Structural Biochemistry/Proteins/Structures
Here is a summary for the primary structure of a protein:
Primary Structure:
- It is a sequence of amino acids.
- It is a linear polymer: linking the alpha-carboxyl group of one amino acid to the alpha amino group of another amino acid => PEPTIDE BOND (covalent bond).
- In some proteins, the linear polypeptide chain is cross-linked: Disulfide bonds.
The primary structure is a polypeptide, in which:
- each amino acid in the peptide is a residue
- there is a regularly repeating segment called the main chain or backbone, and a variable part, comprised of the side chain.
Primary Structure
[edit | edit source]The primary structure of a protein is a linear polymer with a series of amino acids. These amino acids are connected by C-N bonds, also known as peptide bonds. The formation of peptide bonds produce water molecules as a by-product when an amino acid N-terminal loses hydrogen and another amino acid C terminal loses -hydroxyl group. Thus, polypeptide, or polypeptide chain, is a term that describes the multiple connected peptide bonds between numerous amino acids. Each amino acid in a polypeptide chain is a unit, commonly known as a residue. These chains have a planar backbone, as the peptide bonds have double bond characteristics due to the existence of resonance between the carbonyl carbon and the nitrogen where the peptide bonds form. The primary structure of each protein has been precisely determined by the specific genes. The C-N bond in an amino acid's chain has the character of a double bond. This bond has a short length and stable. It cannot be rotated. This double-bond character can be explained structurally, in that the R groups in amino acid chains avoid steric clash.
Amino acids are linked by peptide bonds to form polypeptide chain; each amino acid unit is known as a residue; a polypeptide chain constructed by the same unit is known as the main chain or backbone and a changing R group, side chains.
Forces that stabilize Protein Structure
[edit | edit source]Protein structures are governed primarily by hydrophobic effects and by interactions between polar residues and other types of bonds. The hydrophobic effect is the major determination of original protein structure. The aggregation of nonpolar side chains in the interior of a protein is favored by the increase in Entropy of the water molecules that would otherwise form cages around the hydrophobic groups. Hydrophobic side chains give a good indication as to which portions of a polypeptide chain are inside, out of contact with the aqueous solvent. Hydrogen bonding is a central feature in protein structure but only make minor contributions to protein stability. Hydrogen bonds fine tune the tertiary structure by selecting the unique structure of a protein from among a relatively small number of hydrophobically stabilized conformations. Disulfide bonding can form within and between polypeptide chains as proteins fold to its native conformation. Metal ions may also function to internally cross link proteins.
Factors that cause denaturing
[edit | edit source]1) Temperature
2) pH
Extreme temperatures will result in the unfolding of a polypeptide chain leading to a change in structure and often a loss of function. If the protein functioned as an enzyme denaturing will cause that protein to lose its enzymatic activity. As the temperature of a solution containing the protein is raised, the extra heat causes twisting and bending of bonds. As proteins begin to denature the secondary structure of the protein is lost and adopts a random coil configuration. Covalent interaction between amino acid side chains such as disulfide bonds are also lost.
At high or low pH levels the protein will denature due to the lose or gain of a proton and, therefore, will lose their charge or become charged, depending on which way the pH is changed and by how much. This will eliminate many of the ionic interactions that were necessary for maintenance of the folded shape of the protein. As a result the change in structure will cause a change or loss of function.
Determination of Primary Structure: Amino Acid Sequencing
[edit | edit source]After the polypeptide has been purified, the composition of the polypeptide should be established. To determine which amino acid and how much of each is present, the entire strand is degraded by amide hydrolysis (6N HCl, 1100C, 24hr) to produce a mixture of all free amino acid residues. The mixture is separated and its composition recorded by amino acid analyzer. The amino acid analyzer establishes the composition of a polypeptide by giving a chromatogram, which records the peaks of each amino acid presents in the sequence. However, the amino acid analyzer can only give the composition of a polypeptide, not the order in which the amino acids are bound to one another.
To determine the amino acid sequence, it usually starts from the determination of the amino terminal of the polypeptide. The procedure is known as Edman degradation, and the reagent employed is phenyl isothiocyanate.

In Edman degradation, the terminal amino group adds to the isothiocyanate reagent to produce a thiourea derivative. Treating with mild acid, the tagged amino acid is turned into a phenylthiohydantoin, and the remainder of polypeptide is unchanged. Since the phenylthiohydantoins of all amino acid are known, the amino terminal of the original polypeptide can be identified easily. However, Edman degradation can only be used to identify the amino end of the polypeptides; therefore, for polypeptides that are made up by hundreds of amino acids, it is not a practical method in general. In addition, multiple degradation rounds will build up impurities which will seriously affect the yield of peptide. High yield means not completely quantitative, and with each step of degradation, incompletely reacted peptide will mix with the new peptide, resulting in a intractable mixture.
In other words, secondary structure refers to the spatial arrangement of amino acid residues that are nearby in the sequence. The alpha helix, and beta strands are elements of secondary structure.
Secondary Structure
[edit | edit source]Secondary structures of proteins are typically very regular in their conformation. They are the spatial arrangements of primary structures. Alpha Helices and Beta Pleated Sheets are two types of regular structures. An interesting bit of information is that certain amino acids making up the polypeptide will actually prefer certain folding structures. The Alpha Helix seems to be the default but due to interactions such as sterics, certain amino acids will prefer to fold into Beta pleated sheets and so on. For example, amino acids such as Valine, Isoleucine, and Threonine all have branching at the beta carbon, this will cause steric clashes in an alpha helix arrangement. Glycine is the smallest amino acid and can fit into all structures so it does not favor the helix formation in particular. Therefore, these amino acids are mostly found where their side chains can fit nicely into the beta configuration.
The structure of polypeptide main chains is mostly of hydrogen-bonding; each residue has a carbonyl group that is a good hydrogen- bond acceptor; nitrogen- hydrogen group, a good hydrogen- bond donor.
Alpha helix look like the outside of structure. + Right hand appeared in right bottom of Rachamanda plot often
+ Left hand (LOOP): rare on the left top of Ramachandran plot
Alpha Helix
[edit | edit source]Structure
[edit | edit source]The general physical properties of an alpha helix are:


- 3.6 residues per turn
- Translation (rise) of 1.5 A
- Rotation of 100 degrees
- Pitch (or height) of 5.4A (1.5A*3.6 residues)

- Screw sense = clockwise (usually) because it would be less sterically hindered
- Inside the helix consist of the coiled backbone and the side chains project outward in helical array
- Hydrogen bonding between the 1st carbonyl to the hydrogen on the 4th amino
- The shorthand drawing of the alpha helix is a ribbon or rod


- Alpha helix falls within quadrant 1 (left-handed helix) and 3 (right-handed helix) in the Ramachandran diagram
Supersecondary Structure of Alpha Helix
[edit | edit source]I. Coiled coil
An alpha coiled coil consists of two or more alpha helices intertwined, creating a stable structure. This structure provides support to tissues and cell, contributing to the cell cytoskeleton and muscle proteins such as myosin and tropomyosin. Alpha keratin consists of heptad repeats (imperfect repeats of 7 amino acid sequences). This facilitates bonding between the two or more helices.
II. Collagen
Collagen is another type of fibrous protein that consists of three helical polypeptide chains. It is the most abundant protein found in mammals, making up a large component of skin, bone, tendon, cartilage, and teeth. Wrinkles are also caused by the degradations of this protein. In the structure of collagen, every third residue in the polypeptide is glycine because it is the only residue that is small enough to fit in the interior position of the superhelical cable. Unlike normal alpha helices, each collagen helix is stabilized by steric repulsion of the pyrrolidine rings of the proline and hydroxyproline residues. However, the three strands intertwined are stabilized by hydrogen bonding.
Alpha Tertiary
[edit | edit source]I. Motifs
Motifs are simple combinations of the secondary structure such as the helix-turn-helix, which consist of two helices separated by a turn. The helix-turn-helix motif are usually found in DNA-binding proteins.
II. Domains
Domains, or compact globulars, consist of multiple motifs.They are polypeptide chains folded into two or more compact regions connected by turns or loops. Their structure is spherical, which is beneficial for the protein because it conserves space. Generally, inside the globular protein consist of hydrophobic amino acids such as leucine, valine, methionine, and phenylalanine. The outside consists of amino acids with hydrophilic tendencies such as aspartate, glutamate, lysine, and arginine. An example of a globular protein is myoglobin, which is the oxygen carrier in muscle. It is an extremely compact molecule made of only alpha helices (70%) except for loops and turns (30%).
Transmembrane and Non-Transmembrane Hydrophobic Helix
[edit | edit source]Studying the topography of transmembrane and non-transmembrane helix have helped answer many questions about membrane protein insertion. Specifically, studying the sequence and lipid dependence of the topography provide insights into post-translational topography changes. Furthermore, studying topography has lead to the design of hydrophobic helices that have biomedical applications. For example, a tumor marker called pHLIP peptide has been designed.
Different tests have been used to show the various effects on the hydrophobic helices. For example, hydrophilic residues such as tryptophan and tyrosine destabilize the transmembrane state. The hydrophilic domains cannot cross the membrane so it blocks any transmembrane and non-transmembrane equilibration. Furthermore, charged ionized residues also destabilize the transmembrane state. Stabilization of the transmembrane is also achieved in helix-helix interaction. Moreover, anionic lipids promote membrane binding of hydrophobic peptides and proteins.
Alpha helices, beta strands, and turns are formed by a regular pattern of hydrogen bonds between the peptide N-H and C=O groups of amino acids that are near one another in the linear sequence. Such folded segments are called secondary structure.
The alpha-helix consists of a single polypeptide chain in which the amino group (N-H) hydrogen bonds to a carboxyl group (C=O) 4 residues away. The alpha - helix is a rod-like structure. The tightly coiled backbone of the chain forms the inner part of the rod and the side chains extend outward in a helical array. This results in a clockwise coiled structure, which is known as a "right handed" screw sense. This folding pattern, along with the beta-pleated sheets were actually proposed by Linus Pauling and Robert Corey half a decade before people could actually see it. Most of the alpha strands are located in the lower left corner or upper right corner of the Ramachandran diagram . Essentially, most of the alpha helices are found in the right-hand helices area. An alpha helix is especially suited for cross-membrane proteins because all of the amino hydrogen and carbonyl oxygen atoms of the peptide backbone can interact to form intrachain hydrogen bonds while its aliphatic side chains can stabilize in hydrophobic environment of cell membrane.
Alanine, leucine and glutamic acid (existed as glutamate at physiological pH) are the most common residues present in alpha-helices.
The alpha-helix content of protein ranges widely, from none to almost 100%.
In general, the alpha helix is the "normal" shape of a polypeptide chain; however, features of certain amino acids disrupt alpha helix formation and instead favor beta strand formation. Amino acids with branching at the beta carbon (i.e. valine, threonine, and isoleucine) are problematic because they crowd the peptide backbone. H-bond accepting/donating groups attached to the beta carbon (i.e. serine, asparagine, and aspartate) can bond with backbone amine and carboxyl groups, again interfering with alpha helix formation.
While individual amino acids may favor one form or another, predicting the 2° structure of even a short (<7 amino acid) peptide strand is only 60-70% accurate. Such variability suggests other factors, like tertiary interactions with amino acids further down the chain, influence the folding into its observed 3° structure.
Beta-strand is:
- Around ʊ = 120° and ϕ = -120°
- You have the angle, and you form the zigzag
The zigzag have the distance between amino acids is 3.5 Angstrom
Beta Pleated Sheet
[edit | edit source]In contrast to the alpha helical structure, Beta Sheets are multiple strands of polypeptides connected to each other through hydrogen bonding in a sheet-like array. Hydrogen bonding occurs between the NH and CO groups between two different strands and not within one strand, as is the case for an alpha helical structure. Due to its often rippled or pleated appearance, this secondary structure conformation has been characterized as the beta pleated sheet. The beta strands can be arranged in a parallel, anti-parallel, or mixed (parallel and anti-parallel) manner.

The anti-parallel configuration is the simplest. The N and C terminals of adjacent polypeptide strands are opposite to one another, meaning the N terminal of one peptide chain is aligned with the C terminal of an adjacent chain. In the anti-parallel configuration, each amino acid is bonded linearly to an amino acid in the adjacent chain.

The parallel arrangement occurs when neighboring polypeptide chains run in the same direction, meaning the N and C terminals of the peptide chains align. As a result, an amino acid cannot bond directly to the complementary amino acid in an adjacent chain as in the anti-parallel configuration. Instead, the amino group from one chain is bonded to a carbonyl group on the adjacent chain. The carbonyl group from the initial chain then hydrogen bonds to an amino group two residues ahead on the adjacent chain. The distortion of the hydrogen bonds in the parallel configuration affects the strength of the hydrogen bond because hydrogen bonds are strongest when they are planar. Therefore, due to this distortion of hydrogen bonds, parallel beta sheets are not as stable as anti-parallel beta sheet (exp: formation of parallel beta sheet with less than 5 residues is very uncommon).
The side chains of beta strands are arranged alternately on opposite sides of the strand. The distance between amino acids in a beta strand is 3.5 Å which is longer in comparison to the 1.5 Å distance in alpha strands. Because of this, beta sheets are more flexible than alpha helices and can be flat and somewhat twisted. The average length of beta sheets in a protein is 6 amino acid residues. The actual length ranges from 2 to 22 residues.

Beta sheets are graphically found in the upper left quadrant of a Ramachandran plot. This corresponds to ψ angles of 0° to 180° and Φ angles of -180° to 0°.

Visual representations in 3D models for beta sheets are traditionally denoted by a flat arrow pointing in the direction of the strand.
Loop is everything, but what is alpha helix and beta-strand does. It is related to secondary structure of protein.
Turn and Loop
[edit | edit source]Polypeptide chains can change direction by making reverse turns and loops. Alpha helices and beta strands are connected by these turns and loops. Most proteins have compact, globular shape owing to reversals in the direction of their polypeptide chains, which allows the polypeptide to create folds back onto itself. In many reverse turns, the CO group of residue i of a polypeptide is hydrogen bonded to the NH group of residue i+3. A turn helps to stabilize abrupt directional changes in the polypeptide chain. Loops are more elaborate chain reversal structures that are rigid and well defined. Loops and turns generally lie on the surfaces of proteins so they often participate in interactions between proteins and other molecules. In a loop, there are no regular structures as can be found in helices or beta strands.
Two hypotheses have been proposed for the role of turns in protein folding. In one view, turns play a critical role in folding by bringing together interactions between regular secondary structure elements. This view is supported by mutagenesis studies indicating a critical role for particular residues in the turns of some proteins. Also, nonnative isomers of X-Proline peptide bonds in turns can completely block the conformational folding of some proteins. In the opposing view, turns play a passive role in folding. This view is supported by the poor amino-acid conservation observed in most turns. Also, non-native isomers of many X-Pro peptide bonds in turns have little or no effect on folding.
Beta Hairpin Turns
[edit | edit source]A motif is when secondary structure elements combine in specific geometric arrangements. Beta hairpin turns are one type of arrangement; they are one of the simplest structures and then are found in globular proteins. Upon turning, the antiparallel strand can bind effectively through hydrogen bonding between the carbonyl carbon and the peptide backbone nitrogen. It has been shown that 70% of beta-hairpins are less than seven residues long; the majority being 2 residues long. There are two types of two-residue beta hairpin turns. The first, Type I, forms a left-handed alpha-helical conformation. This left-handed conformation has a positive phi angle due to the properties of the aforementioned amino acids. Glycine does not have a side chain to sterically interfere with the turned amino acid sequence. Asparagine and aspartate both readily form hydrogen bonds with the carbonyl oxygen as a hydrogen bond acceptor. The second amino acid in the Type I turn is usually glycine due to steric hindrance that would result using any amino acid with a side chain. In a Type II beta hairpin turn, the first residue can only be glycine due to steric hindrance. However, the second residue is usually polar, such as serine or threonine.
Fibrous proteins
[edit | edit source]Fibrous protein such as alpha-keratin and collagen consist of two right handed alpha helix intertwined to form a type of left handed super-helix called an alpha coiled coil. The two helices in this type of protein usually cross-linked by weak interaction such as Van der Waals forces force and ionic interaction. The side chain interaction can be repeat every seven residues, forming heptad repeats. Another form of fibrous protein, that of collagen, exists as three helical polypeptide chains. These chains are relatively long, ~1000 residues, and because of overcrowding, glycine appears once every three residues. While the helix is stabilized by the steric repulsions, the three strands are stabilized by hydrogen bonding. These protein usually serve structural roles in organisms, alpha-keratin is commonly found in the cytoskeleton of a cell, as well as certain muscle proteins. Collagen is often found in teeth, skin, and tendons.
Secondary Structure Prediction
[edit | edit source]The science of predicting what polypeptide chain will conform to which secondary structure group (alpha-helix, beta-sheet/strand or turns/loops) is not particularly exact. However, various frequencies of secondary structure formation of certain amino acids have been recorded in actual scientific experimentation, and these values can allow scientists to predict the folding of a protein based on its amino acid composition with about 60-70% accuracy. Stretches of six or less residues can usually be predicted with this accuracy. Although, certain amino acids tend to fold in its preferred conformation, there are of course exceptions and so secondary structure prediction is not always accurate. Tertiary interactions, interactions with residues further apart from each other, can also determine the folding structures. Each amino acid has a preference for either secondary structure, but it normally is only a small preference towards one in comparison to another, therefore, this unfortunately does not mean much. Amino acids can appear in an alpha-helix in one protein and also in a beta-sheet in another. Due to the unpredictability of the secondary structure based on the sequence of amino acids, secondary structures are being analyzed and predicted in relations to a similar family of sequences.
Various techniques have risen throughout history in the study of secondary structural prediction. With the aid of computers, prediction has been a pursued research topic in bioinformatics and many approaches continue to be proposed. After Linus Pauling and Robert Corey discovered the periodic alpha helix and beta sheet structures within proteins in 1951, further elucidation of protein structure prediction began to grow. A major method in secondary structure prediction was the Chou-Fasman method; it yielded a 50-60% accuracy. This method based its predictions on assigning a set of prediction values to a certain amino acid residue and then applied an algorithm to that value. Shortly after, further improvements were made on this method, the GOR method, which was developed in the late 1970s and utilized information theory|entropy and information concepts for secondary structure prediction. When devised, the method was about 65% accurate, however, improvements have also been made to it. There are deductive techniques in which similar sequences are found in already identified proteins. This method is accomplished by having computer software search databases of identified proteins. Opposite of that would be the Ab initio method, which builds 3-dimensional models without looking at similar residue sequences. This method is based on hydrogen bonding principals and localization.
Other methods and factors of folding prediction include analyzing the basic chemical tendencies of the side chains of amino acids to determine its preference in secondary structure. The alpha-helix is taken as the default structure, thus amino acids that destabilize alpha-helices are often found in beta-pleated sheets or loops and turns. For instance, valine, threonine, and isoleucine will often destabilize the helix because of branching of the beta carbon. These three amino acid residues are more often found in beta-pleated sheets, where their side chains will lie in a separate plane than the main chain. There are also amino acid residues that prefer neither alpha-helices nor beta-pleated sheets, for example, Proline has a restricted phi angle of ~60° degrees and no NH group, all due to the fact that it is cyclic. This will disrupt both alpha-helices and beta-pleated sheets, thus is found mostly in loops and turns. A counter-intuitive example is glycine which, according to its small size, theoretically can fit in any structure easily, but in reality it tends to avoid alpha-helices and beta-sheets also. The folding definitely also relies on chemical interactions between the side chains so the surrounding amino group interactions also affect the tendency of folding. These tendencies are reflected in the frequencies of secondary structure for individual amino acids.
The relative tendencies of secondary structures for particular amino acids are listed below:
alpha-helix: Glu, Ala, Leu, Met, Lys, Arg, Gln, His
beta-sheet: Val, Ile, Tyr, Cys, Trp, Phe, Thr
turns and loops: Gly, Asn, Asp, Pro, Ser
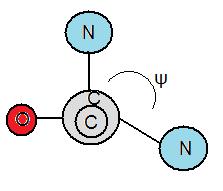
Torsion Angles
[edit | edit source]Torsion angles are also called dihedral angles. The torsion angle is the measure in degrees in bonds between atoms. Folding of proteins are influenced by the degree of rotation amino bonds can hold. There are two different types of torsion angles existing in polypeptide bonds. Phi, φ is the angle between the α-carbon and the nitrogen atom of a peptide bond. The other bond is called psi, ψ which is the angle between the α-carbon and the carbonyl group. To measure φ, one must look from the nitrogen atom towards the α-carbon to measure if the angle is negative or positive. The angle is negative if the α-carbon rotates counterclockwise and vice versa. Furthermore, to measure ψ, one must look from the nitrogen atom towards the carbonyl group. Likewise, the angle is negative if the carbonyl group rotates counterclockwise and vice versa.
Ramachandran Diagram
[edit | edit source]The Ramachandran Diagram, created by Gopalasamudram Ramachandran, helps to determine if amino acids will form alpha helices, beta strands, loops or turns. The Ramachandran Diagram is separated into four quadrants, with angle ϕ as the x axis and angle ψ as the y-axis. The combinations of torsion angles will put the amino acids in specific quadrants, which determine whether it will form an alpha helix, beta strand, loop, or turn. Those that fall in quadrants 1 and 3 a few times in a row form alpha helices, and those that repeat in quadrant 2 form beta strands. Quadrant 4 is generally disfavored because of steric hindrance. Also, it is mostly impossible because the different torsion angles combinations in quadrant 4 can't exist because they cause collisions between the atoms of the amino acids. If the amino acids land in the different quadrants, with no repeats, then they become loops or turns. Furthermore, the principle of steric exclusion states that two atoms cannot occupy the same place simultaneously.
Myoglobin is one of example of tertiary structure. Myoglobin is an extremely compact molecule. It is oxygen carrier in muscle is a single polypeptide chain of 153 amino acids. The capacity of myoglobin to bind oxygen depends on the presence of HEME, a non polypeptide PROSTHETIC group consisting of protoporphyrin IX and a central iron atom.
Tertiary Structure
[edit | edit source]The tertiary structure of a protein is the three-dimensional structure of the protein. This three-dimensional structure is mostly determined by the amino acid sequence, which is denoted by the primary structure of the protein, however the amino acid sequence cannot entirely predict on how the three-dimensional structure is formed. Another contributing factor to the final shape of the tertiary structure is based on the environment in which the protein is synthesized. The tertiary structure is stabilized by the sequence of hydrophobic amino acid residues in the backbone of the protein. The interior consists on hydrophobic side chains while the surface consists of hydrophilic amino acids that interact with the aqueous environment.
Tertiary structure is formed by interactions between side chains of various amino acids - in particular disulfide bonds formed between two cysteine groups. At this stage, some proteins are complete, while other proteins incorporate multiple polypeptides subunits which creates the quaternary structure.
Nucleation-condensation model. The tertiary folding process is very structured with key intermediates. When a protein starts to fold, localized areas of the protein first begin folding. Then, the individual localized folds come together to complete the tertiary structure. The key concept is that when a correct fold is achieved, that fold is retained until all other parts of the protein are also correctly folded. This folding process follows reason because a random trial and error folding process would not only take much more time to complete, but also would require much more input energy.
Tertiary structure refers to the spatial arrangement of amino acid residues that are far apart in the sequence and to the pattern of disulfide bonds. Tertiary structure is also the most important protein structure that is used in determining the enzymatic activity of proteins.
Structure
[edit | edit source]

Cysteine, an amino acid containing a thiol group, is responsible for the disulfide bonds that hold a tertiary structure together. In the tertiary structure, when two helices come together, they may be linked by these disulfide bonds. A tertiary structure with fewer disulfide bonds form less rigid structures that are flexible, but still strong and can resist breakage such as hair and wool. While tertiary structures that contain more crossed disulfide bonds, formed by cysteine residues, lead to stronger, stiffer and harder structures such has exoskeletons. Others examples of protein that contain more disulfide bonds include claws, nails, and horns.
A structure made of two a-helices such as keratin can be found in living organisms. Immunoglobulin, also known as antibodies, is an example of an all beta-sheet protein fold. It consists of approximately 7 anti-parallel beta-strands arranged in 2 beta-sheets. For instance, if a cysteine is mutated to another amino acid it can code to a different protein which would lead to incorrect folding.
Domains
[edit | edit source]Some polypeptide chains fold into several compact regions. These regions in a polypeptide chain are called domains and generally range from 30 to 400 amino acids. On average, domains contain roughly 100 amino acids. Each domain forms its own tertiary structure which contributes to the overall tertiary structure of the protein. These domains are independently stable. Stabilization is caused by metal ions or disulfide bridges that cause the folding of polypeptide chains. Different proteins may have the same domains even if the overall tertiary structure is different.
There are four types of domains:
- All-α domains - Domains made purely from α-helices.
- All-β domains - Domains made purely from β-sheets.
- α+β domains - Domains made both of α-helices and β-sheets.
- α/β domains - Domains made from both α-helices and β-sheets layered in a β,α,β fashion with a α-helix sandwiched in between 2 β-sheets.
Mutations
[edit | edit source]In order for a protein to be functional (except in food), it must have an intact tertiary structure. If a tertiary structure of a protein is disrupted, it is said to be denatured. Once a protein is denatured, it will not be able to perform its intended or original function. A primary cause for an alteration of the tertiary structure is a mutation in the gene encoding a protein. The mutation in the gene can cause a domino effect that will lead to the degradation of the tertiary structure. Degradation can cause several diseases, one of which is called cystic fibrosis. Cystic fibrosis is brought about by a mutation of a genes called cystic fibrosis transmembrane conductance regulator (CFTR). This disease causes the exocrine glands to overproduce mucus. Most commonly, CF patients suffer from lung failure by the age of early 20-30. Diabetes insipidus, familial hypercholesterolemia, and Osteogenesis imperfecta are also diseases that originate from degraded proteins. A mutation in the tertiary structure itself, rather than from a mutation in the nucleotide sequence can also lead to diseases. Such mutated proteins can also aggregate and become insoluble deposits called amyloids, and therefore lose the ability to function. A common mutation is when a hydrophobic R group folds in, rather than out, in a hydrophobic environment. The inherited form of Alzheimer's disease is one disease that is caused by mutated tertiary structure. Another disease includes mad cow disease, which is caused due to a-helix (which are soluble) mutating into b-sheets (which are insoluble and cause amyloid deposits). [1]
Folding
[edit | edit source]The folding of a protein is dependent on the amino acid sequence laid out in the primary structure. It is also dependent on the environment in which the folding occurs. In a hydrophobic environment, the hydrophobic side chains of the amino acids of the protein fold out while the hydrophilic side chains fold in and vice versa for a hydrophilic environment. An example of a protein that is folded in a hydrophobic environment is Porin. Its hydrophilic side chains are folded in which creates a channel for water to pass through. Amino acids that have nonpolar/hydrophobic side chains such as leucine, valine, methionine, phenylalanine, and isoleucine would be folded out in the folding of the protein in a hydrophobic environment. Likewise, in a hydrophilic environment, amino acids with polar side chains such as glutamine and asparagine fold outwards and the hydrophobic side chains would fold inwards.
Determination of Tertiary Structure
[edit | edit source]The tertiary structure of a protein is determined through X-Ray Crystallography and Nuclear Magnetic Resonance (NMR) Spectroscopy. X-ray Crystallography was the first method used to determine the structure of proteins. X-ray crystallography is one of the best methods because the wavelength of an x-ray is similar to that of covalent bonds found throughout proteins, creating a clearer visualization of a molecule's structure. The scattering of x-rays by electrons is analyzed to determine the structure of proteins. In order to use x-ray crystallography, the protein in question must be in crystal form. Some proteins crystallize readily, while others do not. For those proteins that do not crystallize readily, nuclear magnetic resonance (NMR) spectroscopy must be used to determine its structure. NMR spectroscopy uses the spin of nuclei with a magnetic dipole and chemical shifts to determine a molecule’s relative position.
Hemoglobin is one of example of quaternary structure. Hemoglobin, the oxygen-carrying protein in blood, consists of two subunits of one type (designated alpha) and two subunits of another (designated beta).
Quaternary Structure
[edit | edit source]
A quaternary structure refers to two or more polypeptide chains held together by intermolecular interactions to form a multi-subunit complex. The interactions that hold together these folded protein molecules include disulfide bridges, hydrogen bonding, hydrogen bonding interactions, hydrophobic interactions interactions and London forces. These forces are usually conveyed by the side chains of the peptides.
These polypeptide chains are the subunits of a protein, capable of taking part in a variety of functions such as serving as enzymatic catalysts, providing structural support in the cytoskeletons of cells, and even composing the hair on our heads.
The peptides of the protein can be identical or different. Insulin is a dimer consisting of two identical peptides, while Hemoglobin is a tetramer consisting of two identical alpha subunits and two identical beta subunits.
Naming Quaternary Structures
[edit | edit source]In naming quaternary structures, the number of subunits (tertiary structure) and the suffix -mer (Greek for "part, subunit")are used:
- 1 subunit = Monomer
- 2 subunits = Dimer
- 3 subunits = Trimer (These are sometimes viewed as cyclic trimers. For example: aliphatic and cyanic acids)
- 4 subunits = Tetramer
The pattern continues with pent-, hex-, hept-, oct-, and so forth.
Dimers
[edit | edit source]
- Insulin
- Dimer – alpha chain and beta chain
- Linked by 2 disulfide bridges
- HIV Protease
- Dimer
- Composed of identical subunits
Trimer
[edit | edit source]- Collagen
- Composed of 3 helical polypeptide chains
- Glycine appears at every third residue because there is no space in center of the helix
- Stabilized by steric repulsion of the pyrrolidine rings of the proline and hydroxyproline residues
- Hydrogen bonds hold together the strands of the collagen fibers
Tetramer
[edit | edit source]
- Hemoglobin
- Consists of 2 alpha and 2 beta groups
- Has a globular shape
- Has reverse turns that contribute to circular shape of the protein
- Aquaporin
- Made of 6 alpha helices
- Form hydrophobic loops
- Forms tetramers in the cell membrane with each monomer acting as water channels
Breaking Apart the Quaternary Structure
[edit | edit source]The quaternary structure of a protein can be denatured by breaking the covalent and non-covalent forces that keep it together. Heat, urea or guanidinium chloride will denature a protein by disrupting the non-covalent forces, while beta-mercaptoethanol will break disulfide bridges by reducing the bridges.
Protein Folding
[edit | edit source]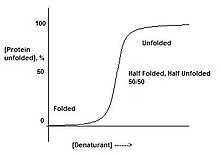
Proteins are either folded, or not. There does not exist a stage where a protein is "half-folded". This can be observed by slowly adding denaturant to a protein. This will result in a sharp transition, from the folded state to the unfolded state, suggesting there only exist these two forms. This is a result of cooperative transition.
For instance, if a protein is put in a denaturant where only one part of the protein is unstable, the entire protein will unfold. This is due to the domino effect where destabilizing one part of the protein will in turn destabilize the remainder of the structure. When a protein is in conditions which correspond to the middle of the transition between folded and unfolded, there is a 50/50 mixture of folded and unfolded protein, instead of 'half-folded' protein.
After all is said about being in one structure or the other, there must be something in between them on an atomic level. Unfortunately, this is an area that is still under development, and much research is still being done. Theories such as the condensation Nucleation Principle are concerned with this area of protein folding.
The properties of quaternary structure:
- Polypeptide chains can assemble into multisubunit structure
- Refers to the spatial arrangement of subunits and the nature of their interactions
Analogy
[edit | edit source]If one takes each student in a class to be a different amino acid, each right hand to be an alpha-carboxyl group, each left hand to be an alpha-amino group, and the head to be the R group; then by joining right hands to left hands, the class will form a polypeptide. The "bonds" joining the hands will be peptide bonds. This can be considered the primary structure of a protein.
If one then takes students and "attract" them to other students 4 "bonds" away, this structure will then fold into a secondary structure; namely the alpha-helix. If the students were put into lines and were attracted to respective students in another line, they would form a beta-pleated sheet.
Now imagine that the heads, or R groups, vary in areas such as personalities, or polarity, like will attract like. The people who are more compatible will then gather together, for instance, hydrophobic areas will usually gather together in the center while surrounded by hydrophilic areas. This makes up the tertiary structure.
Now add in a different class, the people from the new class would have their own tertiary structure, these new people will then come in and react with the original class to form quaternary structures.
Human attempt to manipulate protein assemblies (Quaternary Structures)
[edit | edit source]Controlling the quaternary structures is currently catching more and more interest in academics. There are many advantages in manipulating protein assemblies. Firstly, people are able to grow/synthesize enzymes that are beneficial to human. Yet, to get these enzymes to work is the hard part. For example, nitrogenase, the enzyme that can fix nitrogen gas to yield ammonia, can only work under aerobic environment and coupled with ATP as energy source. In addition, researchers have revealed that nitrogenase is compose of two proteins, one for ATP coupling electron source and the other is the reactive center for nitrogen fixation. The two protein assemble to work as a whole. Recently, scientists remove the ATP coupling protein and replace it with a Ruthenium complex. It turned out that Ruthenium complex can provide electrons with light exposure. Now scientists don't have to deal with the complicate chemistry of coupling ATP, but just shine lights on engineered nitrogenase to get it work! Secondly, protein assemblies can have a lot of clinical/material applications. Ferritin is a family of high-order protein assembly family, usually 12mers or 24mers. Previous researches showed it can absorb large amount of Fe ion. Many researchers are working to control the association and dissociation of Ferritins, seeking for solutions of drug delivery, gas storage, metal harvest and etc. Many approaches have been developed to control protein assembling. Some of them include the following:
1. Transition metal-directed. Metal centers in protein are important, not only because they are reactive centers, but also they help stabilize the shape of protein by coordination. Many amino acids are ligands by themselves. Cysteine, Histidine, lysine are the common ones. Plus, researchers can engineer inorganic ligands onto proteins by cysteine substitution. Thus, introducing inorganic ligands much broaden the horizon of protein assemblies.


Metal-ligand bonding has several properties. Most obviously, it is a strong interaction. It is stronger than hydrogen bond and weaker than covalent bond. Therefore metal-ligand bond is strong yet not so strong that it is still reversible. Spatially speaking, metals have its coordination orientation, mostly, octahedral and tetrahedral. This property provides human great convenience in arranging proteins spatially.


2. Hydrophobic interaction. In aqueous environment, amino acid with hydrophobic side chains tend to aggregate together to minimize the exposure to water. Researchers utilize this character and engineer certain matching pair of non-polar amino acids onto proteins to obtain protein oligomers in water solution.
3. Salt bridges. It is well known that amino acids have different pI's. So at certain pH, some amino acids are negatively charged, some are positively charged. If an area on a protein is occupied by mostly negatively charged amino acid and another area is occupied by positively charged amino acids, proteins can aggregate by electrostatic attraction. However, this technique is usually not so selective.
More technique to direct protein assemblies are being investigated, such as coiled-coil. Mankind's potential to control quaternary structures is promising.