GENtle/Sequence editor

The sequence editor holds the key to several properties of a sequence. It consists of several tabs, depending on the type of sequence, which can be DNA or amino acid.
Properties
[edit | edit source]Here, the title and description of the sequence can be altered. As for feature descriptions, the sequence description will make http references clickable.
For DNA sequences, sticky ends can be entered.
Features
[edit | edit source]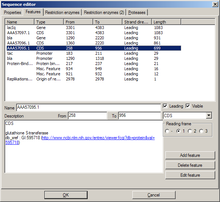 |
 |
This tab shows a list of all features of the sequence. Features can be added, edited, and deleted. Most of the settings should be self-explanatory.
- The setting reading frame is only available when the type is set to "CDS" ("coding sequence").
- A leading sequence is read 5'→3'; leading unchecked, 3'→5'
- Edit feature will invoke an additional "Edit feature" dialog
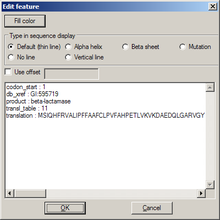
Edit feature
[edit | edit source]- Fill color is the color of the feature; it will invoke a color choice dialog
- Type in sequence display determines how that feature is drawn in the sequence map
- Use offset sets the numbering for the first amino acid of the feature; useful if the feature marks a part of a protein
The list box below contains original data from GenBank format import.
Restriction enzymes
[edit | edit source]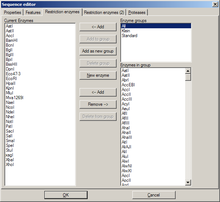 |
 |
When editing a DNA sequence, two tabs with settings for restriction enzymes are available. The first one is identical to the enzyme management dialog. The second one is identical to the global enzyme settings tab, but contains the settings for this sequence alone. By default, its options are disabled, and the global options are used. By activation the options here, global settings are overridden.
Proteases
[edit | edit source]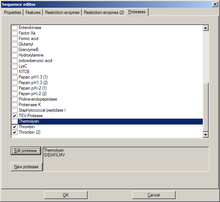
This tab holds a list of available proteases. Potential cleavage sites for selected (checked) proteases are shown in the sequence (not in the DNA map).
New proteases can be added similar to the following examples:
- Example : "Thermolysin"
- Sequence for this protease : "!DE|AFILMV"
- Explanation: "Not D or E", "cut", "then A, F, I, L, M, or V"
- Example: Proline-endopeptidase
- Sequence for this protease : "HKR,P|!P"
- Explanation : "H, K, or R", "then P", "cut", "then not P"