Structural Biochemistry/Protein function/Binding Sites
Protein Properties: Binding Sites
[edit | edit source]A binding site is a position on a protein that binds to an incoming molecule that is smaller in size comparatively, called ligand.
In proteins, binding sites are small pockets on the tertiary structure where ligands bind to it using weak forces (non-covalent bonding). Only a few residues actually participate in binding the ligand while the other residues in the protein act as a framework to provide correct conformation and orientation. Most binding sites are concave, but convex and flat shapes are also found.
A ligand-binding site is a place of chemical specificity and affinity on protein that binds or forms chemical bonds with other molecules and ions or protein ligands. The affinity of the binding of a protein and a ligand is a chemically attractive force between the protein and ligand. As such, there can be competition between different ligands for the same binding site of proteins, and the chemical reaction will result in an equilibrium state between bonding and non-bonding ligands. The saturation of the binding site is defined as the total number of binding sites that are occupied by ligands per unit time. High affinity ligands have a high intermolecular force, are able to reside in the binding site longer since it has a low concentration, and cause the receptors to change.
The most common model of enzymatic binding sites is the induced fit model. It differs from the more simple "Lock & Key" school of thought because the induced fit model states that the substrate of an enzyme does not fit perfectly into the binding site. With the "lock & key" model, it was assumed that the substrate is a relatively static model that does not change its conformation and simply binds to the active site perfectly. According to the induced fit model, the binding site of an enzyme is complimentary to the transition state of the substrate in question, not the normal substrate state. The enzyme stabilizes this transition state by having its NH3+ residues stabilize the negative charge of the transition state substrate. This results in a dramatic decrease in the activation energy required to bring forth the intended reaction. The substrate is then converted to its product(s) by having the reaction go to equilibrium quicker.
Biological Importance of Proteins
[edit | edit source]Proteins are important in our human body. A protein is a chain of amino acids joined by peptide bonds. By understanding protein function, we can solve different problems in our body because proteins are among the main biological components. In order to understand protein functions, there will be four main ideas to study, such as: structure, binding, catalysis, and switching. However, these ideas will be much easier to understand if we know about protein binding. Protein binding has an extremely important role in biochemistry. Protein binding is often reversible and can be stable or unstable depend on the structures and the activities of it. Furthermore, that is also a reason why protein binding can be influence the drug's biological half life in our body. Indeed, many scientists try to perform experiments and research on the binding structure in order to discover diseases and how to destroy it.
Protein binding and the fraction unbound written as the concentration of unbound drug over the total concentration of the drug, which depends on several factors. It is determined by the drug’s affinity for the protein, the concentration of the binding protein, and the concentration of the drug relative to the binding protein.
Protein binding also is very crucial for the hemoglobin and myoglobin. Hemoglobin is involved in the transport of other gases. Also, it carries some of the body's respiratory carbon dioxide, in which CO2 is bound to the globin protein. Hemoglobin exhibits characteristics of both the tertiary and quaternary structures of proteins. Hemoglobin consists mostly of protein, and these proteins, in turn, are composed of sequences of amino acids. Myoglobin is found in the muscle tissues and it was the first protein to have its three-dimensional structure revealed. Even though hemoglobin is found in the blood and myoglobin is found in the muscles, they are have a great connection with each other through functioning and deliver and transport oxygen.
Properties that Affect Binding
[edit | edit source]Complementarity Molecular recognition depends on the tertiary structure of the enzyme which creates unique microenvironments in the active/binding sites. These specialized microenvironments contribute to binding site catalysis.
Flexibility Tertiary structure allows proteins to adapt to their ligands (induced fit) and is essential for the vast diversity of biochemical functions (degrees of flexibility varies by function). Flexibility is essential for biochemical function.
Surfaces Binding sites can be concave, convex, or flat. For small ligands – clefts, pockets, or cavities. Catalytic sites are often at domain and subunit interfaces. Catalytic sites often occur at domain and subunit interfaces.
Non-Covalent Forces Non-covalent forces are also characteristic properties of binding sites. Such characteristics are: higher than average amounts of exposed hydrophobic surface, (small molecules – partly concave and hydrophobic), and displacement of water can drive binding events, and weak interaction can lead to an easy exchange for partners.
Affinity Binding ability of the enzyme to the substrate (can be graphed as partial pressure increases of the substrate against the affinity increases).
Enzyme Inhibitors
[edit | edit source]Overview
[edit | edit source]Enzyme inhibitors are molecules or compounds that bind to enzymes and result in a decrease in their activity. An inhibitor can bind to an enzyme and stop a substrate from entering the enzyme's active site and/or prevent the enzyme from catalyzing a chemical reaction. There are two categories of inhibitors.
- irreversible inhibitors
- reversible inhibitors
Inhibitors can also be present naturally and can be involved in metabolism regulation. For example, negative feedback caused by inhibitors can help maintain homeostasis in a cell. Other cellular enzyme inhibitors include proteins that specifically bind to and inhibit an enzyme target. This is useful in eliminating harmful enzymes such as proteases and nucleases. In addition, many different man made substances are capable of inhibiting enzymes. For example, fluorinated phosphonates (i.e. sarin, VX) bind irreversibly to the enzyme cholinesterase resulting in a buildup of acetylcholine in the body. Moreover, drugs, such as MAOIs (monoamine oxidase inhibitors) bind reversibly to the enzyme monoamine oxidase allowing for the accumulation of monoamine neurotransmitters, which is useful in the treatment of certain medical conditions.
competitive inhibition, the inhibitor and substrate compete for the enzyme (i.e., they can not bind at the same time).[58] Often competitive inhibitors strongly resemble the real substrate of the enzyme. For example, methotrexate is a competitive inhibitor of the enzyme dihydrofolate reductase, which catalyzes the reduction of dihydrofolate to tetrahydrofolate. The similarity between the structures of folic acid and this drug are shown in the figure to the right bottom. Note that binding of the inhibitor need not be to the substrate binding site (as frequently stated), if binding of the inhibitor changes the conformation of the enzyme to prevent substrate binding and vice versa. In competitive inhibition the maximal velocity of the reaction is not changed, but higher substrate concentrations are required to reach a given velocity, increasing the apparent Km.
Irreversible Inhibitors
[edit | edit source]Irreversible inhibitors covalently bind to an enzyme, cause chemical changes to the active sites of enzymes, and cannot be reversed. A main role of irreversible inhibitors include modifying key amino acid residues needed for enzymatic activity. They often contain reactive functional groups such as aldehydes, alkenes, or phenyl sulphonates. These electrophilic groups are able to react with amino acid side chains to form covalent adducts. The amino acid components are residues containing nucleophilic side chains such as hydroxyl or sulfhydryl groups such as amino acids serine, cysteine, threonine, or tyrosine.
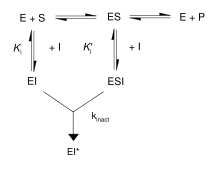
First, irreversible inhibitors form a reversible non-covalent complex with the enzyme (EI or ESI). Then, this complex reacts to produce the covalently modified irreversible comple EI*. The rate at which EI* is formed is called the inactivation rate or kinact. Binding of irreversible inhibitors can be prevented by competition with either substrate or a second, reversible inhibitor since formation of EI may compete with ES.
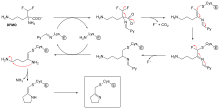
In addition, some reversible inhibitors can form irreversible products by binding so tightly to their target enzyme. These tightly-binding inhibitors show kinetics similar to covalent irreversible inhibitors. As shown in the figure, these inhibitors rapidly bind to the enzyme in a low-affinity EI complex and then undergoes a slower rearrangement to a very tightly bound EI* complex. This kinetic behavior is called slow-binding. Slow-binding often involves a conformational change as the enzyme "clams down" around the inhibitor molecule. It changes shape slightly to accommodate the substrate enough to allow the reaction to kinetically proceed forward. Some examples of these slow-binding inhibitors include important drugs such as methotrexate and allopurinol.
Group Specific Reagents
Group specific reagents inactivate an enzyme by reacting with a certain side chain on an amino acid. One example is Iodoacetamide reacting with a cysteine side chain to completely inactivate the enzyme and creating I- and H+ ions along the way. Another example is diisopropylphophofluoridate (DIPF)inactivating enzymes by reacting with a crucial serine side chain in acetylcholinesterase.
Reactive Substrate Analogs (Affinity Labels)
Affinity labels are specific to the binding site on the enzyme. They covalently bind to active-sites in place of the substrate and thus modify a crucial side-chain and inhibit the activity of the enzyme. For example, bromoacetol phosphate binds to triose phosphate isomerase (TPI) at its active site. The covalent bonding inhibits the enzymatic activity since the active side chain (i.e. glutamic acid) is not active.
Mechanism Based Inhibitors (Suicide Inhibitors)
Mechanism based inhibitors affect the enzyme after the initial substrate has already been bound to an enzyme and has undergone a normal catalytic mechanism. The catalysis generally generates an intermediate that covalently inhibits the enzyme.
Reversible Inhibitors
[edit | edit source]Reversible inhibitors bind non-covalently to enzymes. Many different types of inhibitions can occur depending on the structure of the enzyme the inhibitors bind to. The non-covalent interactions between the inhibitors and enzymes include hydrogen bonds, hydrophobic interactions, and ionic bonds. Many of these weak bonds combine to produce strong and specific binding. In contrast to substrates and irreversible inhibitors, reversible inhibitors generally do not undergo chemical reactions when bound to the enzyme and can be easily removed by dilution or dialysis.
There are three kinds of reversible inhibitors:
1) competitive
2) mixed
3) uncompetitive/mixed
- Competitive inhibitors, as the name suggests, compete with substrates to bind to the enzyme at the same time. The inhibitor has an affinity for the active site of an enzyme where the substrate also binds to. This type of inhibition can be overcome by increasing the concentrations of substrate in order to overcome the inhibitor. Competitive inhibitors are often similar in structure to the substrate. These inhibitors increase the Km.File:Competitive inhibitor.docx
Oxyanion hole stabilizes the tetrahedral intermediate. It is formed by hydrogen bonds linking peptide NH groups to the negatively charged oxygen atom.
Step 3: Instability of the negative charge on the substrate carbonyl oxygen when will leads to collapse of the tetrahedral intermediate, re-formation of a double bond with carbon which breaks the peptide bond between the carbon and amino acid group. The amino leaving group is protonated by His57, facilitating its displacement.
Step 4: The amine component is departed from the enzyme (metabolized by the body) and this completes the first stage (acylation of enzyme). The first product is been made.
Step 5: A water molecule is added.
Step 6: An incoming water molecule is deprotonated by acid-base catalysis, generating a strongly nucleophilic hydroxide ion. Attack of hydroxide on the ester linkage of the acylenzyme generates a second tetrahedral intermediate.
Step 7: collapse of the tetrahedral intermediate form the second product, a carboxylate anion, and displace Ser195.
Step 8: The carboxylic acid is released and the enzyme is reformed to catalyze the next reaction.
- Uncompetitive inhibitors bind to the enzyme at the same time as the enzyme's substrate. However, the binding of the inhibitor affects the binding of the substrate, and vice-versa. This type of inhibition cannot be overcome, but can be reduced by increasing the concentrations of substrate. The inhibitor usually follows an allosteric effect where it binds to a different site on the enzyme than the substrate. This binding to an allosteric site changes the conformation of the enzyme so that the affinity of the substrate for the active site is reduced. These inhibitors lower the K mand Vm.File:Uncompetitive inhibition.docx
- mixed inhibition the inhibitors can bind to the enzyme at the same time as the enzyme substrate. If the concentrate of substrates is higher than the inhibitor, then this type of inhibition can be reduced.File:Mixed inhibition.docx
- Uncompetitive inhibitors are able to bind to both E and ES, but their affinities for these two forms of the enzyme are different. Therefore, these inhibitors increase Km and decrease Vmax because they interfere with substrate binding and hamper catalysis in the ES complex.
- Non-competitive inhibitors have identical affinities for E and ES. They do not change Km, but decreases Vmax.
