, across the axon.
5. There is an absolute refractory period and a relative refractory period.
6. At rest channels for sodium and calcium are closed
7. Membrane is selectively permeable to potassium the most
8. Osmotic pressure is opposed by electrostatic pressure to reach equilibrium for balancing the concentration ions.
Effects of Axon Structure
[edit | edit source]The diameter of an axon influences the speed of an action potential. The larger the width of an axon, the less resistance it provides to the current of an action potential. It acts analogous to how a water hose would, the wider the diameter of a hose offers less resistance to the flow of water. As a result, action potentials are conducted much faster. Another factor that contributes to the speed of action potential are myelin sheaths. Myelin sheaths are made by two types of glia, Schwann Cells in the peripheral nervous system and oligodendrocytes in the central nervous system. Myelin acts as an insulator for the axon and increases the length over which an action potential can be effective. This form of insulation allows the depolarizing current of an action potential to reach threshold sooner. The only problem with insulation is that axons now no longer have access to the environment or extracellular space. The only areas on an insulated axon that do allow for interaction are gaps in the myelin sheaths called Nodes of Ranvier. Action potentials are only formed at these nodes because this is the area where the exposed Na+ voltage gated channels are located. One node will undergo the rising phase of an action potential and the current produced will travel immediately to the next node where the membrane will then be depolarized and action potential will be regenerated. The process in which the action potentials and depolarization jump from node to node is called saltatory conduction.
Reference
[edit | edit source]Carlson, Neil R. Physiology of Behavior. Boston: Pearson Education, Inc., 2007.
Reece, Jane B. Campbell Biology, 2011
Levinthal, Charles, "Drugs, Behavior, and Modern Society", Pearson Education, Inc., 2008

Membrane Potential
[edit | edit source]Membrane potential is the voltage across the plasma membrane. The resting membrane potential is between -60 mV and -80 mV. There are steps in forming the resting membrane potential. First, the known concentration of K+ is 140 mM (millimolar) inside the cell and 5 mM outside of the cell and the Na+ concentration is 150 mM outside of the cell and 15 mM inside the cell. Cl- concentration is 120 mM outside the neuron and 10 mM inside the cell. Such concentrations of chlorine, potassium, and sodium always remain constant because there is sodium potassium in the plasma membrane of the neurons. Also, ATP is used to transport ions against their concentration gradients to maintain the constant concentrations. For instance, ATP helps sodium to go out of the cell and potassium enter the cell. Since the exchange of ions occur at the concentration gradient, it is important to understand that the concentration of gradients of sodium and potassium across plasma membrane represent a chemical type of potential energy. In addition, ion channels in the selective permeability contribute to the electric potential of neurons. For example, there are K+ channels and Na+ channels. There are also ions that move through the membrane through these ion channels to generate a potential across the membrane. Consequently, the diffusion of K+ through the potassium channels is very important for forming the resting potential because the outward flow of potassium results in the negative membrane potential of between -60 mV and -80mV. Excess negative charge inside the cell exerts an attractive force that stops the add’l flow of potassium ions outside of the cell. As a result, there is the creation of equilibrium potential which is the membrane voltage for a moving ion when it is at equilibrium. Nernst created an equation to calculate the resting potential of an ion:
E(ion)= 62mV(log [ion]outside/[ion]inside)
The resting potential for potassium is -90 mV. And the resting potential for sodium is 62 mV. [1]

Signaling allow neurons to communicate with each other. This usually involves long and complex signaling pathways, which results in an action being taken. Examples of different intracellular signaling include paracrine signaling, endocrine signaling, and both chemical and electrical synapses. In paracrine signaling, chemicals are secreted onto local target cells. In contrast, endocrine signaling secretes the chemicals or hormones directly into the bloodstream, which then delivers the chemicals to their intended targets.
A general purpose of chemical signal transduction is to allow for the amplification of a signal. A single ligand can binding to a receptor, which then releases proteins to bind to other signaling molecules and receptors, can exponentially increase the amount of molecules trying to reach a target protein, thereby greatly increasing the binding of the molecules to the targets and the potency of the signal.
Signaling molecules are grouped into three classes: cell-impermeant molecules, cell-permeant molecules, and cell-associated signaling molecules.
Cell-impermeant molecules are molecules that are unable to cross the lipid bilayer plasma membrane and must therefore bind to extracellular receptors. Neurotransmitters are considered cell-impermeant molecules.
Cell-permeant molecules are molecules that are relatively insoluble and are able to cross the lipid bilayer plasma membrane to bind to intercellular receptors. Steroids are considered cell-permeant molecules.
Cell-associated signaling molecules are able to only bind to receptors that are in direct contact with the target.
Receptors
[edit | edit source]There are several types of receptors that receive these different signaling molecules. The binding of a molecule to a receptor will initiate a conformational change within the receptor, which allows signaling to occur.
Channel-linked Receptors
[edit | edit source]
These receptors are also called ligand-gated ion channels. These receptors function by opening or closing their channels when a signal binds to the receptor site. The opening of the channel allows ions to flow across the membrane which leads to an ion gradient across the membrane.
Enzyme-linked Receptors
[edit | edit source]Enzyme-linked receptors are comprised primarily of protein kinases, which phosphorylate target proteins inside the cell. A signal first binds to an inactive enzyme. This activates the enzyme, which allows a product to be made.
Intracellular Receptors
[edit | edit source]Intracellular receptors are usually activated by cell-permeant or other molecules that can pass through the membrane. When the signalling molecule transverses the lipid bilayer plasma membrane and binds to the receptor, the inhibitory complex dissociates and becomes the activated form of the receptor. This begins a signaling cascade that regulates the transcription of DNA.
G-Protein-Coupled Receptors
[edit | edit source]
G-Protein-Coupled receptors (GPCRs) are activated when a signal molecule binds to the receptor, which then binds a G-protein, thereby activating it. There are two types of G-proteins: heterotrimeric G-proteins and monomeric G-proteins.
Heterotrimeric G-proteins
[edit | edit source]Heterotrimeric G-proteins contain three subunits: α, β, and γ. These three subunits are normally inactive when binded together. When a signalling molecule binds to the receptor, phosphorylation occurs and the GDP turns into GTP, which then allows the α subunit to dissociate, thus activating the G-protein. The α subunit then binds to an effector protein and allows for difference responses and mechanisms throughout the cell.
Monomeric G-proteins
[edit | edit source]Monomeric G-proteins are also known as small G-proteins. They use a similar mechanism to heterotrimeric G-proteins. However, instead of the three subunits, a G-protein called ras (named for its discovery in rat sarcoma tumors) is used. The binding of a signaling molecule to the receptor phosphorylates GDP to GTP and actives ras, allowing it to transmit a signal to its target proteins.
Second Messengers
[edit | edit source]Second messengers are used as signaling molecules between neurons.
Calcium
[edit | edit source]The calcium ion (Ca2+) is one of the most abundant second messengers seen in neurons. The influx of Ca2+ into the cell depolarizes the cell so many mechanisms may occur.
Sources
[edit | edit source]There are four methods of increasing Ca2+ in the interior of the cell:
- Voltage-gated calcium channels open in response to the initial depolarization, which allows a further Ca2+ influx.
- Ligand-gated calcium channels open in response to ligand attachments.
- Ryanodine receptors, which are bound to the endoplasmic reticulum are activated in response to the rise in intracellular levels of Ca2+. This causes the channels to open and an efflux of Ca2+ from the endoplasmic reticulum into the interior of the cell.
- Inositol trisphosphate (IP3) receptors, also bound to the endoplasmic reticulum, are activated by IP3 binding to its receptor. Similarly to the ryanodine receptor, this causes to the channels to open and an efflux of Ca2+ from the endoplasmic reticulum to flow into the cell.
Removal
[edit | edit source]There are several ways to remove Ca2+ from the cell:
- Na+/Ca2+ exchanger on the plasma membrane exchanges inflowing Na+ for outflowing Ca2+.
- Membrane Ca2+ pumps use ATP for active transport to transport Ca2+ out of the cell.
- Ca2+ binding proteins bind to Ca2+ and remove their activating abilities.
- Intracellular Ca2+ pumps on the endoplasmic reticulum use ATP to pump Ca2+ back into the endoplasmic reticulum.
- Mitochondria also remove calcium from the cell.
Intracellular Targets
[edit | edit source]These are some examples of targets will initiate some sort of response when bound and activated by Ca2+.
- Calmodulin will bind to other targets. It is one of the main initiator targets of downstream signaling cascades.
- Protein kinases add phosphate groups to proteins (phosphorylation).
- Protein phosphatases remove phosphate groups from proteins (dephosphorylation).
- Ion channels
- Synaptoptagmin is an essential protein involved in trafficking synaptic vesicles, containing neurotransmitters, to the surface of the presynaptic terminal for release.
Cyclic AMP and Cyclic GMP
[edit | edit source]Cyclic adenosine monophosphate (cAMP) is produced when adenylyl cyclase, which is activated by G-proteins in the plasma membrane, acts on ATP to remove two phosphate groups. The most common target of cAMP is the cAMP-dependent protein kinase (PKA) which is often triggers many mechanisms and responses. Cyclic guanosine monophosphate (cGMP) is produced in a very similar process as cAMP, where guanylyl cyclase acts on GTP to remove two phosphate groups. Also similarly to cAMP, the most common target for cGMP is the cGMP-dependent protein kinase (PKG) which also serves a similar function as PKA.
IP3 and Diacylglycerol (DAG)
[edit | edit source]In both the cases of IP3 and DAG, phosphatidylindositol bisphosphate (PIP2) is cleaved by an enzyme called phospholipase C, which is activated by calcium ions and G-proteins. The result of this cleaving produces IP3 and DAG. DAG goes on to target protein kinase C (PKC), within the cell, which causes phosphorylation to occur in its targets, triggering a signaling cascade. IP3 binds to the IP3 receptors, which then allow a calcium efflux from the endoplasmic reticulum.
Modulation
[edit | edit source]- modulation refers to the synaptic transmission that modifies effectiveness of EPSPs generated by other synapses with trasmitter gated ion channels such as activating NE beta receptor. -The mechanisms are: 1. NE, the neurotrasmitter, binds to the corresponding receptors on posynaptic neurons, which activates G-protein in membrane 2. The G-Protein then activates the enzyme, adenylyl cyclase 3. Adenylyl cyclase then converts ATP into second messenger, cAMP. 4. cAMP activates a protein kinase that causes a potassium channel to close by attaching a phosphate group to it.
Dendritic Information Processing
[edit | edit source]- A Cell with an axon can have local ouputs through its dendrites (back propagation) - Dendrites can carry out complex computations with mostly passive properties - Distal dendrites can be closely linked to axonal output (ie. large diameter apical dendrite) - Many neurons show a separation of their dendritic fields
Population Coding
[edit | edit source]- population coding hypothesis states that information within brain is carried by pools of neurons not by a single neuron system
1. The Independent-Coding Hypothesis - each neuron contributes to the pool independently - the "vote" of each neuron gives a population vector
2. The Coordinated-Coding Hypothesis - the relationships among the neurons in a population is an important part of the signal - the signal cannot be decoded without considering spike synchrony, oscillations, or some other relationship among the neurons in the population.
-
Characteristic Patterns of Activity
References
[edit | edit source]Purves, Dale, et al. Neuroscience, 4th Edition. Sunderland, MA: Sinauer Associates, Inc., 2008. Purves, Dale, "Principles of Cognitive Neuroscience", Sinauer Associates, Inc., 2008 Glial Cells (Neuroglial Cells) provide the support, stability, and insulation for neurons. These cells have no role in relaying information but are crucial for the survival of neurons. Glial cells are most abundant in the nervous system with a glial to neuron ratio of approximately 3:1. They are generally smaller in size and do not consist of axons or dendrites. Three types of glial cells exist in a developed central nervous system – astrocytes, oligodendrocytes, and microglial cells.
Astrocytes
[edit | edit source]Astrocytes are star-shaped glial cells that exist only in the central nervous system. These glial cells are used to maintain the best chemical environment for neurons. Astrocytes also help regulate signaling of the neurons by breakdowns of neurotransmitters.
Oligodendrocytes
[edit | edit source]Oligodendrocytes act as myelin. This helps with neuro signaling because it speeds up the process. Signals travel faster down myelinated axons. Again, oligodendrocytes exist in the central nervous system.
Microglial
[edit | edit source]Microglial cells remove cellular waste products from areas of injury or cell death. It helps remove unwanted debris that takes up unnecessary space. These cells are often known as macrophages because of their functional similarities. During head injuries, microglial cells increase in number to help remove the dead cells.
Effects of Glial Cells on Depression
[edit | edit source]After careful analysis of depressed patients’ brain (post mortem), it was discovered that these patients had fewer amounts of glial cells. This caused imbalances in the chemical environment and restricts proper neuro communications. Elsayed and colleagues at Yale University discovered that FGF2 could help produce more glial cells in the brain and reduce the effects of chronic stress. FGF2 (fibroblast growth factor-2 ) was injected in mice that had similar depression symptoms (sadness and loss of interest) and the results showed an increase in glial cell production. This is a newfound discovery for the treatment of depression.
References
[edit | edit source]Elsayed M; Banasr Mounira; Duric V; Fournier NM; Licznerski P.; Duman RS Antidepressant Effects of Fibroblast Growth Factor-2 in Behavioral and Cellular Models of Depression in Biological Psychiatry. Elsevier. 2012, 73. Purves, Dale. "Simple NCBI Directory." Neuroglial Cells. U.S. National Library of Medicine, 18 Jan. 0000. Web. 21 Nov. 2012. <http://www.ncbi.nlm.nih.gov/books/NBK10869/>.
Definition
[edit | edit source]Neurochemistry is a science that describes the study of neurochemicals, which include neurotransmitters and other molecules (like neuro-active drugs) that influence neuron function. It can be said that neurochemistry is the biochemistry of the nervous system.
Reference
[edit | edit source]“Neurochemistry." Wikipedia, The Free Encyclopedia. Wikimedia Foundation, Inc. 31 July 2010. Web. 28 November 2010.
Neurotransmitters
[edit | edit source]A neurotransmitter is a chemical that is released by a terminal button and has an excitatory or inhibitory effect on another neuron. There are many types of neurotransmitters. There are currently over 100 known agents that serve as neurotransmitters.
A brain circuit is a neurotransmitter current or neural pathway in the brain.
By mimicking a neurotransmitter's effect, agonist will enhance and increase the activity of that neurotransmitter while, on the other hand, antagonist will decreases or blocks the effects of a neurotransmitter.
Another chemical substance calls inverse agonist is responsible for producing effects opposite to those of a particular neurotransmitter.
Neurotransmitters Activities:
Neurotransmitters are stored in tiny sacs at the end of the neuron and are released into the synapse as the sacs merge with the outer membrane by an electric jolt. The neurotransmitters travel across the gap to bind with receptors. The neurotransmitters are released from the receptors after the messages has been successfully absorbed by the adjacent neuron. After that, those chemical substances are either degraded or reabsorbed back to the neuron where they come from.
What makes neurotransmitter different from other chemical signaling system?
[edit | edit source]There are many other signaling system of chemicals such as hormones, neurohormone, and paracrine signaling. However neurotrasmitters have advantages in having a greater degree of amplification and control of the signal. It also lengthens the time of cellular integration from milliseconds to minutes and even hours. While hormones are mainly synthesized in gland, neurotransmitters are synthesized and released from neurons. Neurotrasmitters are, as far as we know, only released in response to an electrical signal. There are many mechanisms that must exist to terminate the action of the neurotrasmitters such as chemical deactivation, recapture (endocytosis), glial uptake and diffusion.
Exocytosis of Neurotransmitters Release
[edit | edit source]Exocytosis is a process by which vesicles release their contents. Between presynaptic and postsynaptic neurons, certain neurotransmitters are carried out within vesicles from presynaptic neurons are released to the synaptic cleft. In order to do so, first there is an influx of Ca++ ions takes place through voltage gated calcium channels in the presynaptic site. The calcium ions intering the cells affect the movements of the vesicles toward the active sites by dissolving some actin filaments. It also helps the fusion of the vesicles with the plasma membrane in the presynaptic side. As the vesicles fuse, the neurotranmitters are released into the synaptic cleft, then which binds to the corresponding receptors on the postsynaptic neuron.
-
Caption1
Categories of Neurotransmitters
[edit | edit source]Neurotransmitters are separated into two very broad categories based on their size: neuropeptides and small-molecule neurotransmitters.
Neuropeptides
[edit | edit source]Neuropeptides are generally large molecules that range from 3 to 30 amino acids and consists of over 100 peptides. They are grouped into five categories: brain/gut peptides, opioid peptides, pituitary peptides, hypothalamic releasing hormones, and everything else. Neuropeptides are genetically coded, synthesized from mRNA as prohormones. They are mostly colocalized with and modulate effects of other neurotrasmitters rather than directly presenting the effects. There is no uptake of the NTs, but rather they are broken down by enzymes.
Neuropeptides include large molecules of opioids that include:
1. Beta-Endorphin - made from proopiomelanocortin - produced in pituitary gland, hypothalamus, brain stem
2. Met and Leu Enkephalin - made from proenkephalin - produced throughout brain and spinal cord
3. Dynorphin - ade from prodynorphin - produced throughout brain and spinal cord
Small-Molecule Neurotransmitters
[edit | edit source]Small-molecule neurotransmitters are much smaller than neuropeptides and may consist of a single amino acid or other molecule. The biogenic amines are a group within the small-molecule transmitters that consists of the catecholamines (dopamine, norepinephrine, epinephrine), serotonin, and histamine.
Acetylcholine
[edit | edit source]
Definition and function
[edit | edit source]Acetylcholine is a neurotransmitter that plays a very significant role in the central and peripheral nervous systems. Acetylcholine plays a very big role in the movement of muscles in the peripheral nervous system. Acetylcholine also seems to be released in a variety of areas in the autonomic branch of the peripheral nervous system. In the central nervous system, acetylcholine plays a role in plasticity, arousal, reward, attention, and REM sleep.
Metabolism
[edit | edit source]The synthesis of acetylcholine takes place in the nerve terminals. This process requires acetyl coenzyme A (also called acetyl CoA) and choline. Acetyl CoA is synthesized from glucose during glycolysis, and choline is already present in plasma. The synthesis of acetylcholine further requires choline acetyltransferase. After the synthesis, the ACh is then loaded into synaptic vesicles and released from the presynaptic terminal to the postsynaptic cell. Acetylcholinesterase (AChE) responds to the released acetylcholine and hydrolyzes the molecules back into choline and acetyl CoA. The choline is then transported back to the presynaptic terminal and recycled to resynthesize new ACh. The cleanup of old acetylcholine is the job of acetylcholinesterase.

RCSB PDB Protein of the Month: Acetylcholinesterase
Drugs that affect release of acetylcholine
[edit | edit source]Two drugs that influence the release of acetylcholine are botulinum toxin and black widow spider venom. Botulinum toxin is produced by a bacteria called clostridium botulinum that grows in improperly canned food. Botulinum toxin inhibits the release of acetylcholine. Black widow spider venom is produced by the black widow spider and it increases the release of acetylcholine.
Receptors
[edit | edit source]There are two types of receptor sites that are sensitive to acetylcholine.
1. Muscarinic Receptors - It is named so because they are responsive to the drug muscarine. Muscarinic receptors are mostly located in the parasympathetic autonomic nervous system. If a drug is antimuscarnic, that means that it interferes with the role of acetylcholine in stimulating parasympathetic reations of the body. Examples are atropine and scopolamine. Atropine, when applied to the eyes, for examples, causes the pupils to dialte by inhibiting the parasympathetic tendency for the pupils to constrict.
2. Nicotinic Receptors. - They are responsive to nicotine that are found near the end points of motor neurons where skeletal muscles are innervated as well as thourghout the cerebral cortex. Some antinicotinic drugs such as the poison curare, affect these motor neurons so dramatically that the body can become paralyzed.
Glutamate
[edit | edit source]
Glutamate is a key component in normal brain function. It is believed that over half the synapses that occur in the brain release glutamate as a neurotransmitter. It was discovered in 1907 by Kikunae Ikeda and was identified as a neurotransmitter in 1970s by Peter Usherwood. It is an excitatory relative of GABA. An excessive amount of glutamate (usually come from brain damage or a stroke) is very toxic to neurons and may result in brain cell death. Example for disease of excessive glutamate production is ALS, a degenerative neuromuscular disease.
Characteristics of glutamate 1. Glutamate is a principal excitatory neurotransmitter that is biosynthesized as byproduct of glucose metabolism. 2. Excess of glutamate can be neurotoxic. 3. Glutamate has four receptor types:
a. NMDA receptor
- NMDA is an ionotropic receptor that detects simultaneous events.
- The receptor is gated by comnination of voltage and ligand channels. Glutamate plus glycine binding opens channel to Ca++ for influx
- The effect mediates learning and memory through long term potentiation that essentially deals with psychological addiction, behavioral sensitization, and drug craving.
b. AMPAa Receptor
c. Kainate
d. AMPAb
Metabolism
[edit | edit source]The synthesis of glutamate is done locally from precursors such as glutamine, which comes from glial cells. The glutamine is released into the presynaptic terminal and synthesized into glutamate using an enzyme called glutaminase. The newly synthesized glutamate is then transported from the presynaptic terminal and across the synaptic cleft in synaptic vesicles. After release from the vesicles, the glutamate is then transported to the glial cells and converted into glutamine. This process is called the glutamate-glutamine cycle.

GABA
[edit | edit source]GABA (or gamma-Aminobutyric acid) is used frequently in inhibitory synapses in the central nervous system. It is most commonly found in local circuit interneurons.
GABA is an inhibiting neurotransmitter which is best known for its ability to reduce anxiety by reducing postsynaptic activity. However, GABA's effect is not only to anxiety but has a broader influence. The GABA system is spread throughout the brain. Different types of GABA receptors seem to act in different ways which leads to the conclusion that GABA is not just one system working in only one manner but is composed of several subsystems.
Metabolism
[edit | edit source]GABA synthesis requires glucose, which metabolizes to glutamate. An enzyme called glutamic acid decarboxylase (GAD) then converts the glutamate into GABA. GAD requires a cofactor called pyridoxal phosphate to work properly. A deficiency in vitamin B6, in which pyridoxal phosphate is derived from, would prevent the synthesis of GABA from glutamate. After the GABA has been released and used, the GABA is then transported to glial cells via synaptic vesicles specifically for GABA, called GATs. There, the GABA is then converted into succinate.
Glycine
[edit | edit source]Glycine is a neutral amino acid that is also distributed within the central nervous system. It is synthesized from serine using an enzyme called serine hydroxymethyltransferase and then transported to be released from the pre-synaptic terminal in synaptic vesicles called GATs. After the release of glycine, plasma membrane transporters remove it from the synaptic cleft.
Biogenic Amines
[edit | edit source]

The biogenic amines are sometimes classified as a separate group from the small-molecule neurotransmitters. They regulate many functions of both the central and the peripheral nervous systems. Many psychiatric disorders occur because of defects in the synthesis or the pathways of the biogenic amines. There are five known biogenic amine transmitters: dopamine, norepinephrine, epinephrine (all together known as the catecholamines), histamine, and serotonin. The catecholamines are all synthesized from tyrosine.
Dopamine
[edit | edit source]Dopamine is most present in the corpus striatum, which plays a key role in the coordination of body movements. It is synthesized from tyrosine with the help of DOPA decarboxylase. It is then transported to the pre-synaptic terminals in synaptic vesicles called vesicular monoamine transporter (VMAT).
A defect in dopamine production is a cause for Parkinson's disease. It is also involved in the reward centers of the brain, and many drugs used for abuse target the dopamine synapses in the central nervous system.
Prostaglandins
[edit | edit source]A chemical messengers that work right within the cells where they are synthesized.
Structure:It is an unsaturated carboxylic acids derived from arachidonic acid. The numerous functional groups in prostaglandin contribute to its variety functions in the body; there are 2 alkenes group (one cis and one trans), 2 alcohols, a ketone, and one acid on a 20 carbon skeleton with a five-member ring.
Function: It stimulates inflammation process, responds to injury by producing pain or infection by producing fever. It forms blood clots when blood vessel is damaged. Specific prostaglandins are involved with the induction of labor and reproductive processes.
Norepinephrine
[edit | edit source]Norepinephrine (or noradrenaline) is used in the locus coeruleus in the brain. It is involved in behaviors related to sleeping, attention, and feeding. The synthesis of norepinephrine requires an enzyme called dopamine-β-hydroxylase to convert dopamine to norepinephrine.
Norepinephrine is a part of the endocrine system that seems to stimulate at least two group of receptors called alpha-adrenergic and beta-adrenergic receptors. In the central and peripheral nervsous system, several norepinephrine circuits have been identified which actively helps our body tp control heart rate, blood pressure, respiration. One of the norepinephrine circuit is associated with the emergency reactions or alarm responses. Thus, it may indirectly plays an important role in panic attacks and other disorders. Norepinephrine is concentrated in the hypothalamus and libic system but also found throughout the brain.
Epinephrine
[edit | edit source]Epinephrine (or adrenaline) is also found in the brain. It is the least abundant of the three catecholamines in the brain. Epinephrine is mostly located in the medulla, the hypothalamus, and the thalamus. Phenylethanolamine-N-methyltransferase catalyzes norepinephrine to convert it to epinephrine.
Histamine
[edit | edit source]Histamine is found in the hypothalamus and sends signals to the central nervous system. It is involved in arousal and attention as well as the vestibular system. Histamine is synthesized from the histidine
Serotonin
[edit | edit source]Serotonin (or 5-hydroxytryptamine) is found in the pons and upper brainstem. It is involved in the regulation of sleep and wakefulness. Serotonin is synthesized from tryptophan.
Serotonergic pathways are a topic of interest when it comes to studies of depression and anxiety. Drugs to treat these disorders often target these pathways.
Serotonin (5-hydroxytrytamine or 5-HT) is a neurotransmitter that is associated with the processing of information and coordination of movement, inhibition, restraint, assists the regulation of eating, sexual, and aggressive behaviors. There are at least 15 different serotonin receptors that play different function in our body. Several of drugs affect the serotonin system. For example, serotonin-specific inhibitors (SSRIs),which enhances serotonin's effects by preventing it from being absorbed, are used to treat particularly anxiety, mood, and eating disorder.
Serotonin is very important to psychopathology because it may involves in different psychological disorders. Low serotonin activity will lead to aggression, suicide, impulsive overeating, excessive sexual behavior. Moreover, Its interaction with dopamine is implicated in schizophrenia.
References
[edit | edit source]“Acetylcholine." Wikipedia, The Free Encyclopedia. Wikimedia Foundation, Inc. 15 November 2010. Web. 28 November 2010.
Carlson, Neil R. Physiology of Behavior. Boston: Pearson Education, Inc., 2007.
Purves, Dale, et al. Neuroscience, 4th Edition. Sunderland, MA: Sinauer Associates, Inc., 2008.
Durand, V. Mark, and David H. Barlow. Essentials Of Abnormal Psychology. 5th. 12. Belmont: Wadsworth Pub Co, 2009. Print.
Levinthal, Charles, "Drugs, Behavior, and Modern Society", Pearson Education, Inc., 2008
Boeree, George. "Neurotransmitters." General Psychology. N.p.. Web. 10 Nov 2012. <http://webspace.ship.edu/cgboer/genpsyneurotransmitters.html>.
"Prostaglandins." Virtue Chembook. Chemistry Department, Elmhurst College. Web. 10 Nov 2012. <http://www.elmhurst.edu/~chm/vchembook/555prostagland.html>.
Purves, Dale, "Principles of Cognitive Neuroscience", Sinauer Associates, Inc., 2008 Levinthal, Charles, "Drugs, Behavior, and Modern Society", Pearson Education, Inc., 2008
Axonal Transport
[edit | edit source]
Axonal transport, also called axoplasmic transport, is a cell process not only responsible to the movement of protein and membrane to its axon, but also responsible to the movement of molecules that destined for degradation from the axon back to the cell body. Movement toward the cell body is called retrograde transport; movement toward the synapse is called anterograde transport.
Axonal transport is essential to neuron cell growth and survival. Axon of neuron is 1,000 or 10,000 times the length of the cell body, but contains no ribosomes, which means that it is unable to produce protein. All proteins and membrane must synthesize proteins in neuronal cell body or neuron cell dendrite, and then transport back to the axon. The motor protein kinesin is a motor protein used during anterograde processes while dynein is used in retrograde processes .
Vesicular cargoes move 50-400 mm/day whereas proteins move less than 8 mm/day.
Alzheimer’s Disease
[edit | edit source]Introduction of Alzheimer’s Disease
[edit | edit source]
Alzheimer's disease, also called AD, is a common form of dementia. The reason caused disease is unknown and there is no cure for disease. The patient will get worse as disease progresses, and eventually leads to death.5% 65-year-old people have Alzheimer's disease; 20% 85-year-old people have Alzheimer's disease- percentage of patients’ increases as patients’ age increases.
Type
[edit | edit source]There are 3 kinds of Alzheimer's disease:
1- Dementia AD, also called DAD, occupies 10% of patients. Most patients are younger than 65; some of them even younger than 40-year-old. Researchers find out that some patients have number 14-chromosome abnormalty, which doesn’t show on SDAT patients.
2- Senile dementia with Alzheimer's type, also called SDAT, occupies 85~90% of patients. Most patients are older than 60 years old. Research shows that SDAT is not hereditary.
3- Familial AD, also called FAD, occupies 1% of patients. FAD is considered as hereditary disease, which is based on euchromosome heredity rules of Mendel. In the family of FAD, at least two generation of family members are diagnose as FAD. Most patients are younger than 40-year-old. The disease is barely affected by drugs.
Diagnose
[edit | edit source]Level MMSE Explanation Time to next level Degradation degree
1 29~30 Normal - Adult
2 29 Normal adult aging dysmnesia. - Adult
3 25 Mild neural cognitive dysfunction Reducing working ability and social ability. - Younger adult
4 20 Pre-dementia Alzheimer's disease Reducing ability of calculation and complicated work, attention and dysmnesia. 2 years 8~ Teenagers
5 14 Early Alzheimer's disease Reducing calculation on 2~20, losing ability of normal activity. 1.5 years 5~7 years old
6 5 Moderate Alzheimer's disease Cannot account number from 10 to 1, need help from other people to finish normal activity. 2.5 years 5~7 years old
7 0 Advanced Alzheimer's disease Need help for living, totally depends on other people. 8~9 years 4~15 months
Characteristics
[edit | edit source]There are four stages of the disease:
1- Pre-dementia:

Patients will have mild cognitive difficulties. The test can help doctor to find Alzheimer's disease symbols existing eight years before diagnosis of AD. These symbols, such as short memory loss and inability to acquire new information, will cause patients cannot finish complex living activity independently.
2- Early:
Defined as first 2~3 years, patients will have difficulties with language, executive functions, perception, or execution of movements. Language problem shows obviously in decreased word fluency and shrinking vocabulary, eventually leads to general oral and written impoverishment in language. Memory losing happens as same time, but less prominent than other symbols. Alzheimer's disease doesn’t affect all memory, such as implicit memory, episodic memory and semantic memory.
3- Moderate:
Disease will eventually hinder independence; patients will lose common living abilities. On language perspective, patients are unable to recall the vocabulary, leading incorrect word substitution. Reading and writing skills are also reducing progressively. Memory problem get worse; some patients even cannot recognize close relatives. Emotion changes dramatically, leading to wandering, crying and resistance to caregiving.
4- Advanced:
This is the last stage of Alzheimer's disease. The MMSE will drop 3~4 points every year until it goes to zero. Normally, patients can still live for 8~9 years, but most patients are already lost the living ability - all of their living activities are relied on other people. Their vocabularies reduce to simple phrases; sometimes they can only speak single word. Patients will dead eventually, but the reason caused patients dead is not Alzheimer's disease - is actually some external factors, such as infection of pressure ulcers or pneumonia.
Causation
[edit | edit source]There are several reasons that may cause Alzheimer’s Disease:

1- Heredity
Research report shows that more than 20% 80-year-old people have Alzheimer’s Disease. However, in the Alzheimer’s Disease family, more percentage of patients is observed; the age of having Alzheimer’s Disease even reduces to 13-year-old.
2- Cyanobacteria
Cyanobacteria contain neurotoxin named BMAA(β-N-methylamino-L-alanine). BMAA was verified that is strongly poisonous to animal nerve cells, increasing the speed of deterioration of animal brain cells. Accumulation of small amounts of BMAA can kill all brain nerve cells of rat.
3- Aluminum
Over-absorbing Aluminum ions is considered as the reason that may cause Alzheimer’s Disease. The research report shows that after the World War II, soldiers left on the Territory of Guam had high percentage of Alzheimer’s Disease. The high concentration of Aluminum ions was detected in the underground water of the Territory of Guam. After changing the drinking water for soldiers, the percentage of having Alzheimer’s Disease drops dramatically.
Treating and Drugs
[edit | edit source]There are three drugs that used to treat Alzheimer’s Disease. The first two drugs are already approved by FDA and used in clinical. The third one is still in testing period.
1- Anti-acetylcholinesterase
Anti-acetylcholiesterase is used to treat patients who have early Alzheimer’s Disease. The drug changes patients’ emotion and psychotic symptoms. In order to treat accompany diseases, such as insomnia, doctors suggest that patients should take sleeping pills or other assistant pills with Anti-acetylcholiesterase at same time.

2- Memantine(Namenda)
By stopping excitotoxicity of Glutamic acid, Memantine can stop the destruction of brain cells caused by Glutamic acid, which reduces the speed of losing living ability. Memantine is the only drug that can treat moderate and advanced Alzheimer’s Disease.
3- bFGF
Research report shows that bFGF is useful to treat rats that have Alzheimer’s Disease, but there is no clinical report supporting that bFGF has positive effect on human body.
Prevention
[edit | edit source]There is no evidence to support that any particular behavior is effective to preventing Alzheimer’s Disease. However, researches show that changing a few modifiable factors, such as diet, intellectual activities, and living speed, can reduce the chance of getting Alzheimer’s Disease. On diet, people should eat more vegetable and decrease times of eating meat. And intellectual activities, such as reading, playing board games, completing crossword puzzles, and playing musical instruments, help to keep brain cells healthy.
Reference
[edit | edit source]1. Gorazd B. Stokin and Lawrence S.B. Goldstein, "Axonal Transport and Alzheimer’s Disease"
2.http://en.wikipedia.org/wiki/Alzheimer’s_Disease
3.http://zh.wikipedia.org/zh-cn/阿兹海默病
4.http://baike.baidu.com/view/1340086.htm
5. Wei Fan "Introduction of Alzheimer’s Disease"
6. Meiting Luo "Medicines and Treating Methods of Alzheimer’s Disease"
7.http://en.wikipedia.org/wiki/Axonal_transport
8.http://baike.baidu.com/view/956775.htm
Overview
[edit | edit source]Multiple Sclerosis is a disease that results from demyelination and inflammation along axons. In some cases destruction of axons can also be seen. While the specific cause is unknown, there are several theories. One being that an antigen enters the body with structural similarity to myelin, and as the body launches an immune response, resulting in demyelination. The other hypothesis is that it is caused by a virus, which as the immune system tries to rid of, it consequentially damages myelin.
Symptoms of MS result from lesions to various nerve sites. Some examples being blindness (lesions of optic nerve), motor paralysis (due to lesions of the corticospinal tracts), abnormal somatic sensations (lesions of somatic sensory pathways.
Most of these symptoms are probably due to the fact that demyelination leads to compromises in action potential firing. More severe cases of MS have also shown the destruction of axons as a whole, leading to the functional deficits characteristic of MS.
Reference
[edit | edit source]Purves, Dave, et all. Neuroscience, Fourth Edition. Sunderland, MA: C. 2008, Sinauer Associates, Inc. Text.
Overview
[edit | edit source]Synaptic transmission occurs from a presynaptic terminal at the end of an axon to the postsynaptic specialization at the end of the dendrite on the desired cell, the two of which are separated by a synaptic cleft. The synaptic cleft contains extracellular proteins involved in the diffusion, binding and degradation of the molecule secreted by the axon. Signals are fired across the synapse then binding to a receptor on the dendrite. The specific receptor that binds the neurotransmitter determines what cation in the extracellular cleft will be allowed to enter the cell at that site. The binding of the neurotransmitter to the dendritic receptor then causes the action potential to fire down the axon, then sending a signal to the next cell.
Types of Synapses
[edit | edit source]There are two types of synapses where transmission can occur. The first type is an electrical synapse with gap junctions that allow electrical current to flow from one neuron to another. These synapses are responsible for quick and unchanging behavior. The second type are chemical synapses where presynaptic neurons release chemical neurotransmitters that carry information across a synaptic cleft. These synapses allow room for modification in the case that a change in behavioral response is necessary. Behavioral response can be changed by changing the type of receptor and enhancing or changing the number of synaptic vesicles. The majority of synapses are usually chemical synapses.
Process of Chemical Synapse
[edit | edit source]Inside a synapse, there are many synaptic vesicles located within the terminals of the neuron. These synaptic vesicles are membrane bounded compartments filled with neurotransmitters. They are usually sitting in a synapse waiting to be delivered or taken away. An action potential's journey ends at the synaptic terminal however when the moving depolarization hits the synapse, the Ca2+ voltage gated channels are able to sense the action potential and open. As a result, the Ca2+ ions move into the presynaptic membrane. This action causes the synaptic vesicle to migrate and dock at the bottom of the membrane. The high amount of Ca2+ concentration causes exocytosis where the vesicles fuse with the presynaptic membrane and neurotransmitters are then released into the synaptic cleft. Meanwhile at the post synaptic membrane, there are ligand gated ion channels that are selectively permeable and respond where there is binding of ligand which in this case are the neurotransmitters. The neurotransmitters bind to the ligand gated ion channels and change it's shape causing them to open.
There are also other possibilities for what may occur after neurotransmitters are released from the synaptic vesicle, this includes diffusing out of the synaptic cleft, being taken up by surrounding cells such as astrocytes or getting degraded by enzymes. In the case where neurotransmitters bind to a receptor that is not part of an ion channel such as the example discussed earlier, it undergoes indirect synaptic transmission. This is another form of signaling that can take place at the synapse. The neurotransmitter binding to the receptor activates a signal transduction pathway and while this process is slower, the effects are longer lasting.
Postsynaptic Potential
A postsynaptic potential is a change in the membrane potential generated by the binding of neurotransmitters to ligand gated ion channels in the postsynaptic cell. They are considered graded potentials that rely on the strength of the stimulus and do not regenerate. There are two types of postsynaptic potentials known as the excitatory postsynaptic potential (EPSP) and inhibitory postsynaptic potential (IPSP). EPSP is a depolarization that brings the membrane potential to threshold. On the other hand, IPSP is a hyperpolarization that brings the membrane potential further away from threshold. A single EPSP is too small to trigger an action potential in a postsynaptic neuron. However, when two EPSPS are produced at the same time or within close proximity form the same synapse on the same postsynaptic neuron, it results in a temporal summation. The second EPSP arrives before the depolarization of the first EPSP has a chance to dissipate. When two EPSPS are produced almost simultaneously from two different synapses on the same postsynaptic neuron, they are able to add together and result in a spatial summation. The combination of EPSPS that result from temporal and spatial summation are able to trigger an action potential. Under circumstances where an EPSP and IPSP happen at the same time, they will cancel out and no action potential will arise.
Generation of An EPSP (excitatory postsynaptic potential) -First, there is an impulse arriving in the presynaptic terminal causes the release of neurotransmitter (detailed mechanism is described in Neurotrasmitters page). - Then the neurotrasmitters bind to the trasmitter gated ion channels in the post synaptic membrane. - Na+ enters the post synaptic cell through thte open channels, the membrane will become depolorized (detailed mechanism is explained in the Action Potential page) - The resulting change in membrane potential (Vm) is the EPSP
Generation of an IPSP (inhibitory postsynaptic potential) -First, there is an impulse arriving in the presynaptic terminal causes the release of neurotransmitter (detailed mechanism is described in Neurotrasmitters page). - Then the neurotrasmitters bind to the trasmitter gated ion channels in the post synaptic membrane. - Cl- enters the post synaptic cell through thte open channels, the membrane will become hyperpoloerized (detailed mechanism is explained in the Action Potential page) - The resulting change in membrane potential (Vm) is the IPSP
-
Caption1
-
Caption2
Sources
[edit | edit source]Sources: Purves, Dave, et all. Neuroscience, Fourth Edition. Sunderland, MA: C. 2008, Sinauer Associates, Inc. Text. Purves, Dale, "Principles of Cognitive Neuroscience", Sinauer Associates, Inc., 2008
Overview
[edit | edit source]Channelopathies are genetic diseases due to alterations in ion channel genes. Examples:
Familial Hemiplegic migraines (FHM). These have been shown to occur due to mutations in the Ca²+ channel in the brain. Mutations in pore-forming region of this channel result in FHM characterized by uncontrolled muscle movement. This and other examples suggest a correlation between mutations in the gene for this ion channel and specific implications of FHM symptoms. While this relation is evidenced, the original cause of the migraine is unknown.
Episodic ataxia type 2, or EA2. These mutations cause Ca²+ channels to be cut off, leading to abnormal composition of the channel. Symptoms include uncontrolled muscle movement, vertigo, nausea and headaches.
CSNB (congenital stationary night blindness) is caused by similar mutations as EA2 and leads to abnormal retinal function. It results in numerous affects on vision, including night blindness and lack of acuity.
(GEFS) Epilepsy results from a defect in the Na+ channel. These mutations slow the channel activation, which may explain the hyperexcitability phenoma characteristic of epilepsy occurring here.
(BFNC) seizures occur from mutations in the gene for the K+ channel, leading to hyperexcitability that causes abrupt seizures in the first few weeks-months of life.
Reference
[edit | edit source]Purves, Dave, et all. Neuroscience, Fourth Edition. Sunderland, MA: C. 2008, Sinauer Associates, Inc. Text.
Phases of Brain Development
[edit | edit source]Phases of early embryonic brain development mainly include seven stages such as neural induction, proliferation, migration, differentiation, synaptogenesis, cell death/stabilization and synaptic rearrangement. Neural induction stage takes from the day 18 to 24 since the fertilization. Proliferation stages takes from the day 24 to day 125. Migration stage takes from the day 40 to day 160. And the differentiation stage takes from the day 125 to the postnatal periods. from the differentiation period to the postnatal period, the brain development becomes very sensitive to the environmental factors play a role in addition to the genetic factors.
Neural Induction
[edit | edit source]Neural Induction refers to the process that induces a region of embryonic ectoderm to form the neural plate on the dorsal surface of the embryo. Neural induction goes through several stages such as neurulation and neural patterning.
During the early stages of the embryonic development, newly formed neural plate forms the neural groove, which then forms the neural tube that eventually develops in to the central nervous system. There are numerous enzymes, hormones and proteins that initiates this formation of neural tissue. But the main protein that contributes the most is the Spemmann's organizer. The ectoderm cells are naturally predisposed to become neural tissues. But the ectoderm cells produces BMPS (TGF-β family protein), which causes it to become epidermal (nonneural). However once the Spemann's Organizer molecules produces neural inducers such as follistatin, noggin, and chordin to block the effects of BMPs, ectoderm can be induced to become the neural tissue. Missing such inducers cause many birth defects during the early pregnancy such as spinal bifida and anencephaly. *BMPs are bone morphogenic protens that are members of polypeptide growth factor and transforming growth factor familes.
-
Caption1
-
Caption2
Reference
[edit | edit source]Purves, Dale, "Principles of Cognitive Neuroscience", Sinauer Associates, Inc., 2008
Proliferation
[edit | edit source]In the ventricular zone of the late neural induction stage, the wall of the emerging brain is only one cell deep. During the proliferation period, cells begin to extend their processes across the ventricular zone through the cell division cycle: G1-S-G2_M
-
Caption1
-
Caption2
Migration
[edit | edit source]During the migration stage cells that are yet to become neurons begin to move up along the radial glia cells that provide scaffolding system for the cell migration. Cells along the radial glia cells generally move up vertically forming layered stuctures with predominantly radial migration patterns, including the cerebral cortex, the hippocampal formation, colliculi, and the cerebellar cortex. There are three groups of molecules that are crucial for migration:
1. Adhesion molecules
- adhesion molecules such as Reln, Astrotactin, Neuregulin, Integrins alpha 3,6, play a role in attaching the cells to the radial glia cell.
2. Microtubule binding proteins
- regulate stability of microtubles
3. Actin binding Molecules
- regulate protein to protein interactions. Examples are filamin1, Cdk5/p35
-
Caption1
Differentiation
[edit | edit source]- During the differentiation stage the cells acquire an identity (phenotype). Each cell's axon thus begins to grow toward the target and acquires/expresses specific enzymes to produce neurotransmitters and receptors. And the most importantly, myelination of the neurons begin during this stage. There are numerous ways that axons find its target: 1. pathway selection 2. target selection, address selection 3.direct cell to cell communication 4. detection of extracellular signals from other cells 5.communication via diffusable chemical signals from cells at a distance.
Overview
[edit | edit source]Synaptogenesis is the formation of synapses between neurons and the final step in the development of the central nervous system (CNS). This occurs through two steps. The first step is the contact of an axonal growth cone, an extension of a developing axon, to its partner cell. The second step are the axonal and dendritic protein components coming to the site of contact and ultimately forming a functional synapse.
Neuromuscular Junction
[edit | edit source]NMJ forms in a series of steps that involve the exchange of signals among its three cellular components nerve terminal, muscle fiber, and Schwann cell. The neuromuscular junction (NMJ) is the most well-characterized synapse in that it provides a simple and accessible structure that allows for easy manipulation and observation. The synapse itself is composed of three cells: the Motor neuron, the myofiber, and the Schwann cell. In a normally functioning synapse, when a signal causes the motoneuron to depolarize, the motoneuron releases the neurotransmitter acetylcholine (ACh). Acetylcholine travels across the synaptic cleft where it reaches acetylcholine receptors (AChR) on the plasma membrane of the myofiber, the Sarcolemma. As the AChRs open ion channels, the membrane depolarizes, causing muscle contraction. The entire synapse is sheathed within a myelin cover provided by the Schwann cell to insulate and encapsulate the junction. The NMJ is functional at birth but undergoes numerous alterations postnatally. One step in maturation is the elimination of excess inputs, a competitive process in which the muscle is an intermediary. Once elimination is complete, the NMJ is maintained stably in a dynamic equilibrium that can be perturbed to initiate remodeling.
Factors that Regulate Synaptogenesis
[edit | edit source]In recent studies, TGF-β and Glia are directly related in the regulation of synaptogenesis. For example, the absence of TGF-β ligands or glia results in fewer synapses with other neurons. This was determined when the TGF-β ligands were removed from the NMJ(neuromuscular Junction) of Drosophila melanogaster and the resulting data showed a dramatic decrease in the number of synapses. The removal of glia cells decreased the amount of TGF-β activation therefore, the number of synapses decreased. These are the few factors that effect synaptogenesis.
References
[edit | edit source]Brose, Nils. "Synaptogenesis, Neuroligins, and Autism." Max-Planck-Institut fur Experimentelle Medizin. N.p., 2006. Web. 16 Nov. 2012. <http://www.em.mpg.de/index.php?id=106&L=1>.
"Development of the vertebrate neuromuscular junction" http://www.annualreviews.org/doi/abs/10.1146/annurev.neuro.22.1.389?url_ver=Z39.88-2003&rfr_id=ori:rid:crossref.org&rfr_dat=cr_pub%3dncbi.nlm.nih.gov&
Bialas, A.R. ; Stevens, B Glia: Regulating Synaptogenesis from Multiple Directions. Curr Biol. 2002, 833-835
Overview
[edit | edit source]Up to 50% of neurons that develop die during the course of normal development. The fact that neurons that make incorrect connections are more likely to die suggests that cell death increases the overall accuracy of synaptic connections.
There are three evidences that suggest that neural cells die since they fail to compete with others and fail at life preserving:
- Implantation of an extra target site decreases neuron death
- Destroying some neurons before the period of neuron death increases the survival rate of the remainder
- Increasing the number of axons that initially synapse on a target decreases survival rate of the remainder
Overview
[edit | edit source]Synaptic rearrangement during development characterizes the vertebrate nervous system and was thought to distinguish vertebrates from the invertebrates. However, the examination of the wind-sensitive cercal sensory system of the cricket demonstrated that some identified synaptic connections systematically decrease in strength as an animal matures. Other examinations show that increase in strength over the same period. Moreover, a single sensory neuron could increase the strength of its synaptic connection with one interneuron while decreasing the strength of its connection with another interneuron. Thus, rather than being a hallmark of the vertebrate nervous system, synaptic rearrangement probably characterizes the development of many if not all nervous systems.
reference
[edit | edit source]http://www.ncbi.nlm.nih.gov/pubmed/3363372
Introduction
[edit | edit source]
Neurodegenerative diseases are disorders closely linked to old age and affect more than 120 million people world wide per year. Some neurodegenerative diseases were often identified by the misfolding and clumping of their proteins - which led to neurotoxicity.[2] But, this idea has been controversial and in light of recent research, there is a better understanding of the nature of proteins and their effect on neurodegeneracy. A prominent example of this controversial topic is Alzheimer's Disease and the aggregation of peptide β-amyloid (Aβ) in the brain.[3] β-Amyloid peptide can exist as a non-toxic substance in the brain, however, large deposits of this peptide are found in the brains of patients with Alzheimer's Disease. The connection between these degenerative diseases and protein aggregation have been experimentally studied: genetic mutations, increased gene dosage, and post-translational gene modifications. These experiments are associated to Parkinson's Disease, Alzheimer's Disease, and Huntington's Disease, respectively. Despite these experiments, these relationships are still poorly understood. It is commonly thought that neurodegenerative diseases are caused by aging, proteasomal and mitochondrial disfunction, oxidative stress, and abnormal protein-protein interactions via cytotoxicity, a more radical proposition suggests that macroscopic proteinaceous inclusions are also an important factor in the progression of these diseases. These macroscopic inclusions were included in many models for the diseases, but were only recently observed as oligomeric and prefibrillar species in living cells.
Amyloid Hypothesis - Aberrant protein interactions, which accumulate and result in neural defects and culminate into neurodegeneration, are believed to be causally related to the insertion of aggregated proteins into ordered fibrillar structures (such as amyloid membranes). This is a result of malformed proteins - their designated chaperones cannot recognize them anymore, nor can they be destroyed by ubiquitin-proteasome system. As a result, these proteins are able to remain in the cell and have hazardous effects on the cell. These cells can then form stable and insoluble amyloid assemblies consisting of largely β-sheets. Another assembly, oligomeric species, is smaller and is predicted to be either a precursor for the amyloid fibrils or abnormal intermediates in the amyloidogenic cascade.
Neurodegenerative Diseases
[edit | edit source]Alzheimer's Disease
[edit | edit source]This is one of the most common age-associated diseases and is often associated to dementia. As the age of the population increases, so does the frequency of Alzheimer's Disease. On a molecular level, one of the most characteristic features of Alzheimer's Disease are the extracellular amyloid plaques composed of Aβ and neurofibrillary tangles. Recognizing these hallmark characteristics is not enough, because despite being able to recognize these proteins, the mechanisms of their formations are still unknown. This is partially because most Alzheimer cases occur randomly without a clear signal when the disease takes root. In order to circumvent this, conformation-dependent antibodies have been developed to help examine protein aggregates and identify prefibrillar oligomers and fibrils.
Techniques Used to Study Neurodegenerative Diseases
[edit | edit source]Optical microscopy and live-cell imaging are types of techniques often used to study the development of neurodegenerative diseases.
Bimolecular Fluorescence Complementation (BiFC) - The technique is useful because it can be used in physiological environments (rather than extraction from the body). Furthermore, it helps elucidates protein-protein interaction with spatial and temporal resolution. It also elucidates the function of protein-protein interactions within a living cell. This method involves two non-fluorescent molecules that are fragments of a reporter protein and the two proteins of interest. If the two proteins of interest interact with each other, te the two fragments of the reporter protein come together and form a structure that mimics the protein's native form. This technique has been helpful in recent years because it has helped explain the connection between oligomerization and neurodegenerative diseases. It is also predicted that BiFC will be able to help determine how the inclusion bodies and oligomeric species are formed in the brain. As a result, BiFC can aid therapeutic research by predicting appropriate targets for drugs and by suggesting new methods of treating neurodegeneration.
This technique was used starting in 2002, and has many applications beyond neurodegenerative diseases. For example, it has been used to study the interaction between basic leucine zipper and Rel family transcription factors in their physiological state. This method has also been used on several model organisms such as mammalian cell lines, plants, nematodes, yeast and bacteria. Originally, it was used only to identify singular protein-protein interactions. But in more recent experiments, it has been used to study multiple protein-protein interactions by using multiple fluorescent proteins all with different emission spectra. This advancement has allowed for the study of subcellular localization, assessment of complex formation, analysis of the control over protein-protein interactions, and the ability to simultaneously observe changes in separate protein complexes.
Another variation of BiFC uses the reporter protein in conjunction with bioluminescence resonance energy transfer spectroscopy as an alternative to multicolor BiFC.
The Advantages of BiFC
There are two main advantages of BiFC over many of the older techniques.
1. This technique uses molecules that cause fluorescence after the protein-protein interaction occurs, because this interaction is so specific, it unlikely that any other interaction with be able to cause a strong enough fluorescence in the reporter protein.
2. The use of fluorescence complexes allows the protein-protein interactions to be studied within the organism, rather than removing the proteins of interest from its natural environment to stain and track its actions.
The Disadvantages of BiFC
1. This technique is an indirect method of identifying the protein-protein interaction and requires the use of a very specific, fluorescing reporter protein. As a result, it is not possible to use this method to study the interactions between the proteins if one of them is unknown. This issue as been overcome by producing libraries in which different cDNAs are attached to fluorescent protein fragments.
2. One of the limitations in using this technique to study neurodegeneracy is that when the fluorophore recombines when the proteins interact, than the complexed fluorophore will stabilize the interacting proteins which will prevent the desired interactions from being observed. Despite this disadvantage, it could also be beneficial because it allows for a more selective study of oligomeric and dimeric species. For neurodegenerative studies, this effect might be particularly useful because by stabilizing the protein complex, the complex will be able to exist longer in its interacting state. This will make it easier to study the more ephemeral protein-protein interactions. This technique now makes it possible to study the protein aggregation process because parts of the formation involve these short lived protein-protein interactions. 3. This method does not distinguish between dimers, oligomeric and higher-ordered species. Other techniques have to be used to accomplish this, such as flow cytometry or SDS-PAGE.
Electron cryomicroscopy - In conjunction with nuclear magnetic resonance (NMR) spectroscopy, this technique has be useful in determining the structure of β-Amyloid peptide-derived assemblies. This is important because a better understanding of the assembly of the Aβ peptide will help explain the effect of Aβ aggregation in the brain.
Key Characteristics of Neurodegenerative Diseases
[edit | edit source]Aberrant Protein-Protein Interactions (PPi)
[edit | edit source]Although protein-protein interactions are not yet clearly understood with current research techniques, it is known that they are important in understanding the resultant proteins and how they function in physiological conditions. A common result of aberrant protein interaction is the presence of cytoplasmic, nuclear, or extracellular inclusions. These inclusions are generally a result of a build up of misfolded proteins into insoluble nuclear or cytoplasmic amyloid deposits. The changes in protein-protein interactions are believed to lead to the formation of these inclusions. Although the formation of the inclusion mechanism is not completely understood, studies suggest that the formations of inclusions follow a generally uniform pattern, even when the misfolded proteins are different. As a result, it is necessary for more conclusive data about inclusion bodies to be determined in order to develop therapeutic techniques for neurodegeneration treatment.
The study of Protein-Protein Interactions In the past, protein-protein interactions were studied via co-immunoprecipitation and co-purification. In these procedures, the protein-protein interaction could only be observed by removing the proteins from their physiological conditions. The more recent protein microarrays also work in a similar fashion. Removing the protein from its environment introduces a source of error because protein-protein interaction is also affected by its environment and its removal from physiological conditions means that the proteins won't be acting as they usually do in the cell. In comparison, Protein-Fragment Complementation Assays (PCAs), functional analysis of compensatory mutations, and imaging-based techniques are able to more accurately identify protein-protein interactions by avoiding the removal of proteins from the cell.
Older techniques used to make protein-protein interaction images often times used fluorescence or bioluminescence resonance energy transfer microscopy, fluorescence correlation spectroscopy, and image correlation spectroscopy. Fluorescence resonance energy transfer microscopy is useful because it labels the two interacting proteins with two different fluorophores in vivo. When the donor fluorophore is excited, it will transfer energy to to the second fluorophore, the acceptor fluorophore, which will label the second protein. The distance between the two proteins is determined by the difference between the lifetimes of the fluorophores. This technique is limited though, because the distance between the two proteins must be less than ten nanometers, otherwise the emission spectrum of the two fluorophores will not overlap. This technique has been used to study the effect of mutations in the gene for amyloid precursor protein and its interaction with presenilin-1. This technique can also be used to characterize intramolecular and intermolecular interactions.
Fluorescene Correlation Spectroscopy is a bioimaging technique that uses theoretical analysis to interpret fluctuations and diffusion rates of fluorescently labeled molecules. Specific fluctuations are associated to specific interactions and aggregation patterns of interacting proteins. This method has been used to study how peptide β-amyloid aggregates form.
References
[edit | edit source]- ↑

Exchange of ions in the membrane. - ↑ http://www.sciencedirect.com/science/article/pii/S0968000410000952
- ↑ http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3144754/
Overview
[edit | edit source]Neuroglobin is a recently discovered member of the globin family. Only discovered in 2000, this protein's function is not entirely certain, but there are many hypotheses that define its use. Mainly, it has shown to promote neuron survival under hypoxia, which could potentially limit brain damage. It is an intracellular hemoprotein expressed in the central and peripheral nervous system, cerebrospinal fluid, retina, and endocrine tissues. It has hexa-coordinated heme-Fe atoms that display O2 affinities comparable to those of myoglobin.
Function
[edit | edit source]Neuroglobin's exact physiological role is still uncertain. Over time, this protein has evolved extremely slowly compared to the rate of hemoglobin and myoglobin. However, some likely functions are:
1. Neuroglobin enhances the O2 supply to the mitochondria of the metabolically active neurons. This hypothesis is supported because neuroglobin primarily resides in metabolically active cells and subcellular compartments. Also, the concentration of neuroglobin is closely correlated to the distribution of mitochondria; however, it is not entirely localized in this specific organelle. In neuroglobin, the fast autoxidation of ferrous (Fe2+) neuroglobin to ferric (Fe3+) Neuroglobin observed in vitro would rather inhibit an efficient binding of O2 to neuroglobin, but favors an involvement of neuroglobin in some type of redox reaction (seen below).
2. In cell culture systems, neuroglobin expression may be induced by hypoxia. It is unlikely that many nervous systems of animals are used to low O2 supply, so this theory is still contradictory. In many species of fish that must rely on lower levels of oxygen, the amount of neuroglobin is much higher than in species that do not deal with this.
3. Like other types of globin, neuroglobin associates with other gaseous ligands besides O2. Under an excess of NO applied in vitro, an Ngb-Fe2+-NO form is established by reductive nitrosylation, which then decomposes the very toxic ROS component peroxynitrite, resulting in Ngb-Fe3+-NO. Therefore, neuroglobin was proposed to have a similar role to that of Mb, acting as a NO-dioxygenase when PO2 is low and NO levels are increased. Under low-oxygen conditions deoxygenated Ngb may react with NO2–, resulting in the formation of NO.
Conclusion
[edit | edit source]The recent discovery of neuroglobin, displaying heme hexacoordination has a substantial impact on our understanding of O2 metabolism in man and other vertebrates. The vastly different expression patterns of the four globin types (Hemoglobin, Myoglobin, Neuroglobin and Cytoglobin) strongly suggest diverse roles. Furthermore, it is conceivable that neuroglobin will cast new light on the ancestral function of vertebrate globins in general, and within the nervous system in particular.
Protection against Alzheimer
[edit | edit source]Scientists have learned that neuroglobin protects cells from stroke damage, amyloid toxicity and injury due to lack of oxygen. Neuroglobin occurs in various regions of the brain and at particularly high levels in brain cells called neurons. Scientists have associated low levels of neuroglobin in brain neurons with increased risk of Alzheimer's disease. Recent studies have hinted that neuroglobin protects cells by maintaining the function of mitochondria and regulating the concentration of important chemicals in the cell. However, the exact mechanisms by which neuroglobin protects cells from dying a natural death has, until now, remained unclear. The study led by UC Davis biomedical engineering professor Subhadip Raychaudhuri and University of Auckland biological sciences Professor Thomas Brittain, found that neuroglobin preserves the functioning of a cell's mitochondria by neutralizing a molecule necessary for the formation of a type of protein that triggers the cell's collapse. The scientists think that the fundamental role of neuroglobin found in neurons is to prevent accidental cell death from occurring due to stress associated with normal cell functioning. Cells may protect themselves from triggering the chain of events leading to cell death by expressing a high level of neuroglobin. Mitochondria are tiny "capsules" within a cell that make most of the raw material the cell uses to produce its energy. Mitochondria also play important roles in communication within and between cells and important aspects of cell differentiation and growth. A cell dies quickly when its mitochondria stop functioning. Various kinds of stressors, such as lack of oxygen, low nutrient levels, increased calcium levels or presence of toxic substances can cause mitochondria to rupture and emit a molecule called cytochrome c. Cytochrome c binds with other molecules outside the mitochondria to form a protein called an apoptosome. The apoptosome helps build an enzyme that degrades and eventually collapses the cell. Neural cells can survive damage to the mitochondria if apoptosomes do not form. For their study, the researchers developed predictions from computational modeling and validated them with biological experiments. They found that neuroglobin binds to cytochrome c and prevents it from forming an apoptosome. This finding could offer new approaches to the prevention and treatment of Alzheimer's disease. In Alzheimer's disease, a toxic type of protein accumulates in brain neurons and leads to mitochondrial rupture and cell death. The finding suggests that high neuroglobin levels may buffer neurons against the effect of this protein by preventing apoptosomes from forming.
References
[edit | edit source]1. Burmester, Thorsten. "The Journal of Experimental Biology." What Is the Function of Neuroglobin? Journal of Experimental Biology, 03 Mar. 2009. Web. 22 Nov. 2012. <http://jeb.biologists.org/content/212/10/1423.long>. 2. Pesce, Alessandra. "Neuroglobin and Cytoglobin." US National Library of Medicine. EMPO Reports, 03 Dec. 2002. Web. 20 Nov. 2012. <http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1308314/>. 3. Subhadip Raychaudhuri, Joanna Skommer, Kristen Henty, Nigel Birch, Thomas Brittain. Neuroglobin protects nerve cells from apoptosis by inhibiting the intrinsic pathway of cell death. Apoptosis, 2009; 15 (4): 401 DOI: 10.1007/s10495-009-0436-5
Overview
[edit | edit source]Sensory systems provide the initial inputs to cognitive processes, and motor systems deliver the physical behavioral output that carries out movements. All body movements are generated by the stimulation of skeletal muscle fibers by the lower motor neurons- neurons whose cell bodies are located in the brain stem and spinal cord. The activation of lower motor neurons is coordinated by local circuits consisting of inter neurons, also located in the spinal cord and brainstem. Complex reflecxes and rhythmic locomotor movements can be generated and sustained by the coordinated activation of such local circuits in the absence of inputs from higher motor centers. At higher levels of the motor system, upper motor neurons in the cerebral cortex and brainstem provide descending control of local circuitry in the spinal cord and brain stem. The other two parts, the cerebellum and basal ganglia, modulate the activity of upper motor neurons in order to make online corrections in response to perturbations in ongoing movements and to help initiate goal directed movements.
Reference
[edit | edit source]Levinthal, Charles, "Drugs, Behavior, and Modern Society", Pearson Education, Inc., 2008 Purves, Dale, "Principles of Cognitive Neuroscience", Sinauer Associates, Inc., 2008 Basal Ganglia is a group of nuclei lying deep in the subcortical white matter of the frontal lobes that organize motor behavior. The caudate, putamen, and globus pallidus are major components of the basal ganglia. Basal ganglia appears to serve as gating mechanism of physical movements, inhibiting potential movements until they are fully appropriate for the circumstances in which they are to be executed.
-
Caption1
-
Caption2
Direct and Indirect Pathway
[edit | edit source]The net effect of basal ganglia activation through this so-called direct pathway is thus excitiation of cortical neurons. The subthalamic nucleus, on the other hand, forms part of an internal loop within the basal ganglia that via excitation of a portion of the globus pallidus has a net inhibitory effect on the cortical neurons, so called indirect pathway. The balance of excitatotry and inhibibory effects of the basal ganglia releases and coordinates desired movements.

It is speculated that when a voluntary movement is about to be initiated by cortical mechanisms, the direct pathway selectively facilitates certain motor programs in the cerebral cortex that are adaptive for the present task and the indirect pathway simultaneously inhibits the execution of competing motor programs.
Basal Ganglia Funtions
[edit | edit source]1. Rule-based habit learning system - initiating, stopping, monitoring, temporal sequencing, and maintaining the appropriate movement 2. Braking Function - inhibiting undesired movements and permitting desired ones 3. Action Selection - choosing from the potential actions, cortex plans to perform the ones that actually get executed 4. Other cognitive role - Motor planning, sequencing, learning, maintenance, predictive control, working memoery, attention, switches in behavioral set.
Relation to Parksinson's disease
[edit | edit source]In parkinson's disease, the selective death of neurons in the substantia nigra pars compacta that use the neurotransmitter dopamine increases the excitatotry tone of the direct pathway through basal ganglia. Patients with Parkinson's show a marked disruption in the ability to initiate voluntary movement. This decrease excitation in direct pathway which increase inhibition in indirect pathway, thus resulting more inhibition of thalamaus and less excitotry input to motor cortex.
References
[edit | edit source]Purves, Dale, "Principles of Cognitive Neuroscience", Sinauer Associates, Inc., 2008
Levinthal, Charles, "Drugs, Behavior, and Modern Society", Pearson Education, Inc., 2008
Overview
[edit | edit source]
Cerebellum is the prominent hindbrain structure that is concerned with motor coordination, posture, balance, and some cognitive processes. It is composed of three layered cortex and deep nuclei, and attached to brain stem by the cerebellar penduncles.
Cerebellar Funtions
[edit | edit source]1. Cerebellum combines and coordinates rapid and skilled movements. 2. Controls and corrects compounds, complex movements through feedback and timing 3. Gains control through trial and error. established through error-correction or supervised learning network. 4. With time and practice control, passes from effortful to effortless 5. Effects on equilibrium, posture, and muscle tone.
Organization
[edit | edit source]
The cerebellum is organized into an outer cortex of cells, deep nuclei ontaining output neurons and white matter fiber tracts along which inputs and outputs course. The cerebellum receives both ascending projections from the spinal cord and brainstem and descending projections originating in the frontal motor and parital cortices. Spinal cord inputs come mainly from muscle proprioceptors and convey information about ongoing muscular activity and the position of the joints in space directly to the medial portions of the cerebellar cortex.
Reference
[edit | edit source]Purves, Dale, "Principles of Cognitive Neuroscience", Sinauer Associates, Inc., 2008
Overview
[edit | edit source]Systems biology, network biology, or integrated biology, is an emerging approach applied to biomedical and biological scientific research. Systems biology is a biology based inter-disciplinary field of study that focuses on complex interactions within biological systems, using a wider perspective (holism instead of the more traditional reductionism) approach to biological and biomedical research. In its view, gene (or any molecule of interest) and its product (or smaller molecules, like cofactors, messenger molecules, and metabolites) are seen as “nodes”. The link between the nodes represent a relationship that can be a physical interaction, enzymatic reaction, or a functional connection. This system assumes that perturbations, both internal and external, affect the phenotype. Mass spectrometry (MS) is used to identify the network wiring, such as protein-interaction networks. Particularly from year 2000 onwards, the concept has been used widely in the biosciences in a variety of contexts. One of the outreaching aims of systems biology is to model and discover emergent properties, properties of cells, tissues and organisms functioning as a system whose theoretical description is only possible using techniques which fall under the remit of systems biology. These typically involve metabolic networks or cell signaling networks. Systems Biology has some important inferences: 1) requires a look at the biological processes as an integral; 2)provides new opportunities and depends on technology for advanced computational and experimental approaches.
Old. vs. New
[edit | edit source]In the past, studies have typically been carried out within the “one gene- one protein- one function” standard, called “molecular biology paradigm”. It is a mindset where it is assumes there is 1) a direct link between gene and protein function, where it is implied that genes and their translation products explain biological function, and 2) proteins are looked at individually and in a linear form (downstream and upstream). Although powerful technologies and remarkable technical advances have been made, a genotype-phenotype link still remains a challenge
Applying Mass Spectrometry
[edit | edit source]'Mass spectrometry (MS') is used for this proteomics study. Various strategies have been presented to address each biological need, where each strategy is only a portion of the “total proteome space”. The strategies are:
- • Shotgun/ Discovery: can identify numerous proteins in a sample, but most likely to only detect most abundant
- • Directed: specific, pre-determined set of proteins identified, and quantified at high level of reproducibility
- • Targeted: high reproducibility, and accurate for small, pre-selected portion of proteins, high detection sensitivity and dynamic range
- • Data dependent analysis (DDA): emerging, but attempt to identify all proteins in sample
Technologies to quantify and identify molecules are needed to quantify and identify the edges of networks, which has been address by 2 ways: a direct approach, where the edge is measured by catching an interacting molecule of the node by its higher affinity, and an indirect approach that uses an assay with a node between the molecule and node.
Networks
[edit | edit source]Two types of networks are distinguished in MS-based proteomics: protein interaction networks (PINs) which are undirected networks, so have no preference in directions, and protein signaling networks (PSNs), which are directed networks and have a preferred direction. PINs can be exemplified by the major MS-based proteomics interest, protein-protein interaction networks (PPINs), because the direct links between the nodes and edges in these networks are measurable. In order to study such networks, proteins are used to as a “bait molecule” to catch interacting proteins, then identify them by MS. To analyze PPINs, affinity purification mass spectrometry (AP MS) is preferred because it echoes the multi-directionality complexity of PPINs. Then, AP MS data are interpreted where two or more copurified nodes are connected to edges, forming a network, and can infer protein-signaling networks (PSNs).
Perturbation experiments are done to see the effects of changing an enzyme activity, like activation or repression in a cell that can be used to compare to in vitro experiments, which can give new insights of the rewiring network. There are two types of perturbations: the ones that affect mostly the edges, and those that effect the nodes, which consequently effect their edges. Phosphorylation networks have been heavily studied, where the downregulation of phosphopeptides with a kinase inhibitor lead to the question if it is kinase dependent, but perhaps not kinase mediated. The goal is to know how networks capture and take this information to produce a phenotype or to provoke a cellular response. Connecting network wiring to phenotypes is done by combining static PPINs, which is used to identify genes that have set network properties and can be correlated to a phenotype, with protein-DNA interaction data measured with microarrays. A search was then completed to identify genes with unusual numbers od edges, which could show a change in correlation in phenotypic subset of the samples. Then, an initial set of genes in a How reliable the conclusions from the data and network knowledge depend on multiple factors: coverage and correctness of the network, quality of the data, and method of correlation calculation. Also, the results do not automatically lead to causality.
General Roadmap
- 1) Define starting network of nodes from previous studies
- 2) Disrupt network components and present experimental data
- 3) Use information to improve network model and improve experimental data and phenotypes from network
references
[edit | edit source]Bu Z, Callaway DJ (2011). "Proteins MOVE! Protein dynamics and long-range allostery in cell signaling". Advances in Protein Chemistry and Structural Biology. Advances in Protein Chemistry and Structural Biology 83: 163–221
Bensimon A, Heck AJ, Aebersold R. Mass spectrometry-based proteomics and network biology. Annu Rev Biochem. 2012;81:379-405. Review. PubMed PMID: 22439968.
Overview
[edit | edit source]Systems biology is a computational way of modeling and analyzing complex biological systems and pathways. It has recently contributed largely to the field of biochemistry by serving as a tool to understanding complex systems at large, especially the analysis of programmed cell death (PCD), which has so many interconnected networks with loops and feedbacks that the task of modeling the entire system has been largely avoided until this time. There are largely three main processes of programmed cell death that have been successfully analyzed using systems biology: apoptosis, autophagy, and necrosis. All of these forms of cell death are controlled by the cell, whether they are stimulated or inhibited based on different factors, which are named "inputs" in systems biology. Inputs include protein concentrations, localizations, enzyme activation states, and kinetic factors of the environment.
Systems biology-Why and How
[edit | edit source]Generally, to understand a process of cell death, it would be logical to look at one factor, such as the exposure of the cell to one type of enzyme or a signal, and look at one output, such as cell lysis. However, it turns out that the global PCD network is much more complex and is non-linear. That is, there are multiple independent and dependent variables that affect and determine cell behavior at large. Therefore, one cannot just conclude that a change in one input will cause the cell to die, because there are many other interconnected inputs that may affect the cell more than that specific input. [1]
The systems biological approach is to gather data of numerous single pathway networks provided by bioinformatics or biochemistry, and to computationally connect those networks together into one large system to model cell behavior. To do this, systems biology uses multiple original differential equations(ODEs), which describe a change over a single independent variable, and partial differential equations(PDEs), which integrate multiple independent variables. ODEs generally focus on time as their independent variable, and it looks at the enzymatic activity that is determined experimentally. PDEs are used to look at spatial-temporal reactions(basically where the position of an object or a molecule is at a given time) such as diffusion. Using these equations, a Boolean logic is used to assign a number to each network(called nodes). A value of 1 simply means the network is on, and a value of 0 means the network is off. Thus, using the Boolean logic, one can determine which networks are on or off given certain conditions, or inputs. Combining the boolean values for all the networks, the final output, which for programmed cell death is the activation of caspases that initiate cell death, can be determined. [1]
There is also a data-driven approach to systems biology, which simply uses a large number of experimental data and statistically find correlations between each of the experimental data sets. This method is advantageous in that it does not require using complex differential equations, but the disadvantages lies in that it is limited to and dependent on the number and validity of experimental data. IN this method, linear algebra is used, instead of differential equations, to find correlation between the experimental data sets, clustering techniques are used to group and simplify the data, and partial least square analysis is used to predict data.[1]
3 Types of Programmed Cell Death
[edit | edit source]Apoptosis is mainly characterized by chromatin condensation and fragmentation, followed by blebbing, which causes the cell to be fragmented into apoptotic bodies. The apoptotic bodies are then finally disintegrated by caspase family of cysteine proteases.[2] Apoptosis is the most studied and the best characterized of the three cell death types, and both its intrinsic and extrinsic pathways have been successfully modeled using systems biology. Some of the main contributions to modeling apoptosis have come from the works of Krammer and Eils, who used ordinary differential equations to explain Fas-induced apoptosis. They wanted to predict the death output of the cells(output), in response to the concentrations of the Fas activating antibodies(input). Starting with a complex network, they clustered networks of signaling pathways that behaved similarly into submodules to simplify the system. In doing so, they also found the importance of an intracellular inhibitor pathway named c-FLIP, which induced apoptosis. Through their work, they showed that systems biology can successfully model a complex system of signaling pathways using submodules.
Autophagy is a process in which intracellular contents are engulfed and consumed by autophagosomes, which are multimembrane vesicles. Most of the characterization of autophagy have been made due to yeasst genetics, where a lot of autophagy genes and their functions were identified.[3] It plays an important role in homeostasis of the cell, because it gets removes damaged organelles and misfolded proteins. However, this engulfing process of intracellular contents can actually lead to cell death, and the pathway for this process has been modeled with the help of systems biology.
Necrosis is a process that involves cell swelling, organelle dysfunction, and cell lysis. Originally, it was defined as an uncontrolled event, or an accidental death that did not actually require any gene activity. However, recent research has shown that it is actually a genetically controlled event with specific pathways. Some of the identified regulators include c-Jun N-terminal kinase, apoptosis inducing factor, death-associated protein kinase, and reactive oxygen species.[1] The pathways for these regulators have been also modeled with systems biology, but are not yet fully understood.
Intrinsic and Extrinsic Pathways
[edit | edit source]Intrinsic Pathway is an activation of apoptosis caused by signals originating within the cell. A main trigger of the formation of apoptosome, which triggers apoptosis, is cytochrome c. Cytochrome c is found in between the inner and the outer membrane of the mitochondria. The intrinsic pathway increases the permeability of the mitochondrial outer membrane, which releases cytochrome c into the cytosol, thereby causing apoptosis. Therefore, the mitochondrial outer membrane permeabilization(MOMP) became crucial for understanding the intrinsic pathway to programmed cell death, and the stimili that activates this intrinsic death pathway is known as staurosporine.
Extrinsic Pathway is an activation of apoptosis due to a signal from an external source outside the cell. The stimuli that activates the extrinsic death pathway is called TRAIL, which stands for tumor necrosis factor apoptosis-inducing ligand.
Generation of the spatiotemporal model using partial differential equations for both of these pathways, and a comparison of this model to experimental data have shown that cytochrome c redistribution to the cytosol took longer for the intrinsic pathway than the extrinsic pathway, leading to the hypothesis that apoptosis occurs faster kinetically for the extrinsic pathway.[1]
References
[edit | edit source]- ↑ a b c d e Shani Bialik, Einat Zalckvar, Yaara Ber, Assaf D. Rubinstein, Adi Kimchi, Systems biology analysis of programmed cell death, Trends in Biochemical Sciences, Volume 35, Issue 10, October 2010, Pages 556-564, http://www.sciencedirect.com/science/article/pii/S0968000410000861
- ↑ Cohen, G.M. (1997) Caspases: the executioners of apoptosis. Biochem. J. 326, 1–16
- ↑ Nakatogawa, H. et al. (2009) Dynamics and diversity in autophagy mechanisms: lessons from yeast. Nat. Rev. Mol. Cell Biol. 10, 458–467
What Is Bioinformatics?
[edit | edit source]Bioinformatics is a rapidly developing field of science which use the advantage of computer technology to analyze the molecular biology. The method in bioinformatics field can be derived from statistics, linguistics, mathematics, chemistry, biochemistry, and physics. Sequence or structural data of nucleic acids or peptide chain as well as the experimental data can be used as data by the scientists in the bioinformatics field[1]. Specifically, the area structural biochemistry that involves bioinformatics deals with how sequence alignments are obtained and eventually how the analysis of the sequences can help generate phylogenetic trees. These relations can eventually help contribute knowledge about how structures of macromolecules are displayed and compared with one another.
Properties of a Protein Data Bank
[edit | edit source]Some of the most well known structures of macromolecules are archived as atomic coordinates. These atomic coordinates are data files that contain the three dimensional structure of molecular structures. The link of atomic coordinates further explains the specifics of these data files. The array of molecular structures are archived in the Protein Data Bank also known as (PDB). The link of PDB is the URL to find the many coordinates publicly provided. Many scientific journals that publish results on macromolecular structures now require researchers to upload atomic coordinates to the database. As a result, in this bank, there are almost more than 20,000 macromolecular structures which include proteins, nucleic acids, carbohydrates that were determined through techniques such as X-ray crystallography, diffraction techniques, nuclear magnetic resonance (NMR), electron microscopy, and theoretical models. This bank is growing larger as around 2500 structures are presented each year.
As a structure is determined, a four character identifier is associated with the macromolecular structure known as a Protein Data Bank identification code (PDBid). The first character must be a digit from one to nine, while the remaining three characters can be letters of upper or lowercase. For example, the myoglobin structure is coded as 1MBO in the PDB. However, it is important to note that the identifiers do not necessarily need to have a relationship with the name of the macromolecule.
First, the atom coordinate file begins with information such as the identity and properties of the molecule under study, the date when the file was submitted, the organism from where the macromolecule was obtained, and the author(s) who found the structure along with journal references. Furthermore, the file contains a description of how the structure was determined and symmetry and residues that were not investigated. The sequences of the many chains are presented with one another with a description and formulas that accompany it which are called hetrogen groups (HET). HET are molecules that are not like standard amino acid or nucleotide residues such as organic molecules like the heme group, residues like Hyp, metal ions, and water molecules bound to other molecules. This file continues on to provide elements of the secondary structure along with any disulfide bonds present. Majority of the PDB file contain two series- the standard residue also known as ATOM and heterogens known as HETATM record lines. Each of these series, the ATOM and HETATM provide coordinates for a specific atom in the structure in correspondence to its serial number. Following the series, the atoms Cartesian coordinates (X,Y,Z) are presented relative to the fraction of sites that the atoms space occupies. Normally, this arbitrary origin is quantified as 1.00, but with groups that contain many conformations, or molecules that are not fully bound to a protein, the number is positive and less than one. In addition, an isotropic temperature is described since it can present the thermal mobility of the atom. A larger quantity of isotropic temperature means there is greater motion involved. If the structure was determined through NMR, the PDB would contain ATOM and HETATM series for the most representative member in a coordinate set that was calculated in finding the structure. Finally, the PDB file terminates with connectivity records (CONECT) which present non standard entities between atoms such as hydrogen bonding and disulfide bonds.
Properties of The Nucleic Acid Database
[edit | edit source]Similar to the Protein Data Bank, the Nucleic Acid Database (NDB) contains the atomic coordinates of nucleic acids. The following link of NDB is the direct URL of the database. The format of the file that nucleic acids is like that of the PDB files. The NDB however, has a contrasting organization and algorithms for searching that is specific to nucleic acids. This feature is particularly important since proteins are categorized by names like myoglobin whereas the identity of nucleic acids is defined by their sequences.
Viewing Macromolecular Structures in Three Dimensions
[edit | edit source]Studying the three dimensional structure is relatively important, as it provides much information to reactive sites along with functions of a macromolecule. The most revealing way to investigate the structure of a macromolecule is by utilizing molecular graphic programs. One useful program is called PyMOL. The following link is the direct website to PyMOL, and the programs capabilities of viewing a three dimensional structure. Programs like PyMOL allow a user to actively engage with a molecular structure by rotating it and obtaining an impression of the molecule that can enhance the understanding of the molecular rather than seeing it in two dimensions. PyMOL, along with many other popularly used programs like RasMol utilize PDB files as input for further visualization.
Structural Classification and Comparison
[edit | edit source]Many proteins that discovered are structurally related to other proteins. This similarity is due to evolution conserving the structures of proteins instead of their protein sequences. The following set of descriptions are some of the many websites for the public that have computational tools in order to classify and compare protein structures. By using these tools, the functions, distant evolutionary relationships that are not normally displayed in sequence comparisons, generation of special libraries of unique folds for the prediction of structures, and explanation of why certain structures are more dominant than one another are examined.
1. Class, Architecture, Topology and Homologous superfamily (CATH) categorizes proteins using the four topics into its respective structural hierarchies. First, "Class" is the highest level and has four categories of secondary structure. These are: Mainly alpha, Mainly beta, alpha/beta, and not many secondary structures respectively. Secondly, "Architecture" is the arrangement of the secondary structure that is separate from that of topology. Thirdly, "Topology" refers to both the holistic view of the proteins connectivity and shape. Fourthly, "Homologous superfamily" are proteins that are homologous to the protein that is selected. Furthermore, an interactive or still view of the protein can be displayed. An example of a CATH for myogolobin would be Class: Mainly alpha; Architecture: orthogonal bundle; Topology: globin-like; Homologous superfamily: globin. As a result CATH allows users who access the database to browse up and down to make a comparison of the many structural hierarchies.
What is the advantage of Bioinformatics?
[edit | edit source]1. Create an e-library of biological database
[edit | edit source]- Biological database is the organized biological information stored electrically and able to revive. For example, a biological database can be a record of a nucleic acid sequence with the name, input sequence, the scientific name of the organism it was isolated from[2].
- In this computing era, the storage database give a great convenient for the communication between scientists. The data in the e-library can be used widely by people from scientists , students to knowledgeable laymen.
2. New methods to interact with molecular biology
[edit | edit source]- Since analyzing molecular biology is one of main fields in bioinformatics, bioinformatics researches focus on creation of new tools, the methods to storage, retrieval and analysis the material such as protein sequences.
- The methods to analyze target samples are usually computer programs which will help researchers determine the structure of interesting sample or help scientists enable determine the family group for the sample from storage data. One common program used in bioinformatics is BLAST, Basic Local Alignment Search Tool. The outcome of BLAST search is a list of sequence alignments which will help researchers identify homologous sequences of the sample sequence from the database of known sequences[3].
3. Explore evolution
[edit | edit source]- Proteins with a common ancestor will have resemble amino acid sequences[3]. Therefore, with the information of sequence and structural data, scientists can organize an unknown protein into groups and reconstruct the evolution of the protein. Sequence alignment method is a technique to detect homologous genes or proteins. The evolution relationship of two genes or proteins will determine by calculating the score with identity matrix or substitution matrix . Structural alignment method, comparing tertiary structure of proteins, also can explore the evolution relationship of two protein sequences. Then, scientists can create evolution tree for proteins as well as for the life in this planet[3].
Related Fields
[edit | edit source]Fields that are related to bioinformatics include[4]:
Biophysics- a field where biology is investigated using the techniques and concepts found in the physical sciences.
Pharmacogenomics- as it relates to bioinformatics, a field where the techniques of bioinformatics are used to store and process pharmacological and genetic information of the whole genome.
Pharmacogenetics- similar to pharmacogenomics, it uses bioinformatic and genomic techniques to focus on one to a few genes and identify the correlates of genomes.
Medical informatics- is a discipline where computer applications such as algorithms and structures are used to help effectively convey and process medical information.
Mathematical biology- is a field that focuses on using mathematical tools and methods to represent, evaluate, and model the processes of biology.
Computational biology- much like bioinformatics, involves using computer applications and statistical methods to solve biological problems. As such, biological modeling, simulation, and imaging make techniques such as RNA structure and gene prediction, sequence alignment algorithms, and multiple sequence alignment possible.
Proteomics- is the study of the proteome. The proteome is complete collection of proteins that is expressed by a cell, tissue, or organism. The proteins are complementary for a specific genome.
Genomics- the purpose of this scientific branch is to investigate the genome, an organism's complete DNA sequence, through using methods of DNA sequencing and mapping.
Cheminformatics- is the use of computers and information technology to solve problems found in chemistry.
Reference
[edit | edit source][1] Nelson, David L. and Cox, Michael M. Lehninger Principles of Biochemistry. New York: W. H. Freeman & Company. 2008
[2] National Center for Biotechnology Information <http://www.ncbi.nlm.nih.gov/>
[3] Berg, Jeremy M., Tymoczko, John L. and Stryer, Lubert. Biochemistry. New York: W. H. Freeman & Company. 2007
[4] Bioinformatics Organization. 2010. <http://wiki.bioinformatics.org/Bioinformatics_FAQ>

Homology is a concept that takes into account similarities that occur among nucleic acid or protein sequences of two different organisms. Coined by Richard Owen in 1948, homology is quantized by comparing matches that occur between two different samples of amino acid sequences in proteins or DNA sequences in DNA and assigning a system of point values to identical/similar matches that occur in alignments. This type of analysis is useful in determining relationships between species and can help to trace ancestral descent as well as evolutionary changes that have occurred over time in a given set of species. Today, techniques have been developed to assess the probability of two organisms being homologous and has largely become the main area of focus for bioinformaticians around the world. Homology among nucleic acid are of two major types: orthologous and paralogous. Homologous said to be orthologous if they were separated by an event called speciation. orthologous gene are found in different species, but similar to each other in which they originate from the same common ancestors. orthology often have the same function. paralogous are genes that separate by a gene duplication event. paralogs mostly have the same functions. The genes that encoding hemoglobin and myoglobin are consider paralogs genes. Also hemoglobin A, A2, B, F are paralogs of one another.
Misuse of the Term
[edit | edit source]The term “homology” is often mistakenly used when describing proteins or nucleic acid sequences due to the fact that “homology is a concept of quality and cannot be ‘quantified’[1]”. In a recent analysis, the term “homology” was searched on PubMed in the 2007 database and 1966 abstracts contain the word homology either in the title or the abstract, discarding those which used the term as part of the name of a protein or procedure. Of these abstracts, 57% (1128) properly uses the term while 43% (828) uses the term incorrectly. Some of the incorrect usage of the term includes association with a percentage value and terms such as “high”, “low”, and “significant”. Analyzing the term for the abstracts in the 1986 database shows that the frequency of misusing the term “homology” has slightly decreased.[2]
The analysis of the term was also performed across languages. In the 1986 search of articles containing homology, there was an overall lower percentage of articles that misused the term. However, as other countries have surged in scientific research, more articles from rising countries have been produced and a greater percentage of those articles have misused the term homology. The article "When It Comes to Homology, Bad Habits Die Hard," advocates a solution to this problem by requiring scientific journals to promote guidelines on proper usage of common terminology as well as education of new researchers on terminology in rising countries.[2]
The misuse of the term Homology is considered a problem due to the confusion that it would cause the reader in trying to understand the author's intention. For example, the author may state that two proteins are homologous while also making the statement that the two proteins do not share the same evolutionary origin (which is the definition of homology). The author may also state that two peptide chains are homologous while completely ignoring any discussion if they shared the same evolutionary origin. Authors were also found using the term as evidence that the proteins were from the same evolutionary origin (i.e. "The fact that these proteins are homologous proves that they are from the same evolutionary chain").[3]
An example of the difference between homology and similarity would be comparisons between human and chimpanzee DNA vs a comparison of human and mice DNA. While mice and humans share about 97.5% of their DNA with humans, that does not mean they possess the same evolutionary origin. While very similar, they would not be homologous.[4] Humans and chimpanzees, however, share >98.0% of their DNA and are believed to share the same evolutionary origin. Therefore, human and chimpanzee DNA strands may correctly be stated to be homologous.[5]
Orthologs
[edit | edit source]Orthologs are specific gene sequences that are closely related between two entirely different species, but often have the same functions. The term ortholog stems from the root "ortho" meaning "other" and was coined by Walter Fitch in 1970. Separated by a speciation event where a species diverges into two separate species, divergent copies of a solitary gene result in the orthologous homologous sequence.
An example of orthologous genes are the genes that code for hemoglobins in both cows and humans. The mapping of orthologs can help biologist construct evolutionary trees that are much more detailed and specific. Taxonomy and phylogenetic studies benefit from orthologous sequences. A simple example can be a bat and a bird; a bird and a bat are part of two different species and yet their wings have the same function.
Paralogs
[edit | edit source]Paralogs refer to gene sequences that are shared by organisms in the same species but exhibit different functions. Paralogs are usually the product of gene duplication which can be caused by any number of mechanisms such as transposons or unequal cross-overs. These duplicated genes typically have similar functions and can mutate further to take on other functions which results in the paralogs.
The number of differences or substitutions are proportional to the time that has passed since the gene has become duplicated. Thereby shedding light upon the way genomes evolve. Myoglobin and hemoglobin are considered to be the ancient paralogs which all evolve from.
Suspected paralogs are the genes that encode for hemoglobin and myoglobin as both have similar protein structures but differ in their oxygen-carrying duties. There are four known classes of hemoglobins (hemoglobin A, hemoglobin A2, hemoglobin B, and hemoglobin F) where are all paralogs of one another.Other examples of paralogs are Actin and Hsp-70. Their tertiary structures are similar but their functions are different; actin is part of the cytoskeleton, while Hsp-70 is a heat shock protein.
Sequence Alignments Detect Homologs
[edit | edit source]To test whether or not two molecules are homologous, it is important to examine the nucleic acid or protein sequence for matches that occur between the two sequences. Although forms of sequencing work, the protein sequencing is usually preferable because it's composed of 20 different building blocks (amino acids) while DNA and RNA are each only comprised of four nucleic acids; so having a significant number of matches in protein sequencing is much stronger evidence of a common ancestry than nucleic acid sequencing. Also, redundancy in the genetic code where different genes can encode for the same amino acid (e.g. GCU,GCC,GCA,GCG all code for Alanine) makes the comparison of proteins much more sensitive and useful in determining similarities in protein function than with DNA or RNA.
Two different protein sequences can be compared by analyzing the number of times that their amino acids match when aligned directly above each other or when one sequence is slid past the other. For instance, when assessing the number of matches, amino acid one of the top strand can either be directly above amino acid 1 from the second strand or slid to the left/right of it thus causing different amino acids to align. The number of matches are then plotted against the alignment in order to assess what alignment the maximum number of matches occur. It is important to understand that a large number of matches does not mean the two proteins are homologs.
To account for mutations such as insertions and deletions, gaps may be inserted to create a better match. If two sequence comparisons appear to be a good match, a gap may be inserted to accommodate both comparisons. Scientist score the alignment: +10 points for each match and -25 points for each gap no matter the size. This score must then be plotted against a distribution of other scores obtained by randomly shuffling one of the protein strands and comparing it to the other many times to ensure the amino acid matches were not just due to chance. If the score deviates largely from the majority of the scores, then the two proteins are probably homologs. However, a low score does not rule out homology.
Homolog Sequencing Technology: Matrices
[edit | edit source]

Scores may be calculated using identity or substitution matrices. This process can be more precise by selecting a matrix that adds in gaps to further match the sequences. Examples of matrices include PAM, BLOSSUM (a type of substitution matrix) GONNET (a matrix that specifically targets distance), DNA identity matrix, and a DNA PUPY matrix. Overall, the substitution matrices are most sensitive to protein sequences. By using these matrices, it is possible to detect distant evolutionary relationships. If two sequences are at least 25% homologous identical it can be determined that these two proteins are homologous. However, sequences with percentages lower than 25 are not necessarily not homologous. For example, if protein A is homologous to protein B (based on their identity percentages), and protein B is homologous to protein C, A and C are likely to have similarities in function even if they are only 15% identical. Identity matrices assign a value of one for matches between sequences and zeros for non-matches. This method does not distinguish between likely and rare mutations and therefore does not give a clear answer to homology. Substitution matrices account for conservative mutations that are less likely to be deleterious or seriously change the function, such as switching glycine and alanine, by giving them a large positive score. So in other words, substitution matrices take into account not only if the sequences are identical (giving them the highest possible score), but unlike identity matrices they also assign values for amino acids sequences when they are "substituted" by another amino acid with similarities. The more simililar the amino acid sequence, the bigger the "value" it receives. The more different the sequences are or "rare" the substitution of a given amino acid like A would be substituted for something like P, the bigger their "negative" values get. By making a distinction between the different types of mutations, better matches can be made and alignments based on random chance are avoided.
Identity Matrix : Identity matrices uses scores of one and zero where the matching of identical amino acids or nucleotides results in a score of one and any mismatches are given a score of zero. This is not very as meaningful because random shuffling scores may be in the same area as the original score.[6]
GONNET : Gonnet matrices uses “exhaustive pair-wise alignments” of proteins and measure the distances to estimate alignments. This creates a new distance matrix which refines the alignment score. This type of matrices showed if the proteins were derived from close or distant homologous proteins. This type of matrix was formed in 1993 by Gonnet with the help of Cohen and Benner.[7]

DNA PUPY
: DNA Pupy matrices give scores for purine-purine transitions as well as pyrimidine-pyrimidine. It is believed to be helpful in finding primers for PCR.[8]
PAM : Point accepted mutations (PAM) is a set of matrices used for the scoring in sequence alignments. PAM was introduces in 1978 by Margaret Dayhoff, an American physical chemist and bioinformatist. PAM is used to develop a scoring matrix which is used to determine the homology of two genes or proteins. The matrix is normalized so that PAM1 would give substitution probabilities for sequences that have 1 point mutation for every 100 amino acids. The most commonly used is PAM250, where the probability is determined for 250 point mutations for every 100 amino acids.
BLOSUM 62 : BLOSUM 62 is the most commonly used substitution matrix. A program was developed by the National Center of Biotechnology Information(NCBI) to do this sequence alignment and is available online. This substitution matrix tallies points for different amino acid pairs and accounts not only for the identity but also for the conservation (how similar an amino acid is to another as to not induce a dramatic change in the function of a particular protein) and frequency(how many times the amino acid shows up on the protein sequence) of the amino acid pairs. The matrix will give a higher score if the amino acids are identical but is also going to give points based on the similarities. For example, isoleucine and valine will be given a higher score because although the amino acids are not identical, they are similar in that both are hydrophobic.
Homology Modeling
[edit | edit source]The primary goal of homology modeling is to study the structure of the macromolecules. X-ray crystallography and NMR are the only ways to provide detailed structural information; however, these techniques involve elaborate procedures and many proteins fail to crystallize or cannot be obtained or dissolved in adequate quantities for NMR analysis. Therefore, model building on the basis of the known three dimensional structure of a homologous protein is the most reliable way to obtain structural information about the unknown protein. These are the main steps in homology modeling:
1. Finding homologues protein database files (the template) Template selection is a critical step in homology modeling. Template identification can be aided by database search techniques.
2. Creation of the alignment, using single or multiple sequence alignments.
When more than one known is involved, the knowns will align together, then the unknown sequence aligned with the group; this helps ensure better domain conservation) furthermore, the alignment can be corrected by the insertion or deletion of gaps. Even though introduction of gap complicates the alignment, there are developed methods that use scoring systems to compare different systems and penalize gaps to prevent the unreasonable insertions. Scoring of alignment involves the construction of identity matrices and substitution matrices. Substitution matrices are believed to be the best, theses methods are based on the analysis of the frequency with which a given amino acid is observed to be replaced by other amino acids among proteins for which the sequences can be aligned.
3. Model generation: The information contained in the template and alignment can be used to generate a three dimensional structural model of the protein, which is represented as a set of Cartesian Coordinates.
4. Model Refinement: The major sources of error in homology modeling are the poor selection of template and inaccurate template-target sequence alignment. This can be improved by using multiple sequences and structural alignment.
References
[edit | edit source]- ↑ Lewin, R. (1987) When does homology mean something else? Science 237, 1570
- ↑ a b "When it comes to homology, bad habits die hard." Trends in Biochemical Sciences. Volume 34, Issue 3, March 2009, Pages 98-99.
- ↑ Reeck GR. "Homology" in proteins and nucleic acids: a terminology muddle and a way out of it." Cell Magazine, Volume 50, Issue 5, August 1987.
- ↑ Coghlan A. "Just 2.5% of DNA turns mice into men" New Scientist, http://www.newscientist.com/article/dn2352-just-25-of-dna-turns-mice-into-men.html May 2002
- ↑ Choi C. "Monkey DNA Points to Common Human Ancestor." Live Science, http://www.livescience.com/health/070412_rhesus_monkeys.html April 2007
- ↑ [Berg, Jeremy M., John L. Tymoczko, Lubert Stryer, and Jeremy M. Berg. Student Companion for Biochemistry, 7th Edition, International Edition. New York: W.H. Freeman, 2011.]
- ↑ [Rastogi, S. C., Namita Mendiratta, and Parag Rastogi. Bioinformatics Methods and Applications: Genomics, Proteomics and Drug Discovery. New Delhi: Prentice Hall of India (P), 2006.].
- ↑ Matrices Tutorial
There are over a million different genes that fold into tens of thousands of different protein structures. Homologous structures therefore, must exist. Because there are a limited number of structures, two proteins can have very similar structures, and that's where sequence alignments step in. The theory of homology comes from experimental evidence of similarity in genes and proteins from evolution. Little is known about most genes so homology can be used to predict a gene's function. There are two types of homologs: paralogs and orthologs. Paralogs are found in the same organism with similar genomic structure, such as hemoglobin and myogoblin, but serve different functions. Orthologs are the inverse of paralogs; they are found in different organisms but essentially serve similar functions in its host organism, hinting evidence of evolutionary ancestry.
The human genome consists of over 3 billion base pairs and over 25,000 genes. Alternative splicing is what allows genes to encode numerous proteins.
Sequence Alignments can be used to detect homology between two polypeptide chains. Figuring out sequence alignments can help develop evolutionary origins and trace back the function, structure, and mechanism of a genome. Repeated motifs can be detected by aligning a sequence with itself. More than 10% of all proteins have two or more regions that are similar to one another. An example of this is the protein that binds to the TATA box which is comprised of two similar regions determined by sequence aligning the protein with itself. The three dimensional structure for this protein has been elucidated and the two similar regions have been confirmed.
The percentage of similarity between two gene sequences is known as the best possible alignment among all alignments that can be made to the sequence.
The simplest way to compare protein sequences is to align each strand and count for matching residues. The sequence is slid down one residue and each the sequences are realigned and matched again. Continuing this process for all possible combinations of alignments produces an alignment score for each combination.
Amino acids can be very similar to each other, and therefore replace one another over the course of an evolutionary period. Sequence alignments acknowledge this by including mismatches while accounting for probability and percentage of identities.
Newly elucidated protein sequences can be aligned by inputting the sequence into a large database of previously sequenced proteins. This procedure is a called a BLAST (Basic Local Alignment Search Tool) search. Using blast, homology of a newly sequenced protein can be determined, as well as predict function and tertiary structure of a protein. The first completed genome using the bacteria, Haemophilus influenza, identified roughly 1743 protein sequences. Using a BLAST search, researchers were able to identify possible function and structures for 1007 of these protein sequences.

Sequences are the amino acids for residues 120-180 of the proteins. Residues that are conserved across all sequences are highlighted in grey. Below the protein sequences is a key denoting conserved sequence (*), conservative mutations (:), semi-conservative mutations (.), and non-conservative mutations ( ).[1]
Homology
[edit | edit source]With the thousands of genes that currently exist, it's less feasible to deduce complete information about a gene and more feasible to compare genes are proteins through evolutionary characteristics. Homologous genes and proteins therefore, are the proteins and genes that have markedly similar characteristics.
Two sequences can be extremely similar with identical evolutionary backgrounds, however, over the years the sequence could have lost a set of amino acids or proteins that barely affect the function of the gene or protein. Similar amino acids can also replace each other and have little to no effect on the function of the gene or protein. These substitutions between proteins or genes still are homologous.
Gaps
[edit | edit source]Gaps are introduced when a sequence can be better aligned to encompass an increased amount of matching residues. For example, if two alignments appear to be a good match, a gap may be inserted to accommodate both alignments. Gaps also reflect upon the insertions, deletions, and mutations of nucleotides over time.
Gaps Increase Complexity
In principle, any arbitrary size and number of gaps can be added to any place of a sequence. To avoid an excessive number of gaps and deter further from the original sequence, scoring systems with penalties are used. One example is giving a penalty of -25 a gap of any size. However, each new sequence aligned based on the gaps receives a score of +8. If there are 50 new identities and 1 gap, the score would be [(50*8)-(1*25)], the score would be 375. In a sequence with 86 residues, there would be a 50/86 % identity match. The total score is calculated into a percentage of identity [see below], indicating the statistical probability of sequence similarity.

Shuffling
[edit | edit source]To check if the original sequences are accurate, the original sequences are shuffled randomly. The matching residues of the random sequences to the original sequence to produce an alignment score. Then number of matching alignments are compared between the alignment score of both the original and random sequences.
When comparing the un-shuffled alignment score with the shuffled alignment scores, if the un-shuffled alignment score deviates far from the mean and standard deviation of shuffled scores (is an outlier), this indicates that the sequences are likely homologous and the similarities and not simply due to chance. The probability of the un-shuffled alignment score deviating greatly from the shuffled alignment scores is approximately 1 in 1020,[2] indicating the likelihood of the authentic alignment to be unique in terms of alignment of bases. This method does not rule out homology.
Identity Matrices
[edit | edit source]An identity matrix is a way to evaluate the likeness of two different sequences of amino acids. In an identity matrix, the two sequences are given a point for every time there is an exact match in amino acids. It is all or nothing, the two amino acids either match, or they don't. Identity matrices are not as accurate in evaluating the likely hood of two sequences expressing homology because there are often mutations in amino acid sequence that either doesn't change the function of the protein, or does very little to change the function. These usually occur with similar amino acids such as Leucine and Isoleucine. Because of this factor, other techniques such as the substitution matrix is preferred.
Substitution Matrices
[edit | edit source]Homology is an important tool in evolutionary biology. Substitution matrix is one way to study homology in that it describes the similarities in protein sequences or in DNA Sequences. It accomplishes this by assigning a point system where the two sequences are compared to their randomized sequences. Amino acids have a certain ability to mutate into another amino acid. Hydrophobic amino acids, (i.g. Valine) have greater chance to mutate to another hydrophobic amino acid (i.g. Leucine). Substitutions that are made often receive a high positive score and rare substitution have given negative scores. Identical amino acid matches are also given points in a substitution matrix. There many types of substitution matrix that have been developed that have assigned different points for substitution examples are PAM, Blosum, BLAST matrices. These matrices are 20X20 matrices for protein. Blosum ( block substitution) matrices calculates homology by comparing blocks of conserved sequences in many sequence alignments compared to the identity sequence. The blocks are assumed to have functional significance in evolutionary biology.
Sequence analysis using substitution matrices are much more sensitive than identity matrices because it accounts for conservative substitutions that may have happened over time which do not significantly alter the structure of the protein. Substitution matrices can detect homology between sequences that would have otherwise been found not homologous using simple identity matrices.

Probability of Identity
[edit | edit source]If the 2 sequences are greater than 25% alike in a chain of at least 100 amino acids, the likelihood that they are homologs are high. If the 2 sequences are less than 15% alike, the likelihood that they are homologs are low. Between 15% and 25%, other methods, such as comparison of the tertiary structure, must be done to confirm homology.
Sequence Templates
[edit | edit source]In sequence alignment, certain amino acid residues are more important to the function of the protein than others and are more highly conserved throughout evolution. The areas that are critical to function and the amino acid residues comprising that area can be determined by examining the three dimensional structure of the protein. For example, the globin family (hemoglobin, myoglobin, leghemoglobin) of proteins that bind to oxygen, bind oxygen via a heme group that is comprised of a histidine residue that interacts with the iron in the heme group. This histidine residue is conserved in all of the globin family of proteins. This region that is significant to globin proteins can be used as a sequence template that is characteristic of this family of proteins. Newly elucidated protein sequences can then be matched to this sequence template to match that protein with certain families or to determine whether the new protein has similar functions to those families.
Methods of Sequencing
[edit | edit source]The Sanger Dideoxy method is used to sequence DNA. This process is a fast and simple one in which it involves the use of DNA polymerase to synthesize a complementary sequence containing fluorescent tags on the four deoxyribonucletide bases. The fragments of DNA strands containing the fluorescent bases are then separated via electrophoresis or chromatography then sent through a detector. Another method to sequence genomic DNA is the Shotgun method.
Edman degradation is used to sequence proteins. Phenyl isothiocyanate reacts with the amino group in the N-terminal amino acid, then acidified to remove it. High pressure liquid chromatography (HPLC) is used to identify the amino acid. The process is repeated for each of the following proteins.
Databases
[edit | edit source]Isolating and comparing an individual strand with any given strand can be tedious and time consuming. Therefore, there exists databases with homologous sequences that can be readily obtained and utilized. The methods of sequence alignments as listed above are tremendously useful when utilized alongside the broad databases and resources in available on the Internet.
PAM and BLOSUM matrices are two of the most frequently used scoring techniques.
BLOSUM, or Block Substitution Matrix, is a technique that measures local multiple alignments of related sequences. BLOSUM 62 is the default matrix for BLAST. BLOSUM 62 requires 62% sequence identity, while BLOSUM 80 would require 80% identity, etc.
- Basic Local Alignment Search Tool (BLAST) is located at the National Center for Biotechnology information. The individual amino acid sequence can be searched through the web browser. There are over 3 million sequences are in the database. In addition, the amino acid sequence entered can be compared with a chosen genome (such as humans), all the genomes currently in the database. The database gives a list of sequence alignment and a percentage of identity. It will look for similarity between DNA or protein sequences. The website is [4].
PAM stands for Percentage of Acceptable Point Mutation per 10^8 years. This process measures the global alignment of similar proteins. This practice requires the sequence to be less than or equal to 1% divergence. The mutation probability provides the scores over a period of time by column X, representing amino acid mutation, vs. row Y, the product of mutation. By multiplying this matrices by itself repeatedly, new matrices can be made to measure greater evolutionary distances.
There are three main Databases for DNA: Genebank, EMBL,DDBJ. These contain numerous entries that are the DNA sequence of genes and other DNAs such as genetic mapping markers discover and cloned by scientists so far. Each sequence entry was assign a unique accession number.
NCBI(National Center for Biotechnology Information)- a collection of databases and analysis tools. This site is supported by the National Institutes of Health and free for researchers or anyone who is interested in it. You can simply get on website: http://www.ncbi.nlm.nih.gov/ and search for a sequence of protein, DNA, RNA...etc. Many of database with NCBI are linked through a search and retrieval system called Entrez which allows for text-specific searches using key words.
ExPASy(Expert Protein Analysis System)- a very useful collection of protein and amino acid sequence analysis tools that is part of the server of the Swiss Institute of Bioinformatics. website: http://www.ca.expasy.org
Protein data bank- a database of protein structural information. website: http://www.rcsb.org/pdb/
Clustal W- An online amino acid sequence alignment program that is part of the European Bioinformatics Institute website. This is a powerful website for compaing protein sequences, after align, one can click on "show colors" to view a color based representation of amino acid similarities. website: http://www.ebu.ac.uk/clustalw/
How to look up sequence in Genbank
[edit | edit source]The following will be a step by step guideline of how to use program and website available online:
1. Go to the NCBI home page. (http://www.ncbi.nlm.nih.gov/)
2. The menu bar next to "all databases" should have all the different types of databases available. Pick the appropriate one. For example, if you want to find DNA sequence, you will pick nucleotide.
3. Use "key words" to find the sequence. It will have many varieties of options. which one is the one we are looking for then? If we are trying to find a DNA sequence that contains the entire coding region of the gene then we will have to find something with mRNA which introns were taken out already or complete cds of the coding sequence. It will be easier for one to find the sequence desire by typing down the domestic name of the animal (if you are looking for animal's gene).
4. Accession number is the ID tag for the specific sequence which appears in blue once one find the sequence desire.
5. The DNA sequence is given at the bottom of the page and numbering for the nucleotide in the sequence is given to the right.
6.CDS stands for coding sequence.
If one wants to find a homology then BLAST will be use:
1. Go to the NCBI homepage and click on BLAST. They are many different option of align, in this case, we will pick nucleotide blast.
2. Type the unknown sequence into the large field. For choose search set, one will pick others. then BLAST it.
3. Then a page of summary of the matches query nucleotide sequence is given from highest similarity (top) to least(bottom).
4. Query coverage and maximum identity columns are available too. Query coverage will show us the percentage of nucleotide that were the same or how well they match up. Then the homology of your unknown sequence will be determined.
BLAST can also be use to compare or align two DNA sequences to see how similar they are:
1. Get the entire gene sequence for both sequences one wants to compare (Like mention before.)
2. Open BLAST homepage and click on align under Specialized Blast.
3. In the query sequence box, you can either enter the accession number or the whole sequence.
4. Program selection has many different program you can use. After selecting appropriate one, click BLAST. Then you will align the two selected DNA sequences.
Summarize:
Three stage approach to genome sequencing
The initial stage:
Cytogenetic maps based on this type of information provided the starting point for more detailed mapping. With these cytogenetic maps of chromosomes in hands, the initial stage in sequencing the human genome was to construct a linkage map of several thousand genetic markers spaced throughout the chromosomes. On the stage, the order of the markers and the relative distances between them on such a map are based on recombination frequencies. The markers can be genes or any other identifiable sequences in the DNA. It was also valuable as a framework for organizing more detailed maps of particular regions.
The second stage:
This stage was the physical mapping of the human genome. In a physical map, the distances between markers are expressed by some physical measure, usually the number of base pairs along the DNA. The key is to make fragments that overlap and then use probes or automated nucleotide sequencing of the ends to find the overlap. In this way, fragments can be assigned to a sequencing order that corresponds to their order in a chromosome. In working with large genome, researchers carry out several rounds of DNA cutting, cloning, and physical mapping. After such long fragments are put in order, each fragment is cut into smaller pieces, which are cloned in plasmids or phages, ordered in turn, and finally sequenced.
The last stage:
The ultimate goal in a mapping a genome is to determine the complete nucleotide sequence of each chromosome. For the human genome, this was accomplished by sequence machines, using chain-termination method.
Sequence Alignment Programs: Geneious
[edit | edit source]There are may programs that are used to align sequences that have already been processed by sequencing companies. The most accredited sequencing program is Geneious. Geneious is a program that is a suite of cross platforms for bioinformatics, applications involved in sequence alignment, and sequence BLAST searches in correspondence with NCBI. Geneious comes with many features that involve everything from split viewer genome browsing for easy restriction analysis and cloning workflows to PCR priming design, allowing one to design and test degenerate primers capable of mismatching multiple primers in order to search for implementable DNA sequencing.
References
[edit | edit source]- ↑ "Clustal FAQ #Symbols". Clustal. Retrieved 8 December 2014.
- ↑ Berg, Jeremy M. John L. Tymoczko. Lubert Stryer. Biochemistry Sixth Edition. W.H. Freeman and Company. New York, 2007.
1. Berg, Jeremy M. John L.,Tymoczko, and Lubert Stryer. Biochemistry Sixth Edition. W.H. Freeman and Company. New York, 2007.
2. Coleman,Aaron Gould Meredith Stephano Luis Jose. Biochemical Techniques. University of California, San Diego. 2009
3. “Genomes and their evolution.” Biology. Campbell and Reece. Ed 8th. 2007.500-600. Cases where sequence alignments result in less than 25% of identical amino acids usually rule out homology. However, even if the sequences do not demonstrate homology, comparisons of their structures may. A protein’s three-dimensional structure is more related to its function than is its primary sequence. As a result, the tertiary structure is generally more conserved than the sequence. Though proteins in a family group may differ in amino acid sequence, they are characterized by similar structure. An example of this can be seen in the proteins hemoglobin and myoglobin. Though the two proteins differ in sequence, there are both characterized by a similar heme group that allows for the binding and transport of oxygen.
Paralogs
[edit | edit source]It may also be the case that proteins have structural similarities, but play different biochemical roles and have different amino acid sequences. This is indicative of a common ancestor that evolved into different pathways through what is termed divergent evolution. The proteins are then considered paralogs, which often have different functions within one species. An example of this can be seen in actin and Hsp-70. Though both resemble each other in structure, actin plays a role in muscle contraction and cell motility while Hsp-70 plays a role in preventing stress and heat shock. Another easier example is between our hands and the legs of frogs. Same ancestor and same origin, but came to divert and have different function.

Evaluating Sequence Alignments
[edit | edit source]The method of sequence alignment can be used to detect the similarity of homology. Two sequences of proteins can be compared by sliding one sequence past the other and record the number of matched residues.
Upon analysis of similar structures, the sequence alignments can be re-evaluated for a better fit. Gaps can be inserted to compensate for amino acids that have been inserted or deleted through evolution in order to obtain a better sequence alignment score between structurally similar proteins. Despite overall sequence differences that can be observed between proteins, those in the same family generally contain regions that play crucial roles in function and are therefore more strongly conserved. From this, a sequence template can be created which maps out the conserved amino acid residues that are critical to structure and function.The method of scoring include penalties for gaps to prevent the insertion of an unreasonable number of gaps.
By using substitution matrices, we can detect distant evolutionary relationships. Some substitutions are structurally conservative substitutions. They have similar size and chemical properties. They do not have major effects on protein structure. Other substitutions are amino acids that are not similar to each other. In the substitution matrices, "a large positive score corresponds to a substitution that occurs relatively frequently, whereas a large negative score corresponds to a substitution that occurs only rarely" (Berg 169).
Also, we can use database to identify evolutionary relationships. For example, the National Center of Biotechnology Information (www. ncbi.nih.gov)
Detecting Repeats
[edit | edit source]In some cases a gene may be duplicated and developed into a new gene after mutations are accumulated. However, because the new gene originated from the original gene, it still retains similar domains. Similar domains can be detected by sequence and structural alignments of the repeats by aligning a protein with itself.
Convergent Evolution
[edit | edit source]
Some proteins may contain similar structures but originate from different ancestors. This is known as convergent evolution, where separate proteins evolve through separate pathways and converge on a similar structure and function. In this case, proteins may share similar structures/sequences in regions that prove significant to their function. These proteins often have similar binding sites/active sites. Homology is ruled out due to differences in the overall structures. An example of convergent evolution between our eyes and the eyes of flies. The eyes of flies are from a totally different origin, but has the same function as our eyes.
RNA Sequences
[edit | edit source]Structural alignments of homologous RNA sequences can dictate its structure. The comparison of RNA can be a source of studying evolutionary relationships. Though two RNA strands may differ in its base sequences, a similar structure may be obtained through the conservation of base pairing. Though different bases may be involved, base pairing still occurs in the same general locations allowing for similar secondary structures. An interesting look at a paper by Hitomi Hasegawa and Liisa Holm:
The advances and pitfalls of protein structural alignment
Small Introduction
[edit | edit source]It is apparent that structural comparison can open a window into the past as proteins taken part in a sort of evolution process. Strictly using protein sequence comparison has provided limited information and sometimes generate contradictory information for even slightly distant structures. This has opened the eyes of scientists and researchers to engage in structure comparison methods that allow for more flexible procedures in terms of generating the most biologically meaningful alignments.It is indeed common knowledge that analysis of protein sequences and structures have paved the way for a higher understanding of proteins and their important functions. However, what is important to note is the fact that the variety of protein structures is far less than that of protein sequences, which is mainly due to physical limitations of natural proteins. The foundation for structural proteins has been placed by the concept of visual analysis through the illustration of known structures of proteins. Though there may not be a universal code as to when a protein is structurally similar, we all do know when we see it. The generation of three-dimensional structures has made visualizing proteins far easier. As a matter of fact, the most widespread use of structural alignments is the ability to identify homologous residues that are encoded by the same codon in the genome of a common ancestor. The field of protein structural alignment has maintained its active nature through the years and the number of new methods has actually doubled every five years.
Advances and Technological Development
[edit | edit source]In terms of scores, different scoring themes can be classified into different types depending on whether one is dealing with a three-dimensional, two-dimensional, or one-dimensional structure. When it comes to 3D structures, similarities are drawn from "positional deviations of equivalent atoms upon rigid-body superimposition." The balance between the size of the common core and gap penalties can define sets of optimal configurations. Flexible aligners come into play and serve as the chief identifiers of proteins with large conformational changes by chaining together a series of substructures within the particular protein. When 2D structures are compared, the similarities are drawn from the relative distance differences of "intramolecular C(alpha)-C(alpha) distances." In words we can understand, this essentially means that for the same level of similarity, larger deviations are allowed for tertiary contacts than local ones. Lastly, when it comes to 1D structures, profiles classify each residue according to its amino acid type and highly specific backbone conformational state. Once a score has been identified, the alignment is then determined by finding optimal sets of correspondences. Fragment assembly algorithms generate nonsequential alignments while consistent scoring has been utilized in the generation of multiple alignments from vast databases of pairwise alignments. New structures have been placed in a Protein Data Bank and there are certain parameters such as size and molecular shape of proteins that are initially considered before insertion into this data bank. Evaluations of structural alignments play a pivotal role, as they first measure the accuracy of the alignments, the ability of the alignment score to differentiate homologous from unrelated proteins in database-wide comparisons, and finally, the 'quality' of alignments.
Pitfalls of protein structural alignment
[edit | edit source]A few empirically parameterized models of structural evolution have been previously proposed, but most structural aligners are based on "ad hoc" scores of structural similarity. However, it has been proven that some "ad hoc" scores have worked. In terms of evaluations of structural alignments, the problem with reference-independent evaluation is that it is simply a test of the similarity between scoring functions, rather than the observance of actual rotations that promote alignment optimization. It is also crucial to note that not all programs using the same type of score generated similar alignments. This results in the necessity for developers to pay special attention to the robustness of optimization protocols. Sure enough, despite the few pitfalls of the structural alignment process, the respective models' versatility does advance the study of interplay of sequence and structure evolution in the future.
Reference
[edit | edit source]Curr Opin Struct Biol. 2009 Jun;19(3):341-8. Epub 2009 May 27. Advances and pitfalls of protein structural alignment. Hasegawa H, Holm L.
Programs Used for Structural Alignment
[edit | edit source]
Although somewhat complex, the programs and methods used in structural alignment are quite interesting and hold a wealth of information to be learned. Most of the programs involve matrices and seemingly complex mathematical procedures. Although this holds the difficulty of complex mathematics, it is interesting to see how the structural alignment of proteins is determined and also what each method specifically finds out.
The goal of structural alignment techniques has been to compare individual structures, sets of structures, or an "all-to-all" comparison database that measures the divergence between every pair of structures present in the Protein Data Bank (PDB). The worldwide protein data bank can be found here at this website. These databases generally classify proteins based on their folding.
Certain methods differ in the number of points that are given to each correct protein alignment and the number of points deducted from each incorrect protein alignment. For instance, Glutamine and Asparagine are both polar and have a very similar hydropathy index, so, if Glutamine is present where an Asparagine should be, less points would be deducted than if, say, Valine were there. This type of method allows the maximum number of points to be granted for alignments which change the structure or function of the protein the least. The points granted for each alignment can be compared to other alignments in order to better understand how closely related certain proteins are to one another in structure or function.
VMD
[edit | edit source]A way to actually compare two structures is to use VMD (http://www.ks.uiuc.edu/Research/vmd/). VMD stands for Virtual Molecular Dynamics. One can load a pdb file using VMD and go to file->add a structure. VMD uses RMSD structural alignment to compare two structures. RMSD stands for Root Mean Square Distance and it compares the distances of the atoms. Lower the RMSD values of two proteins, the more aligned.
DALI
[edit | edit source]DALI involves the breaking of the input protein structure into hexapeptide fragments which are then inputted in a distance matrix that evaluates contact patterns between successive fragments. The DALI method has been used to determine structural neighbors and fold classification.
Combinatorial Extension
[edit | edit source]Combinatorial Extension (CE) breaks each protein into a series of fragments and attempts to reassemble them into a complete alignment. This method can be used for structural superpositions, inter-residue distances, secondary structure, solvent exposure, hydrogen-bonding patterns, and dihedral angles.
SSAP
[edit | edit source]SSAP uses double dynamic programming to produce a structural alignment based on atom-to-atom vectors in structure space. In the first step, SSAP will perform inter-residue distance vectors and adjacent non-related neighboring proteins. The dynamic programming on each matrix produce local alignments that are then recorded onto a summary matrix to determine the overall structural alignment. A SSAP score ranging 80-100 explain highly similar structures whereas scores falling between 70-80 are slightly similar with few deviations. Scores 60-70 may contain the same tertiary structure, but the class may vary.
- Beiber*
The *Beiber* method is a combinatorial algorithm for non-sequential structural alignment of proteins and similarity search in databases. This method focuses on secondary structure to evaluate similarities between two different protein structures based on contact maps.
MAMMOTH
[edit | edit source]MAMMOTH's purpose was originally developed for comparing models coming from structure prediction, but now also works well with experimental models. MAMMOTH has been used to create a large database covering predicted structures of unknown proteins for 150 genomes which allows for genomic scale normalization.
RAPIDO
[edit | edit source]RAPIDO is a web based program for analyzing three dimensional crystal structures of different protein molecules in the presence of conformational changes. This method involves the calculation of difference distance matrices between fragments that are structurally similar in two different proteins.
SABERTOOTH
[edit | edit source]SABERTOOTH uses structural profiles to perform structural alignments. This tool recognizes structural similarities with accuracy and quality compared to that of other established alignment tools based on coordinates.
BLOSUM
[edit | edit source]BLOSUM stands for Blocks of Amino Acid Substitution Matrix uses an assigned score based on the observed frequencies of such occurrences in alignment related proteins. Certain scores may be added with values of either positive or negative value. This scale is then run by a log odd ratio. In essence, two matrices are compared and evaluated by the ratio of similar or identical sequences to the ratio of unknowns missed by one.
TOPOFIT
[edit | edit source]TOPOFIT analyzes protein structures based on three dimensional Delaunay triangulation patterns derived from backbone representation. TOPOFIT produces a structural alignment of proteins based on the fact that proteins have a common spatial invariant part (a set of tetrahedrons) which is mathematically described as a common spatial sub-graph volume of three dimensional contact graph derived from Delaunay tessellation (DT).
InsightII
[edit | edit source]insightII is a molecular modeling package developed by Biosym. The programs included include Insight II, BioPolymer, Analysis, and Discover. InsightII therefore is a comprehensive program that can not only build any class of molecule or molecule system, but with the molecular mechanics program Discover, can manipulate these same molecules.
Insight II is primarily used for visualization. It creates, modifies, manipulates, displays, and analyzes molecular systems. Insight II essentially provides the core requirements for all software modules. Analysis revolves around mathematical and geometric modeling of molecular properties. Molecular properties are defined interactively, evaluated dynamically, and visualized interactively through spreadsheets, graphs, and graphic representations. BioPolymer constructs models of polymers—peptides, proteins, carbohydrates, and nucleic acids—for visualing complex structures and use in further simulation work. CHARMM is a simulation program available within insight II that uses energy functions to describe the forces on atoms in molecules. This allows for calculation of interaction and conformational energies, free energy, and vibrational frequencies.
Through use of the Discover program, one can optimize the structure of the molecule or protein being viewed. This is due to the fact that it incorporates a range of well validated forcefields for dynamics simulations, minimization, and conformational searches. This allows for the ability to extrapolate the structure, energetics, and properties of systems, be they organic, inorganic, organometallic, or biological. Because of this program, it is possible to take the sequence of a protein and from that extrapolate a rudimentary structure from it. Discover also implements Inter Process Communications which allow for Discover turn over control to external programs and retrieve those results, inforperating them into continuing Discover computations.
DALI
[edit | edit source]DALI stands for Distance Alignment Matrix Method. DALI is a common and popular method that breaks down the protein that is inputted into hexapeptide fragments and then calculates a distance matrix through the understanding of the contact pattern between successive fragments. Secondary structure features with residues that are contiguous in sequence are shown on the matrix's main diagonal. The other diagonals represent residues that are not next to each other in sequence. When the diagonals are parallel to the main diagonal, the features they represent are also parallel. When the diagonals are perpendicular, however, their features are anti parallel. If two proteins' distance matrices are the same or share similar features in almost the same positions, they can be said to have similar folds and length loops connecting the secondary structure elements. When two matrices are built, DALI's actual alignment process involves conducting a series of overlapping sub matrices that are 6x6. These 6x6 matrices are then reassembled to a final alignment through a standard score-maximization algorithm. The DALI method has been used to construct the FSSP database (Fold classification based on structure-structure alignments of proteins, or families of structurally similar proteins). This database holds all known protein structures and aligns them with each other to determine structural neighbors and fold classification. This database information can easily be found on the internet for ease of access. A DALI online database can found here. A downloadable program and web search are other available DALI resources.
Combinatorial Extension
[edit | edit source]The combinatorial extension (CE) method is like the DALI method in that it breaks each structure into a series of fragments and attempts to reassemble them into a complete alignment. Pairwise combinations of fragments, named aligned fragment pairs, or AFPs, are used to make a similarity matrix where an optimal path is generated to identify the final alignment. To reduce the amount of searching and increase efficiency, only AFPs that meet a criteria for local similarity are used in the matrix. The first CE method only included structural superpositions and inter-residue distances but has been grown to include local environmental properties like secondary structure, solvent exposure, hydrogen-bonding patterns, and dihedral angles. An alignment path is the optimal path that is calculated through a similarity matrix by linear progression through the sequences and extending the alignment with a possible high-scoring AFP pair. The initial AFP pair that nucleates the alignment can occur at any place in the sequence matrix. This is then extended with AFPs that meet the given distance criteria which restricts the alignment to low gap sizes. The size of each AFP and maximum gap size are required input parameters that are usually set to the empirically determined values of 8 and 30, respectively. An all-to-all fold classification database from known protein structures in the PDB has been constructed using CE.
SSAP
[edit | edit source]
SSAP stands for Sequential Structure Alignment Program. SSAP method uses double dynamic programming to produce structural alignment based on atom to atom vectors in structure space. SSAP constructs the vectors using the beta carbons for all residues by glycine, instead of the alpha carbons typically used, which thus takes into account the rotameric state of each residue and its location along the backbone. SSAP first constructs a series of inter-residue distance vectors between each residue and its nearest non-contiguous neighbors on each protein. A series of matrices is then constructed containing the vector differences between neighbors for each pair of residues for which the vectors were constructed. Using dynamic programming on each resulting matrix determines a series of optimal local alignments, which are then mixed together into a "summary" matrix to which dynamic programming is again used to determine overall structural alignment. SSAP originally gave only pairwise alignments, but can now provide multiple alignments. It has been used to produce a hierarchical fold classification known as CATH (Class, Architecture, Topology, Homology) through an all-to-all comparison of proteins. This has then been used to put together a CATH protein structure classification database. The constructed database can found here.
GANGSTA+
[edit | edit source]GANGSTA+ is a combinatorial algorithm for non-sequential structural alignment of proteins and similarity search in databases. This method uses a combinatorial approach on the secondary structural level to find similarities between two protein structures based on contact maps. Different SSE (Secondary Structure Element) assignment modes can be used for this program. This assignment of SSEs can be performed with respect to the sequential order of SSEs in the polypeptide chain of the considered protein pair (sequential alignment), or by ignoring this order completely (non-sequential alignment). SSE pairs can also be aligned in reverse orientation if desired. The highest ranking SSE assignments are transferred to the residue level by a point matching approach. In order to obtain an initial common set of atomic coordinates for both proteins, pairwise attractive interactions of the alpha carbon atom pairs are defined by inverse Lorentzians and the energy is minimized. The GANGSTA+ database can be found here.
MAMMOTH Introduction
[edit | edit source]MAMMOTH stands for MAtching Molecular Models Obtained from THeory. This program was originally developed for comparing models produced from structure prediction since it is tolerant of large unalignable regions. However, it has proven to work well with experimental models, especially when looking for remote homology. Benchmarks on targets of blind structure prediction and automated GO annotation have shown it to tightly rank correlated with human curated annotation. A largely complete database of MAMMOTH based structure annotation for the predicted structures of unknown proteins covering 150 genomes facilitates genomic scale normalization.
MAMMOTH Method
[edit | edit source]MAMMOTH based structure alignment methods decompose the protein structure into short peptides (heptapeptides for example) which are compared with the same length peptides of another protein. Similarity scores between the two polypeptides is then calculated using a unit-vector RMS (URMS) method. These cores are stored in a similarity matrix, and with a hybrid (local-global) dynamic programming, the optimal residue alignment is calculated. Protein similarity scores calculated with MAMMOTH are derived from the likelihood of obtaining a given structural alignment by chance.
MAMMOTH-mult
[edit | edit source]MAMMOTH-mult is an extension of the MAMMOTH algorithm used to align related families of protein structures. This algorithm is very quick and produces consistent high quality structural alignments. Multiple structural alignments calculated with MAMMOTH-mult produce structurally implied sequence alignments that can be further used for multiple-template homology modeling, HMM-based (Hidden Markov model) protein structure prediction, and profile-type PSI-BLAST searches.
RAPIDO
[edit | edit source]RAPIDO stands for Rapid Alignment of Proteins in terms of DOmains. RAPIDO is a web sever for 3D alignment of crystal structures of different protein molecules in the presence of conformational changes. Similar to CE's first step, RAPIDO identifies fragments that are structurally similar in two different proteins using an approach that uses difference distance matrices. The MFPs (Matching Fragments Pairs) are represented as nodes in a graph which are chained together to form an alignment by means of an algorithm for identification of the longest path on a DAG (Directed Acyclic Graph). The last step of refinement is performed to improve quality of the performed alignment. After aligning the two structures, the server applies a genetic algorithm for identification of conformationally invariant regions. These regions correspond to groups of atoms with constant inter atomic distances (within a defined tolerance). By doing this, RAPIDO takes into account variation in the reliability of atomic coordinates by employing weighting-functions based on the refined B-values. The regions that are identified as conformationally invariant by RAPIDO represent reliable sets of atoms for the superposition of the two structures that can be used for detailed analysis of changes in the conformation. RAPIDO can identify structurally equivalent regions even when these consist of fragments that are distant in terms of sequence and separated by other movable domains.
SABERTOOTH
[edit | edit source]SABERTOOTH uses structural profiles to perform structural alignments. The underlying structural profiles express the global connectivity of each residue. Although the vectorial representation is extremely condensed, this program recognizes the structural similarities with accuracy and quality comparable to established alignment tools based on coordinates. Also, the algorithm has favorable scaling of computation time with chain length. Since the algorithm is independent of the details of the structural representation, the framework can be generalized to sequence-to-sequence and sequence-to-structure comparisons within the same setup, and is therefore more general than other tools. SABERTOOTH can be accessed online via this website.
TOPOFIT
[edit | edit source]The TOPOFIT method analyzes protein structures using three-dimensional Delaunay triangulation patterns derived from backbone representation. Structural related proteins have been found to have a common spatial invariant part, a set of tetrahedrons, mathematically described as a common spatial sub-graph volume of the three-dimensional contact graph derived from Delaunay tessellation (DT). Due to this property of protein structures, a novel common volume superimposition (TOPOFIT) method is presented to produce structural alignments of proteins. The superimposition of the DT patterns allows the common number of equivalent residues to be objectively identified. In other words, TOPOFIT identifies a feature point on the RMSD/Ne curve, a topomax point, until which two structures correspond to each other including backbone and inter-residue contacts, while the growing number of mismatches between the DT patterns occurs at larger RMSD (Ne) after the topomax point. The topomax point is present in all alignments from different protein structural classes. Therefore, the TOPOFIT method identifies common, invariant structural parts between proteins. The TOPOFIT method introduces new opportunities for the comparative analysis of protein structures and for more detailed studies on understanding the molecular principles of tertiary structure organization and functionality. This helps to detect conformational changes, topological differences in variable parts, which are particularly important for studies of variations in active/binding sites and protein classification.
RNA Structural Alignment
[edit | edit source]Structural alignment techniques have traditionally only been applied to proteins, as they are the primary biological macromolecules that form characteristic three dimensional structures. However, large RNA molecules also form characteristic tertiary structures, which mostly consist of hydrogen bonds formed between base pairs as well as by base stacking. Functionally similar noncoding RNA molecules can be particularly difficult to extract from genomics data because structure is more strongly conserved than sequence in RNA as well as proteins, while the more limited alphabet of RNA decreases the information content of any given nucleotide at any given position. A recent method for pairwise structural alignment of RNA sequences with low sequence identity has been published and implemented in the program FOLDALIGN. This method is not truly analogous to protein structural alignment techniques because it computationally predicts the structure of the RNA input sequences rather than requiring experimentally determined structures as input. Although computational prediction of the protein folding process has not been particularly successful to date, RNA structures without pseudoknots can often be sensibly predicted using free energy-based scoring methods that account for base pairing and stacking.
History
[edit | edit source]Early signs of branching evolutionary trees or phylogenetic trees are paleontological charts. This kind of chart was illustrated in Edward Hitchcock's book called the Elementary Geology, which showed the geological relationships between that of plants and animals. However, going way back in time, the whole idea of tree life first started from the ancient notions of a ladder-like progression from the lower to the higher forms of life. An example of a ladder-like progression would be that of the Great Chain of Being.
In addition, a well-known man named Charles Darwin from the 1850s produced one of the first drawings of evolutionary tree in his seminal book called "The Origin of Species". Basically in this book, he showed the importance of evolutionary trees. After many years later, many evolutionary biologists studied the forms of life through the use of tree diagrams to depict evolution. The reason for this is that these types of diagrams prove to be very effective at explaining the concept of how speciation happens through the random splitting of lineages and the idea of adaptive. Overall, for many centuries, many biologists used the tool evolutionary trees as a way to study the idea of life.
Phylogeny
[edit | edit source]
Evolutionary trees, or Phylogeny, is the formal study of organisms and their evolutionary history with respect to each other. Phylogenetic trees are most commonly used to depict the relationships that exist between species. In particular, they clarify whether certain traits are homologous (found in the common ancestor as a result of divergent evolution) or homoplasy (or sometimes referred to as analogous, a character that is not found in a common ancestor but whose function developed independently in two or more organisms, known as convergent evolution). Evolution Trees are diagrams that show various biological species and their evolutionary relationships. They consist of branches that flow from lower forms of life to the higher forms of life.
Evolutionary trees differ from taxonomy. Whereas taxonomy is an ordered division of organisms into categories based on a set of characteristics used to assess similarities and differences, evolutionary trees involve biological classification and use morphology to show relationships. Phylogeny is shown by evolutionary history using the relationships found by comparing polymeric molecules such as RNA, DNA, or protein of various organisms. The evolutionary pathway is analyzed by the sequence similarity of these polymeric molecules. This is based on the assumption that the similarities of sequence result from having fewer evolutionary divergence than others. The evolutionary tree is constructed by aligning the sequences. The length of the branch is proportional to the amount of amino acid differences between the sequences.
Phylogenetic systematics informs the construction of phylogenetic trees based on shared characters. Comparing nucleic acids or other molecules to infer relationships is a valuable tool for tracing an organism's evolutionary history. The ability of molecular trees to encompass both short and long periods of the time is hinges on the ability of genes to evolve at different rates, even in the same evolutionary lineage. For example, the DNA that codes for rRNA changes relatively slowly, so comparisons of DNA sequences in these genes are useful for investigating relationships between taxa that diverged a long time ago. Interestingly, 99% of the genes in humans and mice are detectably orthologous, and 50% of our genes are orthologous with those of yeast. The hemoglobin B genes in humans and in mice are orthologous. These genes serve similar functions, but their sequences have diverged since the time that humans and mice had a common ancestor.
Evolutionary pathways relating the members of a family of proteins may be deduced by examination of sequence similarity. This approach is based on the notion that sequences that are more similar to one another have had less evolutionary time to diverge than have sequences that are less similar.
Evolutionary trees are used today for DNA hybridization. They are used to determine the percentage difference of genetic material between two similar species. If there is a high resemblance of DNA between the two species, then the species are closely related. If only a small percentage is identical, then they are distant in relationship.
Construction of a Evolutionary Tree
[edit | edit source]Each point at which a line in a evolutionary tree branches off is known as a node. A node is a common ancestor to the species that come off that branch. Relationships between species in an evolutionary tree include monophyletic, paraphyletic, and polyphyletic. A monophyletic group is a branch of species which contain a common ancestor and all descendants. Paraphyletic groups consist of a common ancestor but not all its descendants. Polyphyletic groups consist of organisms that do not have a (recent) common ancestor and are usually compared to study similar characters among relatively unrelated organisms.
These nodes are calculated by using a computational phylogenetic program that calculates the genetic distance from multiple sequence alignments. However, there are limitations, primarily not being able to account the actual evolutionary history. While the sequence alignment shows comparatively how related two species are, there is no indication as to how they evolved. Therefore for these the origins of the three domains came from the same ancestor and then branches out to the two distinct groups Eukarya and Prokarya. However, the Archaea branches out of the Eukarya domain, even though they are single-celled.

Evidence for Phylogeny construction: 1. By looking at the fossil record. The problem is that the record is incomplete and only hard structures were preserved. 2. By studying recent species, looking at the shared characters, both homologous and analogous characters, give evidence of evolutionary past.

Types of Evolutionary Tree
[edit | edit source]There are many different types of evolutionary trees and each one represents something different. One type of evolutionary tree is called the rooted phylogenetic tree. This tree is a direct type of tree that contains a special node corresponding to the most recent common ancestor of all the entities at the leaves of the tree. The use of an uncontroversial outgroup is one of the most common techniques used for rooting trees. In other words, rooted trees typically illustrate the sequence or trait data of a particular outgroup.
Another type of evolutionary tree is called the unrooted tree. Unrooted trees often illustrate the relationship between different leaf nodes. This is done without making any assumptions about the ancestry. Unlike that of the rooted trees where a sign of ancestry identify is needed, unrooted trees can always be created from rooted ones by omitting the root. Basically, an unrooted tree is generated by introducing assumptions about the relative rates of evolution on each branch or including an outgroup in the input data. An application that is often used in this process is called the molecular clock hypothesis.
Last but not least, bifurcating tree is also a type of evolutionary tree. In bifurcating tree, both rooted and unrooted phylogenetic trees can be multifurcating or bifurcating, and can be shown as labeled or unlabeled. For example, a rooted tree that has been multifurcated may have more than two children at the nodes, unlike that of the unrooted multifurcating tree where more than three neighbors can appear at the nodes.
Furthermore a rooted tree that has been bifurcated has exactly two descendants arising from each interior node that typically forms a binary tree. On the other hand, an unrooted bifurcating tree is similar to that of an unrooted binary tree. An unrooted binary tree often has a free tree with exactly three neighbors at each internal node. In terms of labeled and unlabeled trees, labeled trees has unique values assigned to its leaves, while an unlabeled tree also known as a tree shape is define as a topology only.
Overall, the number of possible trees for a given number of leaf nodes depends on the specific type of tree one is looking at. However, there are always less bifurcating trees than multifurcating trees. Similarly, there are less unlabeled than labeled trees and less unrooted than rooted trees. In terms of labeling the tree, it is always important to know that the letter "n" represents the number of leaf nodes.
Sequence Alignment in Evolutionary Trees
[edit | edit source]Evolutionary trees can be made by the determination of sequence information of similar genes in different organisms. Sequences that are similar to each other frequently are considered to have less time to diverge. Whereas, less similar sequences have more evolutionary time to diverge. The evolutionary tree is created by aligning sequences and having each branch length proportional to the amino acid differences of the sequences. Furthermore, by assigning a constant mutation rate to a sequence and performing a sequence alignment, it is possible to calculate the approximate time when the sequence of interest diverged into monophyletic groups.
DNA can be amplified and sequences with the development of PCR methods. Mitochondrial DNA from a Neanderthal fossil was identified to contain 379 bases of sequence. When compared to the Homo sapiens, only 22 to 36 substitutions in the sequence was found as opposed to 55 differences between homo sapiens and chimpanzees over common base in the same region. Further analysis suggests that the Homo sapiens and Neanderthals shared common ancestry 600,000 years ago. This reveals that Neanderthals were not intermediate between chimpanzees and humans, but rather an evolutionary dead end, which became extinct.
Sequence alignments can be performed on a variety of sequences. For constructing an evolutionary tree for proteins, for example, the sequences are aligned and then compared via likeness to construct a tree. In the globin tree above, it is then possible to see which protein diverged first. Another example typically uses rRNA(ribosomal RNA) to compare organisms, since rRNA has a slower mutation rate, and is a better source for evolutionary tree construction.
This is best supported by Dr. Carl Woese's study that was conducted in the late 1970s. Since the ribosomes were critical to the function of how living things operate, they are not easily changed through the process of evolution. Significant changes could allow the ribosomes to not do its role, therefore having the gene sequence of it is conserved. Taking advantage of this, Dr. Woese compared the minuscule differences in the sequences of ribosomes amongst a great array of bacteria and showed how they were not all related. Looking at extreme bacteria such as methanogens, was not able to connect them to eukaryote or proselytes because they fell within their own category of archaea.

Example of Phylogeny Tree for the Domain Eukarya
[edit | edit source]Constructing the phylogeny tree requires systematists to search for synapomorphies (shared derived characters) and symplesiomorphies (shared ancestral characters) characteristics.
Three Domain
Reading the Phylogeny tree: The numbers in the diagram indicate the synapomorphies or shared derived characters unique only for those group or groups . The diagram shows that there are three domains: Bacteria, Archaea, and Eukarya. The domain Eukarya and Archaea possess (1) have introns, histones, and RNA polymerase similar to eukaryotic RNA polymerase. Furthermore, the domain Archaea possesses (2) unique lipid content in membranes and unique cell wall composition.
Domain Eukarya Four supergroups phylogeny The synapomorphies for Eukarya are (5) have a nucleus, membrane-bound organelles, sterols in their membrane, cytoskeleton, linear DNA with genomes consisting of several molecules, and a 9+2 microtubular ultrastructure flagella. Based on DNA sequence data, there are four Supergroups to this domain: Excavata, Chromalveolata, Unikonta, Archaeplastida (red algae, green algae, plants).
Supergroup Archaeplastia has (7) chloroplasts by primary endosymbiosis. Major clades: Rhodophyta (red algae), Chlorophyta (green algae), Plantae (land plants). Archaeplastida phylogeny
Supergroup Excavata: Excavata There are three phyla for this Supergroup: Parabasalia, Euglenophyta, and Kinetoplastida. Euglenophyta doesn’t have cell wall, but have (8)flexible pellicle within the cell membrane. It has chlorophyll A amd B as in plants - which it obtained by secondary endosymbiosis, green lineage. Parabasalia possesses (9) a reduced or lost mitochondria. Kinetoplastida has (10) single large mitochondrio/kinetoplast, which edit mRNAs.
Supergroup Chromalveolata:
Chromalveolata phylogeny
There are three clades: alveolata, straminopila, and rhizaria.
Rhizaria's Phylum Foraminifera possesses (11) multichambered shells made of organic material and CaCO3.
Synapomorphies for straminopila and rhizaria are (12) chloroplasts by secondary endosymbiosis, red lineage.
Stramenopila has (13) two unequal flagella, one longer tinseled. There are three major phyla for Stramenopila. Phylum Bacillariophyta (diatom) has (14) cell walls of hydrated silica in organic matrix, made up of two halves: “box and lid.” Phylum Phaeophyta (Brown algae)has (16) multicellular sea weeds. Phylum Oomycetes (water molds, downy mldews) has (15) a loss of chloroplasts.
Alveolata has (17) a membrane-bound sac under plasma membrane. There are three phyla. Phylum Dinoflagelleta possesses (19) plates of cellulose-like material, grooved. Phylum Ciliophora has (20) two types of functionally different nuclei: macronucleus (controls metabolism) and micronucleus (function in sexual reproduction). Phylum Apicomplexa has (18) apical structure for penetrating host cells.
Supergroup Unikonta has (6) a triple pyrimidine biosynthesis fusion gene and has one flagellum
Unikonta phylogeny Three major clades: Amoebozoans, Fungi, Animals.
Amoebozoa has (21) broad pseudopodia. It's Phylum Gymnamoeba (22) feed and move by lobed pseudopodia.
Opisthokonta (Fungi and Animalia) possess (23) flagellum posterior. Fungi has (24) cell walls made of chitin, absorptive heterotrophy, and are multicellular. There are four Phyla of Fungi. Asides from Phylum Chytridiomyota (water molds), all other Fungi Phyla has a (25) loss of flagellum and time separation between plasmogamy and karyogamy. Phylum Zygomycota has (26) heterokaryotic state of reproduction limited to zygosporangium. Both Basidiomycota and Ascomycota possess (27) conidia, extensive (n+n) state, size and duration, septate hyphae, and macroscopic fruiting bodies. Phylum Basidiomycota has (28) long-lived dikaryotic mycelium in dikaryotic state, meiospores produced in special cell called basidium, and are sex predominant (asexual spores rare). Phylum Ascomycota has (29) meiospores produced in special cell called ascus.
Kingdom Animalia are (30) multicellular, possess extracellular matric with collagen, proteoglycans, and special types of junctions between cells (cell adhension proteins).
Phylum Porifera (sponges) has (31) spicules, internal aquiferous system. Subkingdom Eumetazoa has (32) body symmetry, primary germ layer (true endoderm and ectoderm), true tissues and organs, epithelial tissue, and nervous tissue. Radiata has (33) primary radial symmetry. Phylum Cnidaria has (34) a mesoglea, and a cnidocytes (with nematocysts). Bilateria has (35) bilateral symmetry, body cavity (coelom), mesoderm (triploblastic), and muscle. Two mahor phylogenetic branches are Protostomia and Deuterostomia. Protostomia has (36)a schizocoelous, and the blastopore becomes its mouth. Deuterostomia has (37) entercoelous, indeterminate cell fate, radial cleavage, and the blastopore becomes the anus. Phylum Echinodermata has (38) a water vascular system, tube feet, and has radial symmetry in adults (bilateral larvae). Hemichordata has (39) pharyngeal slits at some state of life. Phylum Chordata has (40) muscular post-anal tail, dorsal follow nerve cord, and notochord.
Phylogeny tree for Protostomia. Protostomia phylogeny
Phylogeny tree for Chordata. Chordata phylogeny
References
[edit | edit source]Waggoner. Ben. "Archaea: Systematics." 15 Nov. 2009. <http://www.ucmp.berkeley.edu/archaea/archaeasy.html> http://en.wikipedia.org/wiki/Phylogenetic_tree
Light microscopy
[edit | edit source]In light microscopy, glass lenses are used to focus a beam of light on to the specimen under investigation. The light passing through the specimen is then focused by other lenses to produce a magnified image. Standard (bright-field) light microscopy is the most common microscopy technique in use today and uses a compound microscope. The specimen is illuminated from underneath by a lamp in the base of the microscope, with the light being focused on to the plane of the specimen by a condenser lens. Incident light coming through the specimen is picked up by the objective lens and focused on to its focal plane, creating a magnified image. This image is further magnified by the eyepiece, with the total magnification achieved being the sum of the magnifications of the individual lenses. In order to increase the resolution achieved by a compound microscope, the specimen is often overlaid with immersion oil into which the objective lens is placed. The limit of resolution of the light microscope using visible light is approximately 0.2*10-12m.
Fluorescence microscopy
[edit | edit source]In fluorescence microscopy, the light microscope is adapted to detect the light emitted by a fluorescent compound that is used to stain selectively components within the cell. A chemical is said to be fluorescent if it absorbs light at one wavelength (the excitation wavelength) and then emits light at a longer wave length (the emission wavelength). Two commonly used compounds in fluorescent microscopy are rhodamine and Texas red, which emit red light, and fluorescein, which emits green light. First, an antibody against the antigen of interest (so-called primary antibody) is added to the specimen. A fluorescent compound is chemically coupled to a secondary antibody that recognized the primary antibody. Then the fluorescently-tagged secondary antibody is added to the tissue section or permeabilized cell, and the specimen is illuminated with light at the exciting wavelength. The structures in the specimen to which the antibody has bound can then be visualized. Fluorescence microscopy can also be applied to living cells, which allows the movement of the cells and structures within them to be followed with time.
Green fluorescent protein
[edit | edit source]The discovery of a naturally fluorescent protein found in the jellyfish Aquorea victoria. In this 238 amino acid protein, called green fluorescent protein (GFP), certain amino acid side-chains has spontaneously cyclized to form a green-fluorescing chromophore. Using recombinant DNA techniques, the DNA encoding GFP can be tagged on to the DNA sequences encoding other proteins, and then introduced into living cells in culture or into specific cells of a whole animal. Cells containing the introduced gene will then produce the protein tagged with GFP which will fluoresce green under the fluorescent microscope. The localization and movement of the GFP-tagged protein can then be studied in living cells in real time. Multiple variations of GFP have been engineered which emit light at different wavelength. They allows several proteins to be visualized simultaneously in the same cell.
Transmission electron microscopy
[edit | edit source]In contrast with light microscopy where optical lenses focus a beam of light, in electron microscopy electromagnetic lenses focus a beam of electrons. Because electrons are absorbed by atoms in the air, the specimen has to be mounted in a vacuum within an evacuated tube. The resolution of the electron microscope with biological materials is at best 0.10 nm. In transmission electron microscopy, a beam of electron is directed through the specimen and electron magnetic lenses are used to focus the transmitted electrons to produce an image either on a viewing screen or on photographic film. As in standard light microscopy, thin sections of the specimen are viewed. However, for transmission electron microscopy the sections must be much thinner (50-100 nm thick). Since electrons pass uniformly through biological material, unstained specimens give very poor images.Therefore, the specimen must routinely be stained in order to scatter some of the incident electrons which are then not focused by the electromagnetic lenses and so do not form the image. Heavy metals such as gold and osmium are often used to stain biological materials. In particular osmium tetroxide preferentially stains certain cellular components, such as membranes, which appear black in the image. The transmission electron microscope has sufficiently high resolution that it can be used to obtain information about the shapes of purified proteins,viruses and subcellular organelles. Antibodies can be tagged with electron-dense gold particles in a similar way to being tagged with a fluorescent compound in fluorescence microscopy, and then bound to specific target proteins in the thin sections of the specimen. When viewed in the electron microscope, small dark spots due to the gold particles are seen in the image wherever an antibody molecules has bound to its antigen and so the technique can be used to localize specific antigens.
Scanning electron microscopy
[edit | edit source]In scanning electron microscopy, an (unsectioned) specimen is fixed and then coated within a thin layer of a heavy metal such as platinum.An electron beam then scans over the specimen, exciting molecules within it that release secondary electrons. These secondary electrons are focused on to a scintillation detector and the resulting image displayed on a cathode-ray tube. The scanning electron microscope produces a three-dimensional image because the number of secondary electrons produced by any one point on the specimen depends on the angel of the electron beam in relation to the surface of the specimen. The resolution of the scanning electron microscope is 10 nm, some 100-fold less than that of the transmission electron microscope.
Reference
[edit | edit source]David Hanes,Nigel Hooper.Biochemistry. Taylor and Francis Group. New York. 2005. Combinatorial Chemistry represents a mixture of chemistry and biology. It involves the use of computers and technology to create a massive array of different, but structurally related/similar, molecules in order to investigate a specific molecule and its derivatives. In common use, a "virtual library" is first created via computer in order to create a compilation of the different molecules that are to be investigated. From this library will specific molecules be chosen to be synthesized in the lab and further analyzed for desired characteristics. This technique is extremely useful in that it can mass produce the variety of diverse collection of molecules with limited condition variability needed to conduct the mass scale assays required to screen for specific characteristics. You can imitate the process of evolution by creating large sets of molecules and select for a specific function.
Combinatorial chemistry is a method used to synthesize different substances rapidly and at the same time. Compared to those time-consuming methods of traditional chemistry where compounds are synthesized individually one at a time, combinatorial chemistry is a very useful and faster method. Combinatorial Chemistry is used to synthesize large number of chemical compounds by combining sets of building blocks. It synthesizes different substances rapidly and at the same time.
Combinatorial chemistry creates a large sets of molecules and select for a specific function. It requires three steps in process: - the generation of a diverse population - selection of members based on some criterion of fitness - reproduction to enrich the population in these more-fit members
Combinatorial Chemistry has had the largest impact and is mostly used in the pharmaceutical industries such as drug discovery and research, as well as in agrochemical industries. In an area of research in which the optimization of a certain activity or characteristic is the goal, combinatorial chemistry can be used to track down molecules and reactions that will accomplish this task.
Pharmaceutical companies had the need of improving drugs and being able to offer them to the public faster. Usually only 1 of 10 000 synthesized drugs are considered for marketing and the process may take up to 12 years to be settled, meaning millions of dollars spent. Since only few compounds could be synthesized by a scientist per year, a new technique to synthesize drugs faster was needed, and combinatorial chemistry was born.1
Combinatorial chemistry is considered the synthesis of large numbers of molecules reacting together all possible combinations of certain reagents at the same time.2 These combinatorial reactions are done on microtiter plates, which have several short-test-tube-like wells. By making note of exactly which variation of the reagents was placed in each well, one can determine the separate simultaneous reactions that took place to give the products obtained in each cell.1
The execution of combinatorial reactions in these microtiter plates give way to chemical libraries that are screened and analyzed to determine the properties of each of the compounds in it. Computational chemistry then comes into play in order to maintain an organized database of the different characteristics revolving around each of the compounds in the library, such as the structures of the reagents and their quantities, reaction features, the location of the cell in the well, results of tests done, among other information.1
Combinatorial Chemistry Technology
[edit | edit source]At first, combinatorial chemistry was conceived as a technology for synthesizing and characterizing collections of compounds and screening them for useful properties. Its main focus was on the synthesis of peptide and oligonucleotide libraries. Then after a while, the focus of field changed to synthesis of small drugs as organic compounds, rather than large chemical compounds. Over past years, the combinatorial chemistry has emerged as an exciting new paradigm for the drug discovery for many pharmaceutical companies.
Meanwhile, researchers continue to find ways to improve the capabilities of combinatorial chemistry, with developments such as:
1. A growing trend toward the synthesis of complex natural-product-like libraries
2. An increased focus on "phase trafficking" techniques aimed at integrating synthesis with purification
3. strategies for purification and analysis, like use of supercritical fluid chromatography
4. the application of combinatorial chemistry to new targets, such as nuclear receptors
Even though combinatorial chemistry is more efficient in both cost and time wise, it, similar to traditional drug design, still relies on organic synthesis methods. Yet, the large libraries of compounds used for combinatorial chemistry do not produce active compounds, resulting in the necessary straightforward method to locate the active components. Such method is called combinatorial organic synthesis (COS), systematic and repetitive method that uses chemical building blocks to form numerous chemical compounds. There are three different COS approaches: arrayed, spatially addressable synthesis, encoded mixture synthesis, and deconvolution.
The first approach, arrayed, spatially addressable synthesis, groups chemical building blocks according to their individual positions; thus, allowing active compounds to be identified by their locations. The second strategy, encoded mixture synthesis is used to identify each compound using inert chemical tags such as nucleotides or peptides. The third technique, deconvolution, combinatorially synthesizes numerous compound mixtures in order to pursue the most active combination.
Soft Organic Biomolecules
[edit | edit source]Drug research and design has made significant advancements within the past few decades, and still remains an industry in progress. With many different sub-categories in the world of drug design, one particular branch has received a growing amount of attention: creating disease-specific drug delivery vehicles. Drug delivery devices would be predominantly used to treat diseases such as cancer, because current treatments (chemotherapy) also cause harm to essential and non-pathogenic bodily tissues.
Soft, amphiphilic polymers may hold much utility in creating efficient drug delivery vehicles. The desired characteristics of the polymers are as follows: biodegradable, biocompatible, and high molecular weight. Being so, the widely abundant Poly Lactic Acid polymer serves a viable foundation for modeling the polymers after. In utilizing several known synthetic methodologies, researchers have proposed a way in creating potentially novel amphiphilic biomolecules [5].
Reaction of an aldehyde with a specifically designed isocyanide yields the necessary α-hydroxy acid monomer needed to create a Poly(α-hydroxy acid) polymer [6]. Since any aldehyde can be used, a lot of functionality can be incorporated into the polymer backbone. In particular, a norbornene moiety can be used. Norbornene can participate in a widely used Ring Opening Metathesis Polymerization (ROMP). Using two modes of polymerization in tandem opens up a vast amount of functionalization and drug incorporation, as well as fine-tuned adjustments on amphiphilicity. The picture at the right shows a synthetic scheme in obtaining the bifunctionalized α-hydroxy acid monomer.

Amphiphilic polymers will create micelle aggregates with hydrophilic components exposed in the body’s aqueous environment. Poly Ethylene Glycol can be used for the amphiphilic zone, which will allow the drug-encapsulated aggregates to dissolve and be compatible with a patient’s biochemistry. With disease-specific antibodies incorporated into the polymer’s functionality, the micelle aggregates can seek out the cancer, and inflict a lethal dose of medicine.
RNA Synthesis
[edit | edit source]A useful application of RNA analysis can allow scientists to synthesis specific RNA sequences for intended functions. The process itself is rather simple. First, a very large amount of completely random RNA sequences are synthesized with really no thought to actual use. Then the collection of RNA are eluted down a column that specifically selects for RNA that fits a desired function such as ATP-binding where the column will just be filled with ATP bonded molecules. Naturally, only the RNA that bind to the function will stay in the column and those can be isolated for further testing by just eluting the rest of the column. Through this random selection process, scientists can pretty much create RNA to conduct any function. The following synthesis can be summarized in a few steps:
1. Initially, a randomized RNA pool containing of any variety molecules is present in a given population inserted into an ATP affinity column.
2. A selection process occurs in which only specific molecules are isolated with desired binding or reactivity properties.
3. The remaining RNA population that have remained the isolation process are then intensely amplified through Polymerase Chain Reaction (PCR).
4. Errors that occur within the course of replication are seen as additional variations introduced into this generation of RNA. Errors are induced through reverse transcription of RNA to DNA and back to RNA to increase the likelihood of error and mutation. The new population would consequently be introduced back into the column for further ATP analysis. Eventually structures that emerge out of the tube are assumed to be plausible structures of what the RNA could have existed in the past.

Amplified Ancient DNA
[edit | edit source]The use of PCR to amplify DNA has been used to sequence the DNA of ancient organisms, such as the Neanderthal. Results showed that the DNA of homo sapiens have between 22 and 36 substitutions, as opposed to 55 from a chimpanzee. These results showed that the Neanderthal and homo sapiens shared a common ancestor.
Types of Libraries
[edit | edit source]The first combinatorial libraries where based on peptides (consisting of few of their monomeric units, the 20 animo acids). Other oligomeric libraries (from carbohydrates, nucleic acids, etc.) were also made out of adding component A to B, resulting in AB, then adding C to obtain ABC, adding D...1
This was the main technique for creating combinatorial libraries until the synthesis of benzodiazepine (fusion of benzene ring with diazepine ring) libraries was performed by DeWitt3. Instead of using the same technique as the oligomeric libraries, these libraries were based on the attachment of several R substituent chains to a molecule in different positions. The number of products obtained in the library is calculated d by multiplying the number of substituent chains used on the main molecule, in this case benzodiazepine. The R groups are referred to as functionalities, while the points where they attach in the central molecule are called points of diversity.1
Library Synthesis
[edit | edit source]There are two ways of synthesizing libraries through combinatorial quemistry: 1. Using solid-phase synthesis 2. Through solution-phase synthesis
Solid-phase synthesis This technique allows an easy isolation and manipulation of the product, as well as having simple control systems (automation). Excess reagents can be used to complete reactions. Still it has certain disadvantages, such as the specificity of the solid support that don’t allow certain chemical reactions to be performed.
Solution-phase synthesis On the contrary, solution-phase synthesis allows a better manipulation of the sample (once isolated) in any amounts without being concerned about the solid support. Its disadvantage is difficult sample isolation, reason why scientists are working on implementing the use of reactants that give pure products (or at least products that can be easily purified).1
Diversity-oriented Libraries
[edit | edit source]Even though combinatorial chemistry has been an essential part of early drug discovery for more than two decades, so far only one combinatorial chemistry-synthesized chemical has been approved for clinical use by FDA (sorafenib, a multikinase inhibitor used for renal cancer) (Newman & Cragg 2007). The analysis of poor success rate of the approach has been suggested to connect with the limited chemical space covered by products of combinatorial chemistry. When comparing the properties of compounds in combinatorial chemistry libraries to those of approved drugs and natural products, it was noted that combinatorial chemistry libraries suffer particularly from the lack of chirality, as well as structure rigidity, which are both widely regarded as drug-like properties. Even though natural product drug discovery has not been the most fashionable trend in pharmaceutical industry in recent times, a large proportion of new chemical entities still is nature-derived compounds, so it has been suggested that effectiveness of combinatorial chemistry could be improved by enhancing the chemical diversity of screening libraries. As chirality and rigidity are the two most important features distinguishing approved drugs and natural products from compounds in combinatorial chemistry libraries, these are the two issues emphasized in so-called diversity oriented libraries, for example compound collections that aim at coverage of the chemical space, instead of just huge number of compounds.
Library management
[edit | edit source]As we all know, integrating all types of chemistry, biology, and mixed information is exceptionary difficult and important combinatorial chemistry techniques. Thus, without managing all the information, library can not be function as one expected. For instance, if one scientist are trying to synthesize product "A," library must be able to show all the information necessary to synthesize the "A" such as, key reagent, cost, or if the reaction procedures are reliable etc... it is because the question that one will ask is different depends on the researchers.
Molecular Evolution
[edit | edit source]Evolution requires three processes:
1. The generation of a diverse population
2. The selection of members based on some criterion of fitness
3. Reproduction to enrich the population in these more-fit members.
Molecular evolution primarily occurs due to mutations, which are changes in the genetic material of a cell. They can occur during copying errors or due to exposure to chemicals and radiation. Natural selection removes the less favorable mutations. Key topics under molecular evolution are the study of evolution of enzyme function and the study of nucleic acids. Nucleic acids have shown the divergence of species because they are denoted as the "molecular clock."
References
[edit | edit source]- W.A. Warr, Encyclopedia of Computational Chemistry. Combinatorial Chemistry, John Wiley & Sons, Ltd.
- S. R. Wilson and A. W. Czarnik (eds.), Combinatorial Chemistry. Synthesis and Applications, Wiley, New York, 1997.
- S. H. DeWitt, J. S. Kiely, C. J. Stankovic, M. C. Schroeder, D. M. R. Cody, and M. R. Pavia, Proc. Natl. Acad. Sci. USA, 1993, 90, 6909–6913
- http://www.netsci.org/Science/Combichem/feature02.html
- Gianneschi, C. N., Rubinshtein, M., James, R.C., Kobayashi, Y., Yang, J., Young, J., Yanyan, J.M. Org. Lett., 2010, 12 (15), pp 3560–3563
- Kobayashi, Y., Buller, J.M., Gilley, B.C., Org. Lett., 2007, 9 (18), pp 3631–3634
Comparative Genomics
[edit | edit source]Nowadays, there is a vast numbers of genomes were sequenced. It is stunning how similar the genome of other species to human. For example, Drosophila (fruit fly) has more than half of the genes have human counterparts, even though the species does not look “human” at all. The result is even more stunning when scientists compared human’s genes with mammals.
Information about different genomes requires sciences to have a new field: comparative genomics. Comparative genomics studies the relationship between genomes of different species and tries to discover more genomes. Moreover, this field also attempts to answer many evolutionary questions. Recently, the draft sequence of chimp, out closest living relative, has been completed. By comparing the genome of chimp and human, scientists may have the answer of how we evolve from chimp in a biological aspect.
Example: Fruit fly - genome
Processes of Comparing Genomes
[edit | edit source]Comparative genomics works by aligning sequences [5] of different organisms to identify patterns that operate over both large and small distances. Aligning mouse chromosomes with human chromosomes, for example, shows that 99% of our protein-coding genes align with homologous sequences in mice. Underlying such analyses is the principle that DNA sequences that are highly conserved are likely to be functionally important. A common assumption is that adding more comparative genomes to the alignment helps distinguish functionally significant from irrelevant conserved sequences.
If a desired genome is unsequenced, it would take a long time to completely sequence species’ genome from beginning, and then compare it to others. However, science can make use of its relative species to finish this task quicker and easier. In order to compare unsequenced genomes, genome science uses shared synteny, a conserved arrangements of DNA on chromosomes of related species. Take a look at wheat for an examples, wheat has even more genomes than human; therefore, sequencing it would be too abundance.
Benefits of Comparitive Bioinformatics
[edit | edit source]1. Helps our understanding of the genetic basis of diseases in both animals and humans.
2. Increases our basic knowledge of the evolutionary pathways of related species.
3. Helps find new medical treatments and other means of benefiting human health.
4. To determine the function of human genes → for example, researchers can look for genes in humans in other animals whose functions are known. If scientists have identified a particular gene in another animal and know what it does, a gene in a human with a similar sequence probably has a similar function as that in the animal. This is called "annotating" - defined as creating a set of comments, notations, and references describing the experimental and inferred information about a gene or protein.
Examples
[edit | edit source]1. Chloroplast genome and mitochondrial genome -- Chloroplast and mitochondria also contains a significant amount of genetic information for a cell. Therefore, it should also be considered in the studies of genomics. Chloroplast is the organelle in plant that functions in photosynthesis. However, it also possesses its own genome, and thus can be independent of the cell in replication. Compared with the DNA in the nucleus, DNA in chloroplast does not evolve or evolves extremely slowly. Moreover, this DNA also cannot be modified or mutated over generation since recombination does not happen. As a result, scientists can easily provide the detail about evolution. It was found that genetic exchange had occurred between the chloroplast and nucleus. Some of the proteins are also encoded from the chloroplast genome. Similar to chloroplast genome, mitochondrial genome does not change significantly overtime. Mitochondrion in a cell is made by both its nucleus and mitochondria DNA. This shows the interaction between the two genomes, the nucleus’ and the mitochondria’s.
2. Fruit flies -- Although fruit flies have a genome that is 25 times smaller than the human genome, many of the flies' genes are similar to those in humans and control the same biological functions. Research on fruit flies has led to discoveries on the influence of genes on diseases, animal development, population genetics, cell biology, neurobiology, behavior, physiology and evolution.
They are also used for Parkinson's research. Researchers have found that two-thirds of human genes known to be involved in cancer have counterparts in the fruit fly. When scientists put a human gene associated with arkinson's disease into fruit flies, they displayed symptoms that humans have. This might mean that they could serve as a new model for finding a cure for Parkinson's.
DNA as an ID tool if do not know what type of organism it is. Take what species DNA that one would like to sequence and then first look on Genbank- which is the public bank of DNA to see if it has sequences before and since that is public knowledge. It is also somewhat of a Taxonomy browser. If for example seahorses are to be sequenced to see if a certain species at an aquarium is a certain one, the sequences that are available are the Cytochrom B sequence. Just need to sequence the DNA at the cytochrome area of the genome. In order to do this, must amplify that region- the purpose is in order to make millions of copies of that region. Take the primers that will complement that region and the people who have already sequenced that region will already have specified primers that can be used to determine the sequence with. All sorts of primers may be used even ones that have recently died just ones without any smell. Take the DNA that was sequenced and then compare with the DNA from the bank. Input what was sequenced and ask the search engine for the best match. The results will show species that have the highest similarity to the one that was sequence and give the best match.
| Organism | Estimated Genome Size (MB) | Estimated Number of Genes | Year Sequenced |
|---|---|---|---|
| Human | 2,900 | 20,000-25,000 | 2001 |
| Mouse | 2,600 | 30,000 | 2002 |
| Pufferfish | 365 | 33,609 | 2002 |
| Rat | 2,750 | 20,973 | 2004 |
| Chimpanzee | 3,100 | 20,000-25,000 | 2005 |
| Red Jungle Fowl | 1,000 | 20,000-23,000 | 2004 |
| Fruit Fly | 137 | 13,600 | 2000 |
| Mosquito | 278 | 46,000-56,000 | 2002 |
| Fission Yeast | 13.8 | 4,824 | 2002 |
| Brewer's Yeast | 12.7 | 5,805 | 1997 |
| Protist | 23 | 5,300 | 2002 |
| Wall Cress | 125 | 25,498 | 2000 |
| Rice | 430 | 41,000 | 2002 |
- (Reference: Biology, Eighth Edition by Raven and Johnson)
Comparison of human genome with other species':
1. Human vs. Pufferfish: Pufferfish was the first vertebrate that has its sequence compared with human. The latest shared ancestor of humans and pufferfish was 450 million years ago. However, their genomes still have many some similarities. Only about ¼ of human genes have no counterparts in pufferfish. However, 97% human DNA is repetitive while the number is only about 17% in pufferfish.
2. Human vs. Mouse: This is the first genome comparison made between two mammals. The similarity is much more significant. Both humans and mouse have about 25000 genes; and surprisingly, they share 99% genome. Human and mouse have the same ancestor about 75 millions years ago, much shorter compared with human-pufferfish ancestor. However, it was found that mouse DNA mutated two times faster than human’s DNA. That created 300 genes unique (only 1%) to both organism; and human genome has 400 million more nucleotides than mouse’s.
3. Human vs. Chimpanzee: Chimpanzee is one of the closest relative to human. We shared a common ancestor only about 35 millions years ago. In 2005, the genome of chimpanzee was completely sequenced and compared with humans’ genome. Only 1.06% difference in substitution and 1.5% difference in insertion and deletion were detected. Those insertion and deletion may provide us distinct characteristics from chimps, including lack of body hair and larger cranium.
4. Human vs. Plants: Estimated 1/3 of the genes in plants are not found in mammals. Those genes encoded plants’ distinguished characteristics such as photosynthesis and photosynthetic anatomy. The other 2/3 is very similar to human and animal genome. The similar genes encoded for basic metabolism, genome replication and repair, RNA transcription and protein synthesis.
References
[edit | edit source]Overview
[edit | edit source]Transgenic animal are animals that have had foreign genes from another animal introduced into their genome. A foreign gene (such as a hormone or blood protein) is cloned and injected into the nuclei of another animal’s in vitro fertilized egg. Cells are then able to integrate with the transgene, and the foreign gene is expressed, upon which the developing embryo is surgically implanted in a surrogate mother. The result of this process, if the embryo develops, is a transgenic animal housing a particular gene from another species.
Applications of transgenic technology are for example, improving upon livestock, such as higher quality wool in sheep, or increasing the amount of muscle mass of an animal so that it can produce more meat for consumption. Conversely, transgenic animals can also be utilized for medical purposes such as producing human proteins by inserting a desired transgene into the genome of an animal in a manner that causes the target protein to be expressed in the milk of the trangenic animal.
The expression of a transgene can also be engineered to take place in plants, such as obtaining the bio-luminescent gene that gives fireflies their glow in the dark ability, and introducing it to a plant.
Transgenic Mice in Alzheimer's Research
[edit | edit source]
Recently, transgenic mice have been used to conduct research concerning Alzheimer’s Disease. Researchers have created a new transgenic mouse that carries the gene for human Alzheimer’s disease, which represents a breakthrough in the development of effective treatments for a currently incurable disease. Researchers developed these transgenic mice by harvesting DNA that activate genes in the brain affected by the disease, including the hippocampus and cerebral cortex. This DNA was then inserted into mice embryos via microinjection techniques. This new DNA then became part of the mouse’s genotype through hybridization of this foreign DNA with the mouse’s own DNA.
Observable results in the brains of the mice were not seen within the first six months of life, but between six and nine months, the transgenic mice developed similar characteristics as ones seen in human Alzheimer’s patients. Such changes include damage to the neurons in the brain, reduction in synapse density, and accumulation of pathogenic beta-amyloid deposits. These beta-amyloid deposits increased in size as the mice aged and reached levels proportional to the amount of deposits that develop in human Alzheimer’s patients. The development of the amyloid plaques have been shown in transgenic mice to have adverse affects on memory, which resemble memory deficits seen in human patients.
The Morris water maze is a test conducted on transgenic mice demonstrating the memory loss resulting from damage to the hippocampus. This test involves placing a mouse inside a shallow pool of water that contains a submerged platform, which remains out of site of the mouse for the duration of the test. The mouse is placed in the pool on the opposite of the platform and is allowed to find the submerged platform by swimming around the pool and using visual cues on the surrounding walls. The test is performed multiple times and each time the mouse is placed in the same pool, the time spent swimming around in search of the platform decreases, which is a result of the formation of memories the mouse has stored of the various cues. The Alzheimer’s transgenic mice perform poorly when subjected to this same test, which indicates significant damage to their hippocampus: the location of memory storage in the brain.
One of the main reasons for developing these transgenic mice is to determine whether or not these harmful amyloid plaques are the cause for Alzheimer’s Disease, and what types of treatment can be developed to prevent them from forming and/or limit their growth. These transgenic mice not only provide more insight into the cause of Alzheimer’s but they also function as models for testing drugs to treat the disease.
More transgenic animals: [6], [7]
Resources
[edit | edit source]http://www.accessexcellence.org/WN/SUA02/alzheimers_breakthrough.php Protein regulation is essential for biological balance. Too much or too little of any proteinreactivity can cause severe biological damage.
Specific Diseases
[edit | edit source]Alzheimer's Disease
[edit | edit source]Alzheimer's patients have "plaques" in their brains, which are essentially large chunks of a certain protein believed to contribute to neuronal death. It is also a dementia that gets worse over time. There is no cure for Alzheimer's disease. As you can see in the picture, someone who has Alzheimer's has extreme loss in the different functions of the brain.
Alzheimer's patients have this protein's regulation pathway disturbed, causing it to be overly expressed. Protein regulation is also referred to as Enzyme Regulation. Furthermore, the study of these protein regulation pathways have lead to much growth in the creation of medicinal and pharmaceutical products.
Hunter Syndrome
[edit | edit source]Hunter syndrome is a genetic disease in which mucopolysaccharides do not degrade accurately. As a result, mucopolysaccharides will accumulate inside the body. The main cause of this accumulation is the absence of iduronate sulfatase enzyme. Some of the signs for recognition of the syndrome are big heads and different facial characteristics. The syndrome can be found out with urine test. However, this test is not trusted. The study of fibroblasts that extract from body’s skin is more effective. The gene that contributes to the cause of Hunter syndrome is residing on the X-recessive chromosome. Since males only have one X chromosome, the chance that males inherit this syndrome is higher than that of females.
The problem with Hunter syndrome is how the body is unable to break down mucopolysaccharides, which make up proteoglycan that is part of the extracellular matrix. As a consequence, the adding up of mucopolysaccharides becomes an obstacle for other cells in the body to carry out their jobs. This incident may lead to significant effects that can harm the body. For instance, some early side effects of Hunter syndrome can be the normal sickness such as cold and runny nose.
Hunter syndrome results in a certain effects upon the body. Thus, people who inherit the syndrome tend to have common characteristics. The more extreme consequences of Hunter syndrome include mental retardation, heart problem, joint stiffness, etc. Bone marrow replacement was proven to help expanding the life span of Hunter syndrome patients. Unfortunately, it does not solve the problem of mental retardation. Elaprase, a lysosomal enzyme iduronate sulfatase that can be made by recombinant DNA technique, has demonstrated to be an effective cure for Hunter syndrome through the replace of enzyme. Nevertheless, elaprase is indeed very expensive.
Hurler Syndrome
[edit | edit source]Hurler Syndrome is an inherited disease and is caused by a recessive mutation (both parents would need to have passed down the trait). Historically, it was thought to be caused by the excessive synthesis of 2 mucopolysaccharides: dermatan sulfate and heparan sulfate. Some scientists believed the excessive synthesis of the 2 molecules to be caused by a faulty regulation pathway. Elizabeth Neufeld tested this hypothesis and found it to be false; she found the cause of the disorder to be the inadequate degradation of the 2 sugars. While normal cells leveled off production after a certain point, cells affected by the mutation continued production past normal levels.
Correction of this disease is easily possible in vitro (and works to an extent in vivo.) The addition of healthy cells in the same culture as mutated cells in vitro caused normal levels of dermatan sulfate and heparan sulfate to be created. The normal cells excrete a corrective factor/enzyme in the medium which is taken up by the mutated cells. This enzyme is crucial in the degradation of the 2 mucopolysaccharides; furthermore, very little of it is necessary to fully correct the mutated cells. There are many problems with using α-L-Iduronidase as treatment in vivo; the main issue being that various tissues respond at various degrees to the medication. Most importantly, the central nervous system does not uptake any of the intravenously injected enzyme because of the blood-brain barrier. On top of this issue, a significant number of people with this disease have neurological diseases that need to uptake this enzyme somehow.
α-L-Iduronidase was found to be the key enzyme in restoring proper degradation of dermatan sulfate and heparan sulfate. It was found though that α-L-Iduronidase from some cells was not corrective. Research showed that the carbohydrate mannose 6-phosphate was responsible for proper uptake of the enzyme; a mutation in the structure of it caused the α-L-Iduronidase product to not be taken up properly by cells affected by Hurler Syndrome.
Sanfilippo Syndrome
[edit | edit source]Sanfilippo Syndrome is within a set of neurodegenerative diseases called tauopathies (the most common of which is Alzheimer's Disease). Although there are 4 subtypes of Sanfilippo Syndrome, they are all characterized by reduced degradation of heparan sulfate (see Hurler Syndrome section) due to reduced levels of a lysosomal enzyme.
It was found in a mouse model (MPS IIIB) that there were significantly increased levels of the protein lysozyme; increased levels of this disease were found to cause the creation of hyperphosphorylated tau which is found in the brains of Alzheimer's patients and patients with other tauopathies. Significant research is being done in Alzheimer's disease which may carry over to Sanfilippo Syndrome as well due to their similarities.
1. 4 different subtypes: Each Sanfilippo subtype is caused by the deficiency of each specific enzyme: heparin N-sulfatase for MPS-III A, N-acetyl-alpha-D-glucosaminidase for MPS-III B, acetyl-CoA: alpha-glucosaminide acetyltransferase for MPS-III C, and N-acetylglucosamine-G-sulfate sulfatase for MPS-III D. Among these four subtypes, Sanfilippo syndrome type A is the most prevalent (60%), followed by B (30%), D (5%), and C (6%). In total, 47% of all cases of mucopolysaccharidosis diagnosed is related with Sanfilippo disease.
2. Mortality/ Morbidity: Patients with Sanfilippo syndrome tend to develop Central Nervous System degeneration and usually end up at a vegetative state. They usually die before the age of 20 due to cardiopulmonary arrest because of airway obstruction or infection in the pulmonary pathway. Among these 4 subtypes, MPS-III A is the most severe one due to early death in patients (usually during their teenage years). Sanfilippo syndrome has an equal effect on both males and females as well as on different races since its main cause is the inheritance in an autosomal recessive pattern which has no relations with the sex chromosomes.
3. Diagnosis and History: In terms of diagnosis, these four subtypes are not indistinguishable clinically; therefore, the only determining factor to identify each specific subtype is the different genes that are responsible for that subtype. Usually, affected individuals show no symptoms and develop normally during the first two years of their lives. Onset usually takes place between the age of 2 and 6. Developmental delays in infancy may be shown in some of them. Growth might be slowed down at the age of 3 (e.g. short stature). Patients might also become hyperactive and behave aggressively and destructively. Besides disturbing the sleep pattern in patients, this syndrome also interferes severely into the mental development of affected patients – speech impairment, hearing loss, etc. At the same time, patients might show shortened attention spans and find it challenging to concentrate and to be able to perform academic tasks at schools. By the age of 10, patients’ daily activities and movements are severely limited. They often are in need of wheelchairs to accommodate them and might even have swallowing difficulties and seizures. Some other physical symptoms might also be shown such as carious teeth, enlarged liver and spleen, diarrhea (which is believed is due to lysosomal glycosaminoglycans (GAG) storage in the neurons of the myenteric plexus3.) Respiratory compromise can occur and is related to airway obstruction due to anatomical changes, excessive thick secretions and neurologic impairment. Upper respiratory tract infections and sinopulmonary disease are common.
4. Work-up: In order to diagnose patients with Sanfilippo Syndrome, specific enzymatic assays in cultured skin fibroblasts and in peripheral blood leukocytes are used (e.g. enzymatic cell analysis). One indicating sign for this syndrome is the increase in the level of heparan sulfate in the urinary secretion. Thus a total quantitative or a fractionation test are carried by performing either electrophoresis or chromatography with the purpose of measuring how much Glycosaminoglycans (GAGs) is in the urine. Due to the higher level of GAGs in newborns and infants, age-specific controls and fractionation must be included to accurately quantify the level of GAGs. Imaging studies can also be used to look for changes in brain structure since the spectrum of skeletal changes can be seen in patients with Sanfilippo syndrome.
5. Treatment: Currently, there is no available treatment for the real cause of Sanfilippo syndrome. Bone marrow transplantation and Enzyme replacement therapy only work for patients with mucopolysaccharidosis I, II, and VI (not III). However, some promising therapies are making their ways to be FDA-approved.
References
[edit | edit source]- Defendi, Germaine, [8]
- Neufeld, Elizabeth F, "From serendipity to therapy."
- [9]
- [10]
Overview
[edit | edit source]One of the biggest problems in bioinformatics is the relationship between amino acid sequence, structure, and function of proteins. The three dimensional structure of proteins helps in the development of drugs, engineering of enzymes, and analysis of protein functions.
There is the much-debated controversy of structural space and the conservation of structures and their relationship to the homology of proteins. The prediction of tertiary structures, however, relies on the re-use of subjective fragments without the need for homologous sequences (between target sequence and fragment source).
One of the many methods used by biologists and researchers to predict the tertiary structure of proteins is the GenTHREADER. This method aids in the detection of protein templates and sequence-structure alignment accuracy. Most methods used for fold prediction use known protein structures as a basis of the scoring of alignments calculated among protein sequence profiles. Two versions of GenTHREADER are the pGenTHREADER and pDomTHREADER methods, which are used to recognize and align protein sequences in order to analyze their relationship to the structure and function of the protein. Both of these versions have similar inputs of protein sequence profiles and structural information; the use one core alignment algorithm.
The pGenTHREADER uses profile-profile comparisons from matrices built from sequences, coiled coil regions and filtered trans-membrane fragments. The final step includes two sequence-profile and profile-sequence scores, which enabled profile-profile matches that was higher scoring that other matches. In addition, a hydrophobic burial terms is added that serves the purpose of biasing alignments’ positions in a target sequence.
In determining results, the best results of a template-target pair were used to calculate the number of equivalent residues rather than the results for a method; this differentiates alignment accuracy from template selection. Picking the best result significantly improves performance of chains less than 200 amino acids but shows little improvements in longer chains. pGenTHREADER works significantly better than other methods when the fold recognition relationships are more distant; with shorter distances, other methods tend to show more improvement. These other methods used mixes of fold recognition, side chain optimizations and model quality assessments.
- [11]_(Selected_Internet_Resources)
Fold.it is an online game that combines human brain power and computer science together to solve one of the hardest problems in biology--the protein folding problem.
[edit | edit source]== This game was invented by protein researcher David Baker and his fellow computer scientists from University of Washington. The objective of fold.it is find the best structure(the one that is in the lowest energy level)for desire protein. Players use tools like building hydrogen bonds or rotating the structure around. They collect points as they lower the energy of the structure. Players of fold.it are mostly people who don't have knowledge in protein structures. However, these players has done a lot of amazing works in improving our knowledge of proteins. One of the biggest accomplishments they have done was finding the three dimensional structure of a virus that cause AIDS in monkey in 2011. While the supercomputer tried to solve this protein folding problem for 15 years,a group of gamer found the solution in as short as three weeks. This group of gamer didn't want to reveal their real names but their passion for the game has advanced protein study significantly. If you want to be part of this evolutionary game, you can go to http://fold.it/. The game has a clear interface and is very easy to start. As you learn how to use different tools you can use for the game, you also learn more about all the different amino acids interactions in real actions and see it happen in the game. For more information about professor David Baker and his research work, you can go to http://depts.washington.edu/bakerpg/drupal/. ==
Molecular Modeling Overview
[edit | edit source]Molecular modeling refers to abstract methods and techniques for finding the molecular structures and properties by using computational chemistry and graphical visualization techniques to ‘model’ or copy the behavior of the molecule. Besides computational chemistry, they can also be used in fields such as computational biology and science that studies molecular structures. These methods or techniques can be used to figure out molecules ranging from combination of a few atoms such as CH3 to large macromolecules such as polypeptides by giving possible 3-D representation of those structures. The simplest atomic structures do not necessarily need computers to figure out the molecular model, but large molecules do; Molecular modeling helps to make it easier to understand each specific parts of the complex molecule and also allow more atoms to be considered during simulation. The two most common models that are used in molecular modeling are quantum mechanics and molecular mechanics. A method of molecular modeling, molecular docking, can be used to discover and design new molecules.
Common Models
[edit | edit source]Quantum Mechanics are principles describing the physical reality at the atomic level of matter (molecules and atoms) and the subatomic (electrons, protons, and smaller particles), and it include the simultaneous wave-like and particle-like behavior of both matter and radiation. It is a mathematical description of reality, which is usually different than how humans see a set of bodies or how a system behave. The most complete description of a system is its wavefunction, which is a number varying between time and place. In quantum mechanics, quantum refers to a discrete unit that quantum theory assigns to certain physical quantities, such as the energy of an atom at rest. Particles are discrete packets of energy with wave-like properties which led to the branch of physics that deals with atomic and subatomic systems. This area of study is the quantum mechanics. The principle between classical and quantum mechanics is that all objects obey laws of quantum mechanics, and classical mechanics is just a quantum mechanics of large systems.Quantum mechanics are usually compared with classical physics, but they are not the same because they aren't defined at the same time by the Universe. Quantum theory also provides accurate descriptions for many previously unexplained phenomena such as black body radiation and the stability of electron orbitals. It has also given insight into the workings of many different biological systems, including smell receptors and protein structures.
- three fundamental ways Quantum mechanics differs from classical physics:
- the integration of particle and wave phenomena with the relative equivalence of mass and energy
- the quantization of both wave and particle phenomena
- the uncertainty involved in making physical measurements.
- Quantum mechanics theory was formed by a continuation of important discoveries: 1838 discovery of cathode rays by Michael Faraday, the 1859 statement of the black body radiation problem by Gustav Kirchhoff, the 1877 suggestion by Ludwig Boltzmann that the energy states of a physical system could be discrete, the 1900 quantum hypothesis by Max Planck and then the 1905 postulation of light itself consisting of individual quanta called photons, by Albert Einstein.
Molecular Mechanics refers to the use of classical mechanics (Newtonian mechanics) to describe the physical basis behind the models. Molecular models usually describe atoms as point charges with an associated mass. The interactions between neighboring atoms are described by spring-like interactions (representing chemical bonds) and van der Waals forces. Molecular mechanics can be used to study small molecules as well as large biological systems or material assemblies with many thousands to millions of atoms. Simple energy functions can be quick to solve, can deal with large molecules, accurate for systems which are close to the models used to reproduce the force field, and can be used to specify mandatory bonds in a molecule.
- Molecular mechanics force field: Simple equations to describe the energy cost of deviating from ideal geometry
- E = Es + Eb + Ew + Enb
- Es is the energy involved in the deformation of a bond either by stretching or compression; Eb is the energy involved in angle bending; Ew is the torsional angle energy; Enb is the energy involved in interactions between atoms that are not directly bonded
- Epot=∑Vs+∑Va+∑Vt+∑Vv+∑Ve
- where Epot is the energy of the potential function; Vs the bond stretch potential of all bonds; Va is all the bond angle bending; Vt is torsion; Vv is van der waals of all atoms; Ve is the electrostatic interactions
- E = Es + Eb + Ew + Enb
- Molecular mechanics minimization: a method of minimizing the energy by changing the structure toward optimum geometry
- Properties of molecular mechanics methods:
- each atom is represented as a single particle
- each particle has a radius, has polarizability, and a constant net charge
- Bonded interactions are treated as "springs" with an equilibrium distance equal to the experimental or calculated bond length
Single Molecule Biophysics
[edit | edit source]In the past decade, new methods have helped scientists visualize microscopic molecules. These new techniques such as atomic force microscopy, optical and magnetic tweeezers, and single-molecule fluorescence spectroscopy allow scientists to visualize molecules on a single molecule scale instead of how large gigantic systems interact and taking averages of large moles.
The single molecule method takes advantage of being able to determine distinct structural states or large biomolecules. Though it cannot reveal as much structural information as can X-ray crystallography it can obtain nanometer-scale information on structural features.
Atomic Force Microscopy
[edit | edit source]
This tool was first developed for topographically imaging molecules on a flat atomic surface. AFM is generally used on generating static images of biomolecules and can be performed on dry or samples in solution. Due to these factors AFM has been useful for spectroscopy of protein structure.
IBM in Zurich, Switzerland has improved the AMF technique well enough to capture the most detailed and smallest-scale image of a pentacene molecule.

The main idea behind atomic force microscopes is that an image can be created from a detailed force map of minute atomic scale forces. These are sensed by some sort of probing device; analogous to hands feeling an object in a completely dark room as to gain a sense of it's shape. The atomic probe is thereby creating an image of the surroundings in a see-by-feel manner.
The problem one would expect with this close-up sort of atomic measurement is the attractive force of Van Der Waals. At very close distances, this force is strong enough to potentially pull and subsequently attach the penacene directly to the surface of the probe. Luckily, a phenomena noted as the Pauli exclusion principle prevents this sort of behavior. This priciple states that quantum particles called fermions cannot occupy the same quantum state within a certain range of each-other.
The microscope uses a carbon monoxide "tip" to pan over a penacene molecule (or other desired molecule) which is bound to a silicon surface. The carbon monoxide molecule is bound to the probe directionally, such that the oxygen is aligned along the axis of force measurement. This relatively inactive oxygen is able to measure varying forces along the surface of the penacene molecule. This is accomplished through atomic forces created by the repulsive behaviour described by Pauli's principle. This 2D force map is used to create the corresponding image.
Magnetic Tweezers
[edit | edit source]This approach is often used for studying the structural properties of DNA and protein-DNA transactions.
Methods/ Techniques
[edit | edit source]Classical Molecular Dynamics is a computational method for simulating the motion of particles. It involves calculating the spatial derivatives of the energy to get the force acting on each atom according to Newton's Laws of Motion.
Ab-initio methods may also be used to simulate the motion of atoms by solving the Schroedinger equation to obtain results that are more accurate but much more computationally expensive.
Molecular Docking is a method which predicts the preferred orientation of one molecule that binds to another molecule. It is an important tool in structural molecular biology and computer-assisted drug design because it use computers to figure out the shape and properties of a molecule. An example of molecular docking is when a ligand binds to a protein to form a stable complex. Even when a protein and a ligand does not completely complement each other, they would adjust their conformation so there would be an overall "best fit" (also called induced-fit). The goal of molecular docking is to minimize the free energy of the whole system by achieving an optimized conformation for both the protein and ligand.
Challenges In Molecular Modeling
[edit | edit source]- free energies
- Thermodynamic free energy is the energy in a physical system that can be converted to do work:
- First law: conservation of energy
- Second Law: the universal principle of entropy, stating that the entropy of an isolated system which is not in equilibrium will tend to increase over time, approaching a maximum value at equilibrium.
- Third Law: deals with entropy and how it is impossible to reach the absolute zero of the temperature.
- Thermodynamic free energy is the energy in a physical system that can be converted to do work:
- solvation is the process of attraction and association of molecules of a solvent with molecules or ions of a solute.
- simulating reactions is the modeling of natural systems or human systems in order to gain insight into their functioning and how reactions happens.
References
[edit | edit source]- http://www.chemdiv.com/en/Information/glossary/
- http://en.wikipedia.org/wiki/Molecular_modelling
- http://www.booksites.net/leach1/molecular/
- http://www.bmb.uga.edu/wampler/399/lectures/mm1/sld001.htm
- http://www.ch.ic.ac.uk/local/organic/mod/mo_3.html
- http://www.sdsc.edu/~kimb/molmech.html
- http://en.wikipedia.org/wiki/Molecular_dynamics
- http://en.wikipedia.org/wiki/Quantum_mechanics
- http://www.bookrags.com/research/quantum-mechanics-wsd/
- http://en.wikipedia.org/wiki/Docking_(molecular)
- http://en.wikipedia.org/wiki/Solvation
- http://en.wikipedia.org/wiki/Thermodynamic_free_energy
Overview
[edit | edit source]In studying enzyme catalysis, an important factor to understand is how the substrate and enzyme interact with each other. The affinity of the substrate to the enzyme is a deciding factor to whether or not the binding in the active site occurs. There are many things that affect the affinity of the substrate to the enzyme. The surface, complementarily, flexibility and non-covalent charges are all important factors. Another influencing factor includes the orientation of the molecule. Molecular modeling is a technique used in computational chemistry, computational biology and material science to mimic the behavior of molecules to further understand its systems and how they bind to each other. A type of molecular modeling is docking. Molecular docking is a method that predicts the preferred orientation of a molecule when it binds to another molecule. It uses computers to create a 3D image of two molecules and shows how they fit together. Knowing the preferred orientation that a molecule has for another can help predict the strength of binding affinity that a substrate will have for an enzyme. A common application that molecular modeling is used for is drug design. When creating a new drug for a certain disease, molecular docking can be used to predict the binding affinity that the new drug has on the targeted protein. However, there are certain errors that can occur in relying mainly on this approach. For example, while molecular modeling can predict the preferred orientation that a drug may have for a protein, it is not able to effectively account for conformational changes that may occur. But despite this, molecular docking is still a very important technique that helps us further understand the molecular processes that occurs within the molecular level.
Theoretical Overview
[edit | edit source]
Classical molecular dynamics (MD) is a branch of computational chemistry that focuses on simulating the motion of particles according to Newton's Laws of Motion.[1] Atoms are approximated as "balls on springs," which allows for the application of the following laws:
Where is position, is velocity, is acceleration, and is time.
Classical MD approximations treat bond and angle potentials as harmonic, and dihedrals as periodic. This allows for expression of the energy of a system with an equation similar to the following used by the AMBER molecular dynamics program: [2]
Simulations are conducted in a series of "time steps." In each step, the force on each atom is calculated according to the derivative of the energy equation. The atom is moved a certain distance corresponding to the magnitude and direction of the force over the size of the time step. The process is repeated over many time steps, iteratively calculating forces and solving the equations of motion based on the accelerations from the forces, and produces results that mimic complex behavior from docking of pharmaceutical compounds to the folding of simple proteins.

Limitations
[edit | edit source]Quantum Effects
[edit | edit source]Although atoms are known to behave according to the Schroedinger equation rather than by Newton's Laws, classical methods can be used with reasonable accuracy. A good indication of this can be given by checking the De Broglie wavelength of the particles involved [3]:
where is the atomic mass and is the temperature. The classical approximation is justified when the mean nearest neighbor separation between particles. So for heavier systems, such as organic molecules, the classical approximation is reasonable, but for light systems such as H2, He, or Ne, the method loses considerable accuracy.
Since atoms are approximated as discrete particles, interactions of smaller particles cannot be correctly modeled with classical MD.[4] Making and breaking chemical bonds, noncovalent reaction intermediates, and proton or electron tunneling all cannot be simulated with this method. Simulations at very low temperature are also problematic, as near absolute zero classical mechanics are increasingly disfavored for quantum explanations. Quantum molecular dynamics methods exist for simulations that can be more electronically realistic, but at much greater computational cost.
System size and timescale
[edit | edit source]The number of atoms and amount of time that may be simulated is directly related to available computing power. Although the entirety of a small virus has been simulated before,[5] the majority of fine-grained, all-atom simulations that take place are on the scale of proteins rather than organelles or complex assemblies. This is because the length of time that is necessary to simulate to observe results directly relates to system size:
Table - Timescales of molecular processes
[edit | edit source]| Secs | name | Process |
|---|---|---|
| 10−1 | Surface reorganizations | |
| 10−3 | milliseconds | Protein Folding |
| 10−8 | 10 nanoseconds | NMR |
| 10−9 | to 10+3 | Brownian dynamics |
| 10−9 | to 100 | Lattice Monte-Carlo |
| 10−12 | picoseconds | Fast vibrations |
| 10−15 | femtoseconds | Electronic excitation |
So, the larger the system, the longer the simulation needs to be run to observe relevant behavior. Today, computer hardware specifically designed for molecular dynamics simulations[6] can run approximately 17 microseconds of a 23,000 atom system in one day with a 0.5 femtosecond timestep. On the same system, a general-purpose parallel supercomputer can obtain on the order of a few hundred nanoseconds of simulation time.
Thermostats
[edit | edit source]Using an approach based on Newton's equations of motion, a simulation is obtained in the "microcanonical ensemble", which is a constant energy system (nVE). This is unrealistic, however, and in order to produce accurate comparisons to experiment, simulations are most commonly done in the canonical ensemble, which features constant temperature (nVT).[7]This temperature must be maintained with an algorithm, which is called a thermostat. There are several thermostats that are commonly used, each of which has its advantages and disadvantages in terms of reliability, accuracy, and computational expense.
A typical simulation involves an MD equilibration phase, at which the structure is simulated at constant temperature for a certain amount of time at 0K, a heating phase where the system is gradually heated to the desired temperature (often 300K), and a final production simulation at that constant temperature. The initial equilibration is necessary not to move the structure to a chemical equilibrium but rather to an energetic minimum (which will usually be near the chemical equilibrium state). If the equilibration were omitted and the simulation run in a region with high energy, the simulation will continue to sample highly energetic phase space, producing results that will be inconsistent with experimental data.[8]
The Langevin Theromostat
[edit | edit source]This thermostat follows the Langevin equation of motion rather than Newton's, where a frictional force proportional to the velocity is added to the conservative force, which adjusts the kinetic energy of the particle so that the temperature is correct.[9]
The system is then described by the equation:
Where , and are mass, velocity, and acceleration of the particle. is a frictional constant, and is a random force that will add kinetic energy to the particle. It is sampled from a Gaussian distribution where the variance is a function of the set temperature and the time step. This balances the frictional force to maintain the temperature at the set value.
The Anderson Thermostat
[edit | edit source]This thermostat couples the system to a heat bath at the desired temperature. The bath is represented by collisions with a stocastic particle on randomly selected system particles.
It has been shown[10] that the addition of stochastic collisions to the Newtonian MD system results in the simulation being a Markov chain that is irreducible and aperiodic. This implies that the generated distribution with Anderson's algorithm is not a canonical distribution.
This thermostat will randomly affect velocities of particles, which results in nonphysical dynamics in the simulated system.
The Gaussian Thermostat
[edit | edit source]This thermostat strongly couples the system with a heat bath set to the desired temperature. Here, the change in momentum for each particle is represented as:[11]
This rescaling destroys dynamic correlations very quickly, and therefore does not maintain the canonical ensemble.
The Berendsen Thermostat
[edit | edit source]This thermostat, first presented in 1984[12] involves re-scaling of the velocities of each particle in a manner that controls the temperature.
Similar to the Gaussian thermostat, the system is coupled to a heat bath with the desired temperature, although this algorithm uses weak rather than strong coupling, where the temperature of the system is corrected such that the deviation in temperature decays exponentially with some time constant proportional to the timestep:
The Nose-Hoover Thermostat
[edit | edit source]This thermostat adds an extra term, to the Newtonian equations of motion, producing the following second-order differential:
Where is mass, is the desired temperature, and is the Boltzmann constant. The heat bath is given some mass, , which for Nose's suggested value of produces the above equation for the momentum change of each individual particle.
With this thermostat, the energy of the simulated system fluctuates, but the total energy of the system and heat bath is conserved. If the system is ergodic (all accessible microstates are equally probable), then this system gives the canonical ensemble.[13]
Force fields
[edit | edit source]Molecular dynamics simulations require a "force field," which lists the parameters involved in each type of interaction in the energy equation. For example, the AMBER force field requires parameters for bond equilibrium length, bond force constant for every possible combination of two atom types, angle equilibrium value, and angle force for every combination of three atom types, dihedral equilibrium phase, dihedral force constant, and dihedral periodicity for every combination of four atom types. Additional parameters also describe the nonbonded (electrostatic) forces for each atom as well.
The development and analysis of these parameters, which collectively are called a force field, is a focus of molecular dynamics research.
Force field development
[edit | edit source]Parameters for classical molecular dynamics ideally lead to behavior that closely approximates that of actual molecules on an atomistic level, which are in reality governed by quantum phenomena. The classical parameters are developed such that this is true- once an energy and force equation (such as the AMBER equation) has been made to establish the paradigm for the system, parameters are derived such that calculated values with them match those calculated with quantum mechanical methods or experiment. Due to the lack of availability of experimental data for the wide variety of molecules and molecular conformations necessary to develop a force field, data from quantum calculations on the systems are used instead.
Force fields are developed for specific classes of molecules. For example, AMBER provides the Amber ff99SB forcefield for proteins[14], the Glycam forcefield for carbohydrates,[15] and the newly developed lipid forcefield GAFFLipid.[16] In addition, a "General AMBER Force Field," or GAFF, has been developed for the parameterization of small ligands and other molecules that do not fit into the broad force field categories.
The accuracy of a force field is critical to simulation accuracy. Simple terms, such as dihedral force constants, can have an extremely large effect on overall simulations. In fact, inaccurate dihedral parameters for the protein backbone and angles produced simulations that were excessively disposed to alpha helix formation, a fact which affected numerous results.[17][18][19]
Historically, force field development has been complicated by the lack of availability of energy and force data to fit to. Force field parameters are generated by fitting the parameters such that the energy of a structure or the predicted forces on each atom predicted using the parameters matches the energy or forces generated by ab initio quantum calculations. These calculations are extremely expensive, and as a result considerable extrapolation is usually made from one set of parameters to a general one in order to avoid re-doing the calculations. For example, using parameters generated for the protein backbone of tetra-alanine as describing the backbone of any protein.
The availability of increasing computing power, more efficient algorithms, and knowledge of past methods proves promising for future parameter development. For example, side-chain parameters for various amino acids may be calculated separately and then added in to a general parameter set in order to characterize each amino acid individually.[20]
Force field accuracy
[edit | edit source]Classical force fields have been developed by fitting the classical parameters such that they give results consistent with experimental and quantum data. They are categorized by the type of biomolecule- protein, carbohydrate, and lipid forcefields have been developed separately for all major MD programs. However, this categorization is still quite broad, and can neglect system-specific interactions.

- ↑ Meller, Jarosaw. "Molecular Dynamics" Encyclopedia of Life Sciences, 2001
- ↑ R. Salomon-Ferrer, D.A. Case, R.C.Walker. An overview of the Amber biomolecular simulation package. WIREs Comput. Mol. Sci.(2012)
- ↑ J. P. Hansen and I. R. McDonald, Theory of Simple Liquids, 2nd Ed., Academic, 1986.
- ↑ Ercolessi, Furio. "A Molecular Dynamics Primer" University of Udine Spring College in Computational Physics, 1997
- ↑ Peter L. Freddolino, Anton S. Arkhipov, Steven B. Larson, Alexander McPherson, and Klaus Schulten. Molecular dynamics simulations of the complete satellite tobacco mosaic virus. Structure, 14:437-449, 2006.
- ↑ David E. Shaw, Martin M. Deneroff, Ron O. Dror, Jeffrey S. Kuskin, Richard H. Larson, John K. Salmon, Cliff Young, Brannon Batson, Kevin J. Bowers, Jack C. Chao, Michael P. Eastwood, Joseph Gagliardo, J.P. Grossman, C. Richard Ho, Douglas J. Ierardi, István Kolossváry, John L. Klepeis, Timothy Layman, Christine McLeavey, Mark A. Moraes, Rolf Mueller, Edward C. Priest, Yibing Shan, Jochen Spengler, Michael Theobald, Brian Towles, and Stanley C. Wang (July 2008). "Anton, A Special-Purpose Machine for Molecular Dynamics Simulation". Communications of the ACM (ACM) 51 (7): 91–97.
- ↑ Hunenberger, Phillipe. Thermostat Algorithms for Molecular Dynamics Simulations DOI:10.1007/b99427
- ↑ Finnerty, J. "Molecular Dynamics meets the physical world: Thermostats and Barostats"
- ↑ Adelman, S.A. and J.D. Doll, Generalized Langevin Equation Approach for Atom-Solid-Surface Scattering - General Formulation for Classical Scattering Off Harmonic Solids. Journal of Chemical Physics, 1976.
- ↑ E, W. and Li, D. (2008), The Andersen thermostat in molecular dynamics. Comm. Pure Appl. Math., 61: 96–136. doi: 10.1002/cpa.20198
- ↑ J. Clarke D. Brown. Mol. Phys., 51:1243, 1983.
- ↑ Berendsen, H.J.C., et al., Molecular-Dynamics with Coupling to an External Bath. Journal of Chemical Physics, 1984. 81(8): p. 3684-3690.
- ↑ Palmer, R. G. (1982). Advanced Physics 31: 669
- ↑ Lindorff‐Larsen, Kresten, et al. "Improved side‐chain torsion potentials for the Amber ff99SB protein force field." Proteins: Structure, Function, and Bioinformatics 78.8 (2010): 1950-1958.
- ↑ Kirschner, Karl N., et al. "GLYCAM06: a generalizable biomolecular force field. Carbohydrates." Journal of computational chemistry 29.4 (2007): 622-655.
- ↑ Dickson, C. J., Rosso, L., Betz, R. M., Walker, R. C., & Gould, I. R. (2012). GAFFlipid: a General Amber Force Field for the accurate molecular dynamics simulation of phospholipid. Soft Matter.
- ↑ Best, Robert B., Nicolae-Viorel Buchete, and Gerhard Hummer. "Are current molecular dynamics force fields too helical?." Biophysical journal 95.1 (2008): L07-L09.
- ↑ Freddolino, Peter L., et al. "Force field bias in protein folding simulations." Biophysical journal 96.9 (2009): 3772.
- ↑ Lindorff‐Larsen, Kresten, et al. "Improved side‐chain torsion potentials for the Amber ff99SB protein force field." Proteins: Structure, Function, and Bioinformatics 78.8 (2010): 1950-1958.
- ↑ Lindorff‐Larsen, Kresten, et al. "Improved side‐chain torsion potentials for the Amber ff99SB protein force field." Proteins: Structure, Function, and Bioinformatics 78.8 (2010): 1950-1958.
Interactive 3D Model within a PDF (3D PDF) is a process by which a selected structural data is integrated into a 3D model within a PDF. This process allows a shortened learning curve to learning how to operate it, compared to the more well known software’s that are used to visualize structural data. On top of being user-friendlier, interactive 3D model within a PDF allows for a 3D model to be presented in any publications or writings, which allows for the reader to get a visual perspective on the molecule that they are reading about. The PDF version allows for an integration of other media, such as video and audio. The main advantage of 3D PDF is the ability of the reader to access 3D structures, while also being able to read the article at the same time [1].
Exporting Molecular Models in 3D into PDF format
[edit | edit source]Using PyMOL, which is a popular software tool used for molecular visualization, one can embed 3D images into PDF publications through the process of three steps: 1. Export components of the 3D model as a VRML2 files through the use of PyMOL. The file is in (.wrl) form. 2. Process the model components by commercial 3D software, which allows the user to get a U3D file, which comes with all of the parts of the 3D model. The file is in (.u3d) form. 3. In order to embed the U3D into a PDF file, one can use Adobe Acrobat 9 Pro Extended software suite in order to complete the process. [2]
Alternative ways to create a 3D PDF
[edit | edit source]1. One alternative way to create a 3D PDF is to use the function of the Adobe Acrobat 9 Pro Extended, called “3D capture.” In this, the OpenGL graphics signal is captured. The PyMOL and Adobe Acrobat 9 Pro Extended are opened at the same time and the picture that is displayed in PyMOL is captured by the Adobe Acrobat 9 Pro Extended in a 3D form. After this the file is saved in the PDF form. The advantage to this is that the saved file can be edited and then saved as U3D file, which as pointed out before, is embedded into a PDF file that have all the proper settings, resulting in a 3D PDF model. The downside to this alternative way is that the models are much smaller compared to the original results from other procedures. [2]
2. Another alternative way to create a 3D PDF involves the LaTEX document preparation system. In this approach the LaTEX document preparation system is used with the movie 15 package. The Adobe Acrobat 9 Pro Extended creates files, either a U3D or a PRC file, which contain the 3D molecular models. These files are inputted and from that a PDF document is produced, containing the 3D image. The important part is that the LaTEX and movie 15 script for the integration of the models into the PDF file. [1]
3. A third alternative way to create a 3D PDF is called MeshLab. Using VRML2 files from PyMOL, MeshLab software turns these files into U3D format, which can be used to embed 3D images into PDF files using Adobe Acrobat 9 Pro Extended or the LaTEx/movie15. [1]

Existing Problems with 3D PDF
[edit | edit source]1. Some online submission systems are unable to build the final PDF file using individual files. 2. There are submission systems that do not accept the 3D PDF format files. 3. File size of publication that contain interactive 3D images must be below 10 MB in order to be accessible.
Sources
[edit | edit source]1] Kumar, Pravin, Alexander Ziegler, Alexander Grahn, Chee Hee, and Andreas Ziegler. "Leaving the Structural Ivory Tower, Assisted by Interactive 3D PDF." PubMed. N.p., n.d. Web. 20 Nov. 2012. <http://www.sciencedirect.com/science/article/pii/S0968000410000629>.
[2] Kumar, P. et al. (2008) Grasping molecular structures through publication-integrated 3D models. Trends Biochem. Sci. 33, 408–412
Structural Biochemistry/Mental Inertia in the Biological Sciences/ The field of biology is an experimental science, where the logical theory is only as good as its experimental background. Therefore, there comes a dilemma when accepting theories, should the scientist faithfully accept the known theory for what it is or should the scientist attempt to verify them experimentally? One of the main causes of mental inertia is due to these incorrectly established results. The first form of mental inertia is belief in generally accepted observations, oftentimes due to blind faith in the authoritative source. For example "...Aristotle thought there were eight legs ona fly and wrote it down. For centuries, scholars were conetnt to quote his authority'.[1] This idea could be easily refuted yet due to mental inertia scientists just accepted this theory of years and years.
Another form of mental inertia is when scientists still adhere to the observations made with faulty techniques when a better technique is already available. If an incorrect observation was made before better techniques came around, then that is an mistake, not mental inertia. It is only mental inertia if better techniques came, that would allow the scientists to improve the experiment and observations, and the scientist still uses the old observations for the basis of their research.
References
[edit | edit source]- ↑ Chase, S. (1938) The Tyranny of Words, Harcourt, Brace and Company
Overview
[edit | edit source]Mechanism actions are constantly being improved and updated as new information becomes available. Its a scientists' responsibility to keep up to date with what's new, as well as think critically about what is being stated and implied. However, when scientists don't think critically and merely accepts things for they are, they fall prey to mental inertia. Mental inertia caused by an incorrect understanding of a mechanism action can be split in four categories: naming, linkage groups, scientific paradigms, and improper controls.
Mental inertia can be caused by the naming of a mechanism. While it may not seem like it, the naming of a mechanism is extremely important. A mechanism's name implies its function and the group of mechanisms it belongs to. Therefore if a mechanism is poorly named it could cause confusion or incorrect assumptions. For example, hormone-independent activation of hormone receptors were overlooked for years because these proteins were initially called 'receptors for hormones'. Mental inertia can also be caused by incorrectly looking at linkage groups-when involvement of the first mechanism is considered as evidence for involvement of the second, and likewise the lack of involvement of the first mechanism means the lack of involvement of the second mechanism. This causes faulty thinking when different mechanisms for different stages of the same process are seen as different indications of a single mechanism. Scientific paradigms causes mental inertia by hindering scientists from thinking outside the traditional paradigms. For example, a set scientific paradigm was the idea that the death of a normal mammalian cell only occurs because something is killing it. Because of this paradigm, nobody considered the idea that normal cells can commit suicide for years. Now, this idea of cell suicide is currently one of the cornerstones of biology. Lastly, mental inertia can be caused by improper controls for an experiment, when controls are merely accepted because they are commonly used. This mental inertia is self-perpetuating because as more improper controls are used, the more common they become.
Water
[edit | edit source]Unique Properties of Water
[edit | edit source]- Structure of Water
- Polarity and Hydrogen Bonding
- Cohesive and Adhesive Behavior
- Ability to Moderate Temperature
- Expansion upon Freezing
- Versatility as a Solvent
- High Dielectric Constant
- High Heat of Vaporization
- Phase States of Water
- High Dielectric Constant

The pH of a solution is defined as the negative logarithm of its hydrogen ion (H+) concentration. For example, If the concentration of H+ ion is 10^(-7),
then the pH of the solution = -log (10)^(-7) = 7.
Therefore, as the Hydrogen Ion concentration increases, pH value decreases
& as the Hydrogen Ion concentration decreases, pH value increases
The pH of a solution is a measure of the hydronium ion (H3O+) concentration on a logarithmic scale. The pH scale range is 0-14 from acidic to basic, respectively. The pH of a neutral compound, such as pure water at room temperature, is 7. The concentration of hydronium ions is related to the concentration of hydroxide ions by the dissociation of water:
H2O H+ + OH-
There is some confusion as to the meaning of the "p" in pH. Although many believe it to stand for "power" or "potential," (i.e. power of hydrogen), that may not actually be the case. According to Jens G Norby's research (Dept of Biophysics, University of Aarhus, DK), "power of hydrogen" is actually a false association made by W.M. Clark. In actuality, Sorensen, the originator of the term, "pH" did not explicitly state a meaning for the "p," and it appears to be an arbitrary selection of letters "in his initial explanation of the electrometric method." For more information about this misconception, see, "The origin and the meaning of the little p in pH."
pH is biologically important because it affects the structure and activity of macromolecules. pH is important in homeostatic processes. For example, most animals breathe not because they lack oxygen, but because CO2 buildup in the blood increases the blood acidity beyond normal levels. Enzyme function is especially sensitive to pH: each has an optimum pH at which they have maximum catalytic ability. Extreme pH levels can denature enzymes, completely disrupting their function. Other proteins are also destabilized by extreme pH levels.
The pH and pOH of a solution are related such that: pH+pOH=14. For example, if the pH of a solution is 5, the pOH of the same solution will be 14-5=9.
The pH of distilled water is 7, this is neutral. Any solution with a pH below 7 (i.e. pH 0 to pH 6.9) is acidic and any solution with a pH above 7 (i.e. pH 7.1 to pH 14) is basic.
Acidic solutions have a pH between 0 and 6.9 (the stomach contains HCl, which has a pH ~2). Alkaline solutions have a pH between 7.1-14 (the small intestine is pH 9). Neutral solutions are neither acidic nor alkaline so their pH is 7.
Hydrogen Chloride dissociates into hydrogen and chloride ions:
HCl(aq)= H+ + Cl-
Water dissociates to produce Hydrogen and hydroxide ions:
H2O(l) = H+ + OH-
Sodium Hydroxide dissociates to produce Sodium and hydroxide ions:
NaOH(aq) = Na+ + OH-
In each case the concentration of Hydrogen ions can be measured or calculated.[H+] represents hydrogen ions and the square brackets represent concentration.
• pH = -log[H+]
• pH + pOH = 14
• DECREASING pH, INCREASING pKa
• pH = pKa + log ([A-]/[HA])
• pH = pKa
Mathematics
[edit | edit source]The pH of a solution is calculated by
pH = -log [H+]
By convention H+ is used to represent hydronium ions (H3O+). A log system simplifies the notation of the H+ concentration in a medium. A pH increase of 1 is indicative of a ten-fold decrease of H+ concentration. This negative logarithmic relationship between the hydrogen ion concentration and pH means that a lower pH indicates a higher concentration. Conversely a higher pH indicates a lower concentration of hydrogen ions.
The ionization of water is expressed by equilibrium constant: Keq = [H+][OH-]/[H2O], which is 1.0 * 10−14 at 25°C.
The tendency of an acid, HA, to lose a proton and form its conjugate base, A-, is defined by the acid dissociation constant, Ka. Another quantitative measurement of acidity is the pKa, which is calculated from the Ka (pKa = -log Ka). A smaller Ka means a higher value of pKa, thus a higher value of pKa is equivalent to a weaker acid due to less dissociation of the acid into H+ and its conjugate base. Similar to pH, a single integer difference in pKa represents a tenfold difference.
The pH of a solution is determined by relative concentrations of acids and bases.
Using the reaction of the dissociation of a weak acid, HA:
HA H+ + A-
The Ka can be written as:
Ka = [H+][A-] / [HA]
and rearranged as:
[H+] = K*[HA] / [A-]
Taking the negative log of each term:
-log[H+] = -log(Ka) + log([A-]/[HA])
Letting pH = -log[H+] results in:
pH = -logKa + log([A-]/[HA])
Letting pKa = -logKa gives:
pH = pKa + log([A-]/[HA])
This relationship of pH to pKa is known as the Henderson-Hasselbach equation.
When the concentration of an acid, HA, and its conjugate base, A-, are equal:
log([A-]/[HA]) = log 1 = 0
Yielding:
pH=pKa
meaning that the pH of the solution is equivalent numerically to the pKa of the acid. This point is also known as the half equivalence point.
This equation is useful for calculating the pH of a buffer solution, or a solution containing a known quantity of weak acid (or base) and its conjugate base (or conjugate acid). From a biochemistry perspective, Hederson-Hasselbach can be applied to amino acids. However, this equation does not account for the ionization of water in a solution and thus is not useful for calculating the pH of solutions strong acids or bases.
See Buffer for more information on weak acid or weak base calculations.
pH Calculation for Polyprotic Acid
[edit | edit source]Polyprotic acid such as sulfuric acid, H2SO4 or phosphoric acid, H3PO4, is capable of being deprotonated more than once because of the presence of two or more H atoms within the molecule. However, the pKa of each form of a polyprotic acid determines the contributions of hydrogen atoms, and consequently a change in pH, to a solution.
For example, consider the dissociation phosphoric acid:
H3PO4 H2PO4- + H+ Ka1 = 7.1 x 10−3 (1) Ka1 = H2PO4- HPO42- + H+ Ka2 = 6.3 x 10−8 (2) Ka2 = HPO42- PO43- + H+ Ka3 = 4.5 x 10−13 (3) Ka3 =
Supposing we have 0.01 M H3PO4 we could calculate the pH of the solution based on the Ka values. Comparing the three pKa values of H3PO4, it is clear that the first deprotonation (1) contributes the most H+ because it is the highest value. At equilibrium, x can represent the M [H+] and this value is the same for [H2PO4-] as well. While [H3PO4] can be expressed as 0.01-x M. Therefore,
- Ka1 = = 7.1 x 10-3
After which the quadratic equation can be used: x2 + 0.0071x - 0.000071 = 0
- To solve this equation, we use , and then obtain
- x = 5.59 x 10-3 = [H+] = [H2PO4-]
- pH = -log[H+] = -log 5.59 x 10-3 = 2.25
A less time-consuming (but also less accurate) approach can be used to avoid using the quadratic equation. If an acid dissociates less than 5% or if the acid concentration to Ka ratio is greater than 103, the following equation can be used:
[H+] = ; where Ca is the concentration of acid. (4)
However, in this case we see that 0.01M H3PO4 dissociates more than 5% in the first deprotonation.
Therefore, using quadratic equation to solve this problem is the right choice.
Since phosphoric acid has three acidic protons, the second and third deprotonations may also contribute to pH of the solution.
When phosphoric acid was first deprotonated, the [HPO42-] concentration was approximately 0.00559 (calculated above). The [HPO42-] to Ka2 ratio is greater than 103. In this case equation (4) can be used to approximate [H+] from the second deprotonation. From calculation, [H+] =
As shown from calculation, [H+] contribution from the second deprotonation is much less than that from the first, and makes an insignificant change in pH. Similarly, calculations from the third deprotonation reveal that even less [H+] (approximately 2.9 x 10−9 M) is contributed. As shown by the differing pKa values, the majority of [H+] comes from the first deprotonation.
Now consider the multiple deprotonations of sulfuric acid:
H2SO4 HSO4- + H+ Ka1 = large (5) Ka1 = HSO4- SO42- + H+ Ka2 = 1.2 x 10−2 (6) Ka2 =
Equation (5) shows a large value for the first pKa, implying that sulfuric acid dissociates completely in the first deprotonation. (See acidic substances) Thus, if we have 0.01 M H2SO4, [H+] from the first dissociation would also be 0.01 M. In addition, there would be 0.01 M HSO4- in the solution, and is a source of protons by the second deprotonation (equation 6). Before using equation (4), check to see if the concentration of acid to Ka ratio is greater than 103 or not. Since this simplification cannot be used, the quadratic equation is used to determine [H+] from the second dissociation. The equation is as follows:
- x2 + 0.012x - 0.00012 = 0
- Again, is used to obtain
- x = 6.5 x 10-3 = [H+]
The second deprotonation gives a [H+] more than half of which was obtained from the first deprotonation. In contrast to the example with phosphoric acid, the second dissociation of sulfuric acid is contributes greatly to the concentration of H+. In total, [H+]tot = 0.01 + 0.065 = 0.0165 M
The pH = -log 0.0165 = 1.78
Sulfuric acid is a much stronger acid than phosphoric acid as seen by its H+ contribution in each deprotonation step.
An Alternative Method to the Quadratic Equation
[edit | edit source]The quadratic equation can become messy and time consuming in certain situations. Lets consider the following hypothetical situation:
HA A- + H+
KHA = 6.3 x 10−3
[HA] = 2
[A-] = 0
[H+] = 1x10−1
Our equilibrium equation would therefore be:
= 6.3x10−3
Now we take into account the overall shift in equilibrium from the dissociation of HA and include the given concentrations.
= 6.3x10−3
Because there is already a large concentration of protons in the solution the overall dissociation of HA, given by the variable x, will be very small. Therefore we are going to ignore the change in x in the denominator to yield an equation that is not quadratic.
= 6.3x10−3
Solving for x yields approximately 0.11853 which will call x1 which we will use to successively obtain a more accurate number by plugging this number back into our original equation where we ignored x previously:
= 6.3x10−3
Solving using x1 yield x2 to be 0.05604
In order to simplify the process the following equation is used to solve for xn+1 derived from our original rate expression.
xn+1 =
The following successive approximations are calculated to be:
x3 = 0.07761
x4 = 0.06851
x5 = 0.07207
x6 = 0.07063
x7 = 0.07098
x8 = 0.07107
x9 = 0.07103
x10 = 0.07105
When xn starts repeating itself to a given amount of significant figures, it is safe to assume that the given answer is an accurate approximation and can be used for the final value of x. The overall number of successive approximations that must be calculated varies according to the situation. In some situations it is simpler to use the quadratic equation. An equation that required many successive approximations was used for the purposes of this example. Ingenuity is require to obtain an accurate in a time efficient manner.
Contribution of Water in Dilute Acidic or Basic Solution
[edit | edit source]One more thing we need to consider in case of dilute acid or base is the contribution of H+ or OH- from water. As we have already known, at 25 oC water molecule could dissociate and H+ as well as OH- would form with the amount of 10−7 M each. For example, if we have 0.0000001 M acetic acid, we can determine the pH of the solution as follows:
Acetic acid is a monoprotic acid which has pKa 1.8 x 10−5. To determine the H+ of the solution, let's consider
CH3COOH CH3COO- + H+ Ka = 1.8 x 10−5 Ka =
Regarding the previous section, we need quadratic equation to solve this problem, Thus
- x2 + 0.000018x - 0.0000000000018 = 0
Then, we get x = 1 x 10−7 = [H+] the concentration of [H+] from 0.0000001 M acetic acid is nearly close to the ones obtained from water. In this case, we have to take into account the [H+] from water. Therefore,
- [H+]tot = 2 x 10-7 M
Finally, the pH = -log 2 x 10−7 = 6.7
Remark For dilute basic solution, we can evaluate in the similar way.
pH Scale Diagram
[edit | edit source]The scale is broken up into two regions: acidic (pH 0-7) and basic (pH 7-14) with a pH of 7 being said to be "neutral." Some acidic substances are gastric acid (pH ~ 2), vinegar (pH ~ 3) and coffee (pH ~ 5). Some basic substances are hand soap (pH ~ 10), ammonia (pH ~ 13), and lye (pH ~ 14). An example of a "neutral" substance would be pure water (pH ~ 7). pH values below 0 require measurements using specialized equipment. The reason is that typical pH measurements are done in an aqueous solution, which means that the measured pH will only be as strong as the weakest acid in solution (in this case H3O+ which has a pKa of 0). Therefore, measurements are typically made by calibrating with a strong acid like sulfuric acid. Similar arguments also apply for extremely strong bases.
Acidic Substances
[edit | edit source]There are three definitions of acids based on Arrhenius, Bronsted-Lowry, and Lewis. By Arrhenius theory, acids are molecules that produce hydronium ions (H3O+) in aqueous solution. The Bronsted-Lowry definition of an acid is a molecule that can donate protons (H+). A Lewis acid is defined as a molecule that is able to accept electron pairs.
The most common strong acids include hydrochloric acid (HCl), hydrobromic acid (HBr), hydroiodic acid (HI), sulfuric acid (H2SO4), perchloric acid (HClO4), and nitric acid (HNO3). Where perchloric acid is the strongest with a pKa = -10 In calculations, strong acids are assumed to dissociate completely. For example: By convention 2.5 M HCl produces 2.5 mol of hydronium ions per liter.
Basic Substances
[edit | edit source]
Just as there are three definitions of acids, there are three corresponding definitions of bases. An Arrhenius base is defined as a molecule that produces hydroxide ions (-OH) in water. A Bronsted-Lowry base is a molecule able to accept hydrogen ions. Finally, a Lewis base is able to donate electron pairs.
Bases are usually characterized by having a negative charge (surplus of electron pairs), which according the to Lewis theory makes it capable of donating to another molecule. Bases can also have a neutral charge, but contain a lone pair of electrons (such as ammonia) which can be donated to other molecules.
Common strong bases include: Lithium hydroxide (LiOH), Sodium hydroxide (NaOH), Rubidium hydroxide (RbOH), Cesium hydroxide (CsOH), Potassium hydroxide (KOH), Magnesium hydroxide (Mg(OH)2) Calcium hydroxide (Ca(OH)2) Strontium hydroxide (Sr(OH)2) Barium hydroxide (Ba(OH)2). A strong base, like a strong acid, is defined as a compound that dissociates completely in aqueous solution.
Application in Biochemistry
[edit | edit source]pH range for amino acids have Zwitterionic structure is 2-9. Below pH 2 the carboxyl group is not dissociated. Above pH 9, but amino group is not dissociated.
Zwitterion
[edit | edit source]
Since amino acids have both an acidic carboxylic acid group and a basic amine group, they can be either positively charged, negatively charged, or both at the same time depending on the pH. A zwitterion is a molecule that has both a positively charged group and a negatively charged group at the same time, thus carrying a total net charge of 0. The zwitterion is the most commonly found form of amino acids when the pH is in the neutral range, since in the neutral pH, the basic amine group is protonated (carries a positive charge) and the carboxyl group is deprotonated (carries a negative charge).
Isoelectric Focusing
[edit | edit source]Since animo acids, and therefore proteins, have multiple sites that can be either positive or negative, the total overall charge of a protein depends on the pH. Isoelectic Focusing is a technique where proteins are separated based on the pH at which they become neutrally charged overall. This point is called the pI, or isoelectric point. Isoelectric Focusing is done by loading the sample proteins onto a gel with a pH gradient and then applying an electric current. The proteins will then move across the gel because of the electric current and will stop moving once they reach their pI. Isoelectric focusing can be combined with SDS-PAGE to increase the separation of proteins in a sample, a technique called Two-Dimensional Electrophoresis.
The Bohr Effect
[edit | edit source]Christian Bohr, a Danish physiologist, defined the Bohr effect as the change in binding affinity of oxygen to hemoglobin based on hydrogen ion and carbon dioxide concentrations. Oxygen has higher affinity when the pH of the blood is high. CO2 and H+ exist in the following equilibrium in the blood:
CO2 + H2O H+ + HCO3−
Thus, a change in either concentration will alter the pH. An increase in carbon dioxide will increase the concentration of hydrogen ions, and therefore lower both the pH and oxygen's affinity for hemoglobin. This benefits the body because areas that are high in carbon dioxide are likely deficient in oxygen. The low pH in those areas causes oxygen to disassociate from hemoglobin, releasing it into the body tissues that need it most.
As the pH increases, so does the ability for hemoglobin to bind oxygen. However, as the pH moves to a lower pH (as slight as a drop from 7.4 to 7.2), the ability to bind to oxygen decreases.
Carbon Dioxide
Carbon dioxide also affects the pH inside a red blood cell (which affects the ability of oxygen to bind to hemoglobin). As carbon dioxide flows into a red blood cell, carbonic anhydrase (an enzyme specific for this reaction) facilitates its interaction with water in forming carbonic acid, which then dissociates into bicarbonate and hydrogen ions.
CO2 + H2O H2CO3
H2CO3 H+ + HCO3-
The increase in H+ decreases the pH, and stabilizes the T-state of hemoglobin, also known as the deoxygenated state.
The transport of carbon dioxide into a red blood cell is carried through via membrane transport. It also directly interacts with hemoglobin, affected the binding and release of oxygen. The carbon dioxide decreases the oxygen binding affinity and stabilizes the T-state of hemoglobin even better than the effects of pH on oxygen binding.
Carbamate Group
Carbamate groups are negatively charged groups formed from the interaction of carbon dioxide and the last amine group in an amino acid. The carbamate groups stabilize the decrease in the binding of oxygen by increasing the salt bridge.
References
[edit | edit source]http://www.purchon.com/chemistry/ph.htm http://en.wikibooks.org/wiki/Structural_Biochemistry/Buffer
General Information of Buffers
[edit | edit source]A significant change in pH can lead to harmful reaction to molecular structure, biological activity and function. Protein structure can be disrupted and enzymes denatured due to the effects of pH on cellular structure. Fortunately, nature has evolved a solution to this problem; solutions that resist changes in pH are called buffers. If an acid is added to an unbuffered solution, the pH will change suddenly and proportional to the amount of acid added. However, the pH will drop gradually in buffer solution when acid is added. Buffers also mitigate the pH increase caused by adding base.
Buffers are aqueous systems that resist changes in pH as acid or base is added. They are usually composed of a weak acid and its conjugate base. Biological buffers, mixture of weak acids (the proton donors) and their conjugate bases (the proton acceptors), help maintain biomolecules in optimal ionic state of pH 7. Buffering is a result of two reversible reaction equilibrium occurring in a solution of about equal concentration of proton donor and its conjugate proton acceptor; when acid or base is added to a buffer, it results in a change in the ratio of the relative concentrations of the weak acid and its anion, or the pH. However, this change in pH is significantly greater than it would be without a buffering solution to accommodate excess hydronium or hydroxide ions.
Properties
[edit | edit source]A buffer solution usually contains a weak acid and its conjugate base, but it can also contain a weak base and its conjugate acid. When H+ are added to a buffer, the weak acid's conjugate base will become protonated, thereby "absorbing" the H+ before the pH of the solution lowers significantly. Similarly, when OH- is added, the weak acid will become deprotonated to its conjugate base, thereby resisting any increase in pH before shifting to a new equilibrium point. In biological systems, buffers prevent the fluctuation of pH by processes that produce acid or base by-products to maintain an optimal pH.
Each conjugate acid-base pair has a characteristic pH range as an effective buffer. The buffering region is about 1 pH unit on either side of the pKa of the conjugate acid, where it has the most effectiveness for resisting large changes in pH when either acid or base is added.
Henderson-Hasselbalch Equation
[edit | edit source]The equilibrium constant for the deprotonation of an acid is written as:
(1)
Where is the concentration of a conjugate base and [HA] is the concentration of an acid
Taking logarithms of both sides, we get
(2)
subtract both sides by , we get
(3)
This is the Henderson-Hasselbalch Equation. It describes the dissociation of a weak acid (HA) in the presence of its conjugate base (A-).
The midpoint of the buffering region is when one half of the acid reacts to dissociation and where the concentration of the proton donor (acid) equals that of the proton acceptor (base); the pH of the equimolar solution of acid is equal to the pKa. (When the concentration ratio for conjugated base and weak acid, [A-]/[HA], is 1:1)
Limitations
[edit | edit source]Buffers usually work well at a pH close to the pKa value of the acidic component. If too much acid is added to the buffer, or if the concentration is too strong, extra protons remain free and the pH will fall sharply. This is reflected in the titration curve and is known as the buffer capacity.
Titration curve
[edit | edit source]This curve demonstrated the capacity of a buffer. In the middle part of the curve, it is flat because the addition of base or acid does not affect the pH of the solution drastically - this is the buffer zone. However, once the curve extends out of the buffer region, it will increase tremendously when a small amount of acid or base added to the buffer system. This effect demonstrated the buffer capacity of the solution.
H3PO4 titration curve
HCl titration curve
Physiological Buffers
[edit | edit source]Phosphoric acid system (pKa = 7.4) and carbonic acid system (pKa = 3.6) are two important buffer systems in human body. The phosphate buffer system is an effective buffer in the cytoplasm of all cells. H2PO4– acts as the proton donor and HPO42–- acts as the proton acceptor.
H2PO4– ⇋ H+ + HPO42–
The bicarbonate buffer system is used to buffer blood plasma where the carbonic acid (H2CO3) acts as a proton donor and bicarbonate (HCO3− acts as a proton acceptor.
H2CO3 ⇋ HCO3− + H+
In this buffer system, when the pH of the blood plasma is too high, the H+ of blood plasma is lowered. The H2CO3 is dissociated to H+ and HCO3−. The CO2 from the lungs is dissolved in the blood plasma resulting in a lower pH. On the other hand if the pH of the blood plasma is too low, H+ is added to the blood increasing the concentration of H2CO3. This results in an increase of CO2 in the blood plasma. The increase in the partial pressure of CO2 in the air space of the lungs causes extra CO2 to be exhaled ultimately resulting in a raise in pH. Since the concentration of CO2 can be adjusted rapidly through changes in the rate of respiration, a large reservoir of CO2 can quickly adjust this equilibria to keep blood pH at a nearly constant rate.
Buffers are also important for enzyme activities. There is an optimal pH for each enzyme. For example protein cleavage enzyme pepsin works at pH 1-6 (pH=2 max), trypsin at pH 2-9 (ph=6 max) and alkaline phospatase at pH 4-10(pH=8 max).
In addition to this, some reactions should be carried out at constant pH which is provided by buffer systems.
Osmosis
[edit | edit source]The tendency for aqueous solutions [solvents] to migrate across a semi-permeable membrane from a higher water potential to a lower water potential is known as diffusion. Semi-permeable membranes are selective, allowing only specific particles to flow through and preventing others from invading. Water always flows from lower solute concentration [dilute solution] to higher solute concentration until a balance is produced. Dilute solutions are highly concentrated in water and low in solute. Hence, a low concentration of solute results in a high water potential. The general motion is from more water to less water or from less solute to more solute.
- See Water Potential for more information.
Osmotic pressure
[edit | edit source]
During osmosis, pressure is generated due to the difference in solute potentials of two environments separated by a semi-permeable membrane. This pressure provides the force for the water to move from low solute concentration to high solute concentration. Once the water pressure reaches the osmotic pressure, osmosis stops. The osmotic pressure is measured by the van't Hoff equation: Π = icRT, where "ic" represents osmolarity. Osmolarity is a product of the molar concentration of solute and the van't Hoff factor, i, a measurement of the extent of dissociation of the solute. The van't Hoff factor is equal to how many particles the solute dissociates in to. For example, NaCl dissociates into Na+ and Cl-, so the van't Hoff factor is 2. Glucose does not dissociate, so it's van't Hoff factor is 1. If there are multiple solutes, the total osmolarity is equal to the sum of the osmolarities of the individual solutes. [1]
Reverse Osmosis
[edit | edit source]Reverse osmosis is the process by which excess pressure is placed on one end of a semipermeable barrier in order to drive a solution from an area of high solute concentration to that of a low solute concentration. Opposite from general osmosis, the solvent does not go down a concentration gradient. In this case, the cell membrane serves as a filter. However, the key difference between osmosis and filtration is that in reverse osmosis, separation is done by diffusive mechanisms rather than size exclusion or staining. Reverse osmosis has been used industrially for water treatment. Salt water collected from the ocean is transformed to pure water by setting an external pressure equal to the air pressure at sea level. Water purification is only one industrial use of reverse osmosis. Metals and chemicals are also recycled through this process.
Example of the environmental use of reverse osmosis: County Sanitation Districts
Types of Tonicity
[edit | edit source]
Isotonic
[edit | edit source]- The scenario when the solute concentration inside and outside of a cell (or semi-permeable membrane) is in equilibrium. There is no net movement of water if the cell is in the isotonic environment. Since there is equilibrium between the water inside and outside the cell, no net movement of water occurs.
Hypertonic
[edit | edit source]- When the solute concentration outside of a cell is higher than the solute concentration inside of a cell. Hence the cell shrinks since the water from inside of the cell rushes outside of the cell in order to equilibrate the solute concentrations inside and outside of the cell. Fish such as salmon have to excrete salt from their gills to prevent themselves from being in a hypertonic solution.

Hypotonic
[edit | edit source]- When the solute concentration outside of a cell is lower than the solute concentration inside of a cell. Hence the cell swells because the water from outside of the cell flows inside of the cell in order to make the solute concentrations equal. This may cause the cell to burst. To prevent bursting, cells have developed mechanisms to cope with its environment. Freshwater protists that inhabit hypotonic environments have organelles such as contractive vacuoles to pump water out from the cell. For bacteria and plants, a plasma membrane is surrounded by a rigid and non-expandable cell wall that has the strength to resist osmotic pressure and prevent osmotic lysis. Lastly, for multi-cellular animals to prevent bursting, blood plasma and interstitial fluid are used to maintain the osmolarity level close to that of cytosol.

The Role of Osmosis in Living Organisms
[edit | edit source]Living cells may be thought of as microscopically small bags of solutions contained within semi permeable membrane that allows water to flow in or out. In order for the cell to survive, the concentration of solutes within the cell cannot be changed dramatically. Water passes through the membrane in both directions to generate equilibrium between the cell and its surroundings. If the cells are in a highly concentrated solution, the water in the cell would flow out to maintain the equilibrium between the exterior and interior of the cell. This may cause the cell to shrink due to loss of water and die of dehydration. Oppositely, if the cells are in a more diluted solution, water will enter the cell and cause it to burst and be destroyed. In the molecular level, storage cells maintain osmotic pressures by storing energy in forms of macromolecules, such as polysaccharides, rather than micro molecules, such as glucose. By storing macromolecules, the osmotic pressures are diminished, thus preventing storage cells from bursting. The reason behind this is that osmolarity depends on the amount of solutes in the cell rather than the solute's size and mass. Therefore, the storage of macromolecules prevents a dramatic increase in osmotic pressure.
To survive through harsh conditions in nature, organisms have developed various methods to maintain their solute concentration within safe levels. For example, organisms that live in saltwater have much higher cell solute concentrations than organisms living in fresh water; other animals replace lost water and solutes by drinking and eating, or by removing the excessive water/solutes to decrease the solute concentration through excretion of urine.
Osmosis also plays an important role in plants. It contributes to the movement of water through parts of the plant. As the minerals and other solutions from soil are taken up by root cells to leaf cells, the solute concentrations in the plant cell will increase. It brings the differences of osmotic pressure between the cell and the exterior environment. As a result, water will be drawn upward and spread through the plants cell. When too much water is taken up into the cells, water is evaporated from leaves by regulating the size of the openings in the leaf surfaces to remove the excess water. In addition, osmolarity plays a role in plant rigidity. In a plant cell, the vacuole holds much of the plant volume and solute concentration. Because of this high solute concentration, the osmotic pressure causes water to enter the vacuole. However, because of the immutable cell wall, the cells do not burst from such a hypotonic solution. Instead, the cell stiffens, thus increasing the rigidity of the plant and its tissues.
The Role of Osmosis in Studying Organelles
[edit | edit source]Within the cell, there are organelles that have semipermeable membranes that allow the intake and output of water. Naturally, these organelles, such as mitochondria, chloroplasts, and lysosomes, exist in the cytoplasm where the solute concentration is higher. Keeping that in mind, in order to study organelles and isolate them from the cell, precautions must be taken in order to create an isotonic solution and prevent the organelles from absorbing too much water and bursting. Processes such as Differential Centrifugation depend on this precaution in order to obtain successful separation.
Reference
[edit | edit source]External Links
[edit | edit source]The Structure of Water
[edit | edit source]Water consists of two hydrogen atoms connected with one oxygen atom. The hydrogen atoms are connected to the oxygen in a covalent bond manner by sharing two electrons together. The structure of oxygen is that surrounding it there is the presence of two unpaired electrons. Oxygen is significantly more electronegative than Hydrogen, meaning that has a greater tendency to attract electrons.
Polarity and Hydrogen Bonding
[edit | edit source]As a result of this uneven distribution of electron density, water is polar. Both of the hydrogens have a partial positive(Delta+) and oxygen has a partial negative (Delta-) due to oxygen holding the shared electrons closer to itself and away from hydrogen. This attraction between the partial positive and partial negative creates what is known as a hydrogen bond. Hydrogen bonding only occurs when hydrogen is bonded with electronegative atoms such as oxygen, fluorine, or nitrogen, so this type of a bond is very unique.
The oxygen carrying partial negative charge can form hydrogen bonds with two neighboring hydrogen atoms and each hydrogen atoms in the water molecule can form one hydrogen bond with other electronegative atoms. So typically each molecule of water is able to bond with four of the neighbors in the proximity to form what is known as a tetrahedron network.
Indissolubility of molecules and ions in water because of its polarity. Another unique characteristic of water resulting from hydrogen bonding is that water is lighter in the solid state. The reason for that is because the hydrogen bonds keep water molecules in certain distance away from each other when water freeze. When water is in liquid state, hydrogen bonds form and break, then reform again constantly in the space. Making the density of water higher in liquid state.
In fact, water is extremely polar solution because it has the polar O-H bond. Given that oxygen has an extremely high electron affinity compared to hydrogen, oxygen will tend to pull the electrons away from the hydrogen atoms in the molecule. The massive polarity of the O-H bond is the result of this electron affinity. Water gas reaction happens at elevated temperatures and pressures between water and carbon (usually coke or coal):
H2O + C -> H2 + CO
The product of this is called synthesis gas and can be extremely useful in bonding with metallic heterogeneous catalysts. For example, in the Fischer-Trospsch process, transition metals are used as catalysts to speed up these reactions like H2 + CO -> Alkanes with Co as the catalyst. 3H2 + CO -> CH4 + H2O with Ni as a catalyst. 2H2 + CO -> CH3OH with Zn/Cu as a catalyst. Ni catalyst is used in a process called steam reforming where methane is mixed with steam at high temperature to create hydrogen gas and carbon monoxide.
Some examples are:
Methane: CH4 + H2O (+heat) → CO + 3H2
Propane: C3H8 + 3H2O (+heat) → 3CO + 7H2
Ethanol: C2H5OH + H2O (+heat) → 2CO + 4H2
Gasoline: C8H18 + 8H2O (+heat) → 8CO + 17H2 C7H8 + 7H2O (+heat) → 7CO + 11H2
In a water gas shift reaction, CO + H2O -> CO2 + H2 with the addition of Zn-Cu as the catalyst. This reaction is thermodynamically favorable at about 400 degrees Celsius.

References
[edit | edit source]http://www1.eere.energy.gov/hydrogenandfuelcells/production/natural_gas.html Miessler, Gary. Inorganic Chemistry. 4th Edition.


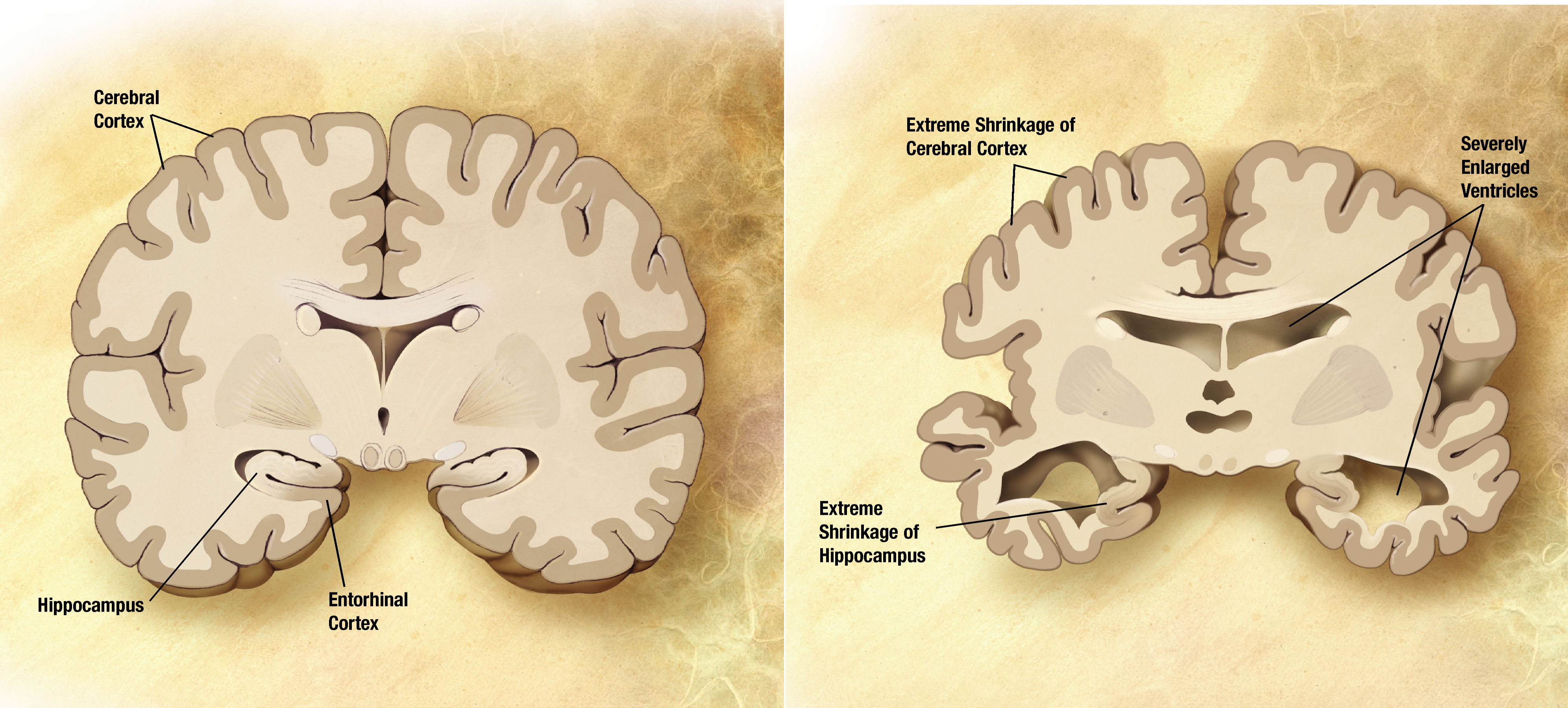







![{\displaystyle E_{\rm {total}}=\sum _{\rm {bonds}}K_{r}(r-r_{eq})^{2}+\sum _{\rm {angles}}K_{\theta }(\theta -\theta _{eq})^{2}+\sum _{\rm {dihedrals}}{V_{n} \over 2}[1+{\rm {cos}}(n\phi -\gamma )]+\sum _{i<j}\left[{A_{ij} \over R_{ij}^{12}}-{B_{ij} \over R_{ij}^{6}}+{q_{i}q_{j} \over \epsilon R_{ij}}\right]+\sum _{\rm {H-bonds}}\left[{C_{ij} \over R_{ij}^{12}}-{D_{ij} \over R_{ij}^{10}}\right]}](https://wikimedia.org/api/rest_v1/media/math/render/svg/07abb3f2d6f16ca1e1dede075b6faf02bc42b5df)



















![{\displaystyle {\tfrac {[H_{2}PO_{4}^{-}][H^{+}]}{[H_{3}PO_{4}]}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/dbdcbfb1f940453804f3957cd991ea06a7e6b8b7)
![{\displaystyle {\tfrac {[HPO_{4}^{2-}][H^{+}]}{[H_{2}PO_{4}^{-}]}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/e9417cf0c9396eac459b76692f305e91b6a57540)
![{\displaystyle {\tfrac {[PO_{4}^{3-}][H^{+}]}{[HPO_{4}^{2-}]}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/b3dd1ea87a6bc82a92ad592c8543723a910094c1)





![{\displaystyle {\tfrac {[HSO_{4}^{-}][H^{+}]}{[H_{2}SO_{4}]}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/029b7b29fef47d220c1f19aaac85276c08dc72c8)
![{\displaystyle {\tfrac {[SO_{4}^{2-}][H^{+}]}{[HSO_{4}^{-}]}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/9973be1f9f730a16cc97934db9e96ec55242d74e)
![{\displaystyle {\tfrac {[A^{-}][H^{+}]}{[HA]}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/a4b01e3812f22bbf95dd8ba6984548d1af6aebdc)
![{\displaystyle {\tfrac {[x][0.1+x]}{[2-x]}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/e7983be1057530f8a4098ec7bad91df4e4b43462)
![{\displaystyle {\tfrac {[x][0.1]}{[2+x]}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/24a7a003f63b3d811e9bf7326f253bbdfa30a138)
![{\displaystyle {\tfrac {[x][0.1+x_{n}]}{[2-x]}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/4af3939d99bae95ad39a9bf0e2437c8f68a55eb3)

![{\displaystyle {\tfrac {[CH_{3}COO^{-}][H^{+}]}{[CH_{3}COOH]}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/155cb7e565d976aa2e430e56d81fcf8e633ea831)


![{\displaystyle \mathrm {K_{a}={\frac {[H^{+}][A^{-}]}{[HA]}}} }](https://wikimedia.org/api/rest_v1/media/math/render/svg/14a41818964da5798042c1d67a87ead832bce748)
![{\displaystyle [A^{-}]}](https://wikimedia.org/api/rest_v1/media/math/render/svg/79a328ebb2b6d512330c0c19632d00c6ecb4082d)
![{\displaystyle \mathrm {\log K_{a}=\log H^{+}-\log {\frac {[A^{-}]}{[HA]}}} }](https://wikimedia.org/api/rest_v1/media/math/render/svg/f61bf9d0b318fcd068fc42aac0fb760552effa58)
![{\displaystyle log([A^{-}]/[HA])}](https://wikimedia.org/api/rest_v1/media/math/render/svg/2040afa61e7aafb3641f362b825c0720310cfe73)
![{\displaystyle \mathrm {pH=pK_{a}+\log {\frac {[A^{-}]}{[HA]}}} }](https://wikimedia.org/api/rest_v1/media/math/render/svg/f5ce641b42bcfbb625b84a4d200a79c43847f69e)



