GENtle/PCR and Primer Design
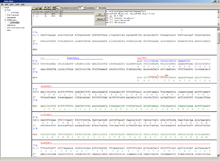
This module allows for designing primers and running virtual PCRs. It can be started from a DNA module via context menu of the DNA or [../Sequence map#Context menu|sequence]] map, or through Tools/PCR. If a sequence is selected in the DNA module, one or more primers can be generated automatically upon startup of the PCR module. These will only be rough suggestions, and are in no way optimized by default.
Toolbar
[edit | edit source]- Enter new primer
- Open primer/sequence
- Print PCR
- Add a primer (you will have to open or enter the primer first)
- Export a primer (generate its sequence)
- Edit mode
- Show/hide features
- Polymerase running length
- Horizontal mode
The polymerase running length is the number of nucleotides the polymerase is allowed to run during the PCR in the elongation step. This is usually measured in minutes, but each polymerase runs at a different speed, which is why this information is given here in nucleotides. The value is initially computed automatically, but can be changed manually.
Primer list
[edit | edit source]The primer list (the upper left) shows all primers used in this PCR, as well as certain key properties of these. Selecting one of these primers will show more detailed information in the box on the right (see here for details). Double-clicking one of the primers will mark and show that primer in the sequence. A selected primer can be removed through the Remove button, or edited via the Edit button. A selected primer can also be exported via the Export button in the toolbar; a new sequence will be generated for that primer.
- Caveat : The generated sequence is not stored anywhere automatically, it needs to be saved manually!
- Caveat : To add a primer, use the Add button in the toolbar, or the Selection as new primer context menu. Merely editing the sequence (see below) is for editing existing primers only, it will not create new ones!
Sequence
[edit | edit source]The sequence consists of the following lines:
- Features of the template DNA (can be turned off in the toolbar)
- 5' primer
- Template DNA (5'→3')
- Amino acid sequence of the template
- Template DNA (3'→5')
- 3' primer
- Restriction sites of the resulting DNA
- Resulting DNA (shown in green)
- Amino acid sequence of the resulting DNA
Some special functions and properties of the PCR sequence display:
- The amino acid reading frame can be set as described here. This will affect both amino acid sequences shown (template and result).
- Only the two primer sequences can be edited; overwrite mode is mandatory, and deleting is disabled.
- To delete a nucleotide, overwrite it with Space.
- The "." key will copy the matching template nucleotide to that position in the primer sequence.
- Matching primer nucleotides (that is, matching with the template) are shown in blue, mismatches in red.
- If (when not in edit mode) an empty span of the primer sequence is selected, it can be turned into a new primer via the context menu (Selection as new primer).
- The sequence of a restriction site can be inserted left or right of a selection (in edit mode, right or left of the cursor) via the context menu. A selection dialog for the desired enzyme will appear.
- A silent mutation can be introduced via the context menu.
Finally, the resulting DNA or amino acid sequence (the green sequence, which will be the one generated by the PCR) can be copied to the clipboard or generated as a new sequence (containing all features, restriction enzymes etc.) via the context menu.