Structural Biochemistry/Nucleic Acid/RNA/Transfer RNA (tRNA)
Overview
[edit | edit source]Transfer RNA (tRNA) have a primary, secondary, and tertiary (L-shaped) structure. tRNA bonds to activated amino acids and transfers them to the ribosomes. Once at the ribosome, an initiator tRNA binds the amino acid to the ribosome to start translation. It carries the amino acids and binds to the Messenger RNA (mRNA) to form proteins.
tRNA's structure contains an amino acid attachment-site and a template-recognition site. The template-recognition site is called a anticodon and contains a sequence of three bases that are complementary to the codon on the mRNA. tRNA travels from nucleus to cytoplasm in a cell. Each tRNA can be used repeatably to be transcribed from DNA in nucleus.
There are 61 different anticodon sequences which code for the 20 amino acids. However, most prokayotic cells only have 30-40 different tRNAs and eukaryotes have about 50 different tRNAs. This is the third nucleotide of the codon, also called a wobble base, allows wobble pairing of the anticodon to the codon.

Role in Protein Synthesis
[edit | edit source]In protein synthesis, a tRNA molecule takes a specific activated amino acid to the site. The amino acid is esterified to the 3' or 2' -hydroxyl group of the terminal adenylate of tRNA. This joining of tRNA and an amino acid forms an aminoacyl-tRNA and is catalyzed by a specific enzyme called aminoacyl-tRNA synthetase (aaRS). There are 20 aminoacyl-tRNA synthetase, one for each amino acid. Similarly, there is a specific aaRS for each tRNA. The esterification reaction also called charging of the tRNA is powered by ATP.
The process of protein synthesis starts out when a charged tRNA (a tRNA with an amino acid attached), mRNA, and the small and large ribosomal subunits come together and form the initiation c complex, which consists of a peptidyl binding site (P site) and an aminoacyl binding site (A site). The first tRNA, otherwise known as the initiator RNA, binds to the mRNA start codon, AUG; thus, the first amino acid in the chain is methionine. To add additional amino acids to the polypeptide chain, a second charged tRNA must come in and have its anticodon bind to the next mRNA codon in the vacant A site. The P site and A site are in close proximity, thus allowing a formation of a stable peptide bond by reacting the carboxy terminus of the amino acid in the P site with the amino terminus of the amino acid on the tRNA in the A site. The reaction is catalyzed by peptidyl transferase. The complex moves along the RNA in a process called translocation which causes the tRNA in the P site to be displaced. The tRNA in the A site then moves into the P site so another charged tRNA can move into the A site. This process continues until the stop codon is reached the polypeptide chain is released from the ribosome.
1. amino acid + ATP --> aminoacyl-AMP + PPi 2. aminoacyl-AMP + tRNA --> aminoacyl-tRNA + AMP
tRNA Structure
[edit | edit source]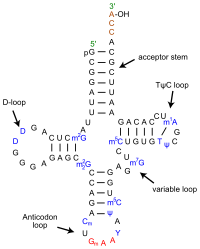
2. Secondary Structure
[edit | edit source]The secondary structure is formed like cloverleaf structure because of four base-paired stems also called arms. The cloverleaf contains three non-base-paired loops: D, anticodon, and TpsiC loop. The terminal CCA is not base paired. It's duplexed between the 5'segment and 3'segment.
The acceptor stem which is not a loop is the site where the enzyme amino-acyl-tRNA synthase attaches an amino acid. It is located opposite of the anticodon arm which reads the mRNA.
There are different types loops. In D loop, D arm ends. Anticodon arms ends in anticodon loop. In the figure, it shows hydrogen bond present inside the loop structure. The hydrogen bonds stabilized the structure.
3. Tertiary Structure
[edit | edit source]For the tertiary structure, it can be described as a compact of L shape. It is three dimensional. The structure is bonded and stabilized by base pairing and base stacking. Base pairs between nucleotides in the D loop and the TΨC loop. At the end of the L shape is the three base sequence called anti codon.
Anticodon
[edit | edit source]The anticodon region of a transfer RNA is a sequence of three bases. They are complementary to a codon in the messenger RNA. In the translation, the pairing between its anticodon and the messenger codon brings the ribosome. The amino acid is attached at its 3' end. And it will be peptide bond. In prokaryote cells, there are about 35 tRNAs with different anticodons present. In eukaryote cells, there are 50 tRNAs with anticodons present. tRNA with the anticodon CCC is complementary to the anticodon GGG. The anticodon AAA is complementary to the anticodon UUU. Since each type of tRNA has a different one, the anticodon of tRNA is able to identify others well.
tRNA Aminoacylation
[edit | edit source]Aminoacyl-tRNA is an amino acid ester of tRNA. It can be called a charged tRNA. When a polypeptide chain is formed by the anticodon of the tRNA, the reaction is thermodynamically unfavorable. So, aminoacyl-tRNA is used to activate the formation. An amino acid is esterified to the 3'-end of a tRNA containing the corresponding anticodon in amynoaclyation of tRNA molecules. As a result, the aminoacyl-tRNA attaches amino acids to the tRNA. These paring of amino acids and tRNAs define the genetic code. The aminoacyl-tRNA synthestase(AARSs)catalyze the aminoacylation of tRNAs. During transfer the genetic information from the nucleotide sequence of a gene to the amino acid sequence of a protein, this process plays an important role. When errors occur, amynoacyl-tRNA synthetases edit mechanisms structurally. Further, it prevents the error synthesis and releases aminoacylated tRNA that shouldn't be placed.
Aminoacyl tRNA synthetases (AARSs)
[edit | edit source]AARSs is an enzyme that catalyzes the esterification of specific amino acid to a tRNA to form an aminoacyl tRNA. AARSs take a major role in translation during protein synthesis. In recent researches, scientists discovered that AARSs also take role in ex-translation.
Role of AARSs in translation
[edit | edit source]The accuracy of the protein translation depends on the exactness of AARSs' recognition of both the amino acid to be activated and the cognate tRNA molecules. That is a crucial step in the fidelity of the translation. All AARSs carry out the same two-steps reaction:
- Step 1: AARSs binds ATP to the amino acid to induce an aminocyl-adenylate intermediate in which a covalent linkage between the 5'-phosphate in ATP and the carboxyl-end of amino acid.[2][3] Next, the AARSs use the generated energy from ATP hydrolysis to activate the amino acid which results in the formation of aminoacyl-AMP as an energy storage.[4]
- Step 2: The amino acid is transferred to the appropriate tRNA and bind either 2'OH or 3'OH of the 3' adenosine terminal of tRNA covalently. The energy that stored in aminoacyl-AMP is used to transfer the amino acid to the tRNA to form aminoacyl tRNA.
Role of AARSs in ex-translation
[edit | edit source]Modified version of AARSs and natural fragments take role in ex-translational functions as confirmed in recent studies. The interplay of AARSs appears to be at the center of homeostatic mechanisma which controls angiogenesis, inflammation, metabolism, and tumorigenesis.[5] Through some recent experiments, ex-translational functions of AARSs was found to be the interplay between natural extracellular fragments of human TrpRS and TyrRS in angiogenesis. TyrRS was found to have a nuclear localization signal that is controlled by its cognate tRNA (called tRNA-Tyr) so that a decrease in level of tRNA-Tyr will increase the level of nuclear import of the AARSs which will induce effects on many gene regulatory mechanisms. In contrast, an increase in level of tRNA-Tyr will decrease the level of TyrRS. Thus, the subcellular distribution of TyrRS is directly controlled by the demands of protein synthesis and this control is an example of homeostatic mechanism that balances a translational with an ex-translational functions.[6].
However, recent reseaches also showed that the ex-translational function of AARSs is regulated which is a contrast to the discussion above. This regulation is considered an auto-balancing process in which a natural fragment takes control in the activity of its own original protein. Thus, further research is needed to confirm the specific role of AARSs in ex-translational functions.
Binding to Ribosome
[edit | edit source]
tRNA will bind at the A, P and E sites of ribosomes. The A site will bind to aminoacyl-tRNA which was signaled by the codon that is binding to that site. The codon will also signify the next correct amino acid that will be in the peptide chain. But the A site will only work when the P site has an aminoacyl-tRNA attaching to it. The P-site is actually occupied by a chain with a few amino acids called peptidyl-tRNA. It carries synthesized amino acid chains. Lastly, the E site carries the empty tRNA.
Three dimensional image of a tRNA.
Coloring:
ORANGE: CCA tail
PURPLE:Acceptor stem
RED: D arm
BLUE Anticodon arm
BLACK: Anticodon
GREEN: T arm
Diseases Caused by Mitochondrial tRNA Gene Mutation
[edit | edit source]Mitochondria are an organelle in the cell, which contains 22 tRNA. Gene mutation of tRNA will cause serious diseases. There are seven kinds of genes diseases caused by mitochondrial tRNA gene mutation:

1- np5601 G->A and np3243 A->G gene mutations related to MELAS (Mitochondrial encephalomyopathy, lactic acidosis, and stroke-like episodes). Most patients get this disease before 40-year-old with epilepsia and lactic acidosis. Some of them will die during 20~30 age.
2- np8363 G->A, np8356 T->C, and np8344 A->G gene mutations related to MERRF (Myoclonic Epilepsy with Ragged Red Fibers). MERRF affects central nerve system, causing epilepsia, Dementia and epicophosis.

3- np4274 T->C gene mutation related to LIMM (Lethal Infantile Mitochondrial Myopathy). Most patients are newborn, having nerve defect and lactic acidosis, and die in one month.
4- np1644 G->T gene mutation related to subacute necrotizing encephalomyelopathy (SNE). This disease is familial autosomal recessive inheritance, happened to newborn baby.
5- np606 A->G gene mutation related to Rhabdomyolysis. Toxin produced by muscle cells is the main reason that causes Rhabdomyolysis.
6- np4500 G->A gene mutation related to the splenic lymphoma. The splenic lymphoma is a common malignant tumor happened on spleen. Normally, the splenic lymphoma caused by advanced stage lymphoma transfer.
7- np4336 A->G, np15927, and np15928 gene mutations related to Parkinson's Disease and Alzheimer’s Disease. Parkinson's Disease is a degenerative disorder of the central nervous system.
Reference:
1. Inheritance of Mitochondrial Disease: http://wenku.baidu.com/view/fe5a99ea172ded630b1cb69b.html
2. Diseases of Human Mitochondria tRNA: http://wenku.baidu.com/view/95fabd1fa300a6c30c229f93.html
3.http://baike.baidu.com/view/1120527.html
4.http://baike.baidu.com/view/2219175.html
5.http://en.wikibooks.org/wiki/Structural_Biochemistry/Nucleic_Acid/Translation
6. http://en.wikibooks.org/wiki/Structural_Biochemistry/Nucleic_Acid/RNA/Messenger_RNA_(mRNA)
7. http://www.wiley.com/college/boyer/0470003790/structure/tRNA/trna_intro.htm
| This page or section is an undeveloped draft or outline. You can help to develop the work, or you can ask for assistance in the project room. |
- ↑ http://www.wiley.com/college/boyer/0470003790/structure/tRNA/trna_intro.htm
- ↑ Desogus, Gianluigi; Flavia Todone; Peter Brick; and Silvia Onesti. "Active Site of Lysyl-tRNA Synthetase: Structural Sudies of the Adenylation Reaction. Biochemistry, 2000 vol 39, 8418-8425.
- ↑ Klug, William, and Michael Cummings. Concepts of Genetics.5th Edition. Upper Saddle River, NJ: Prentice Hall, 1997.
- ↑ Hartweel, Leland; Leroy Hodd; Michael Goldberg; Ann Reynolds; Lee Silver; and Ruth Veres. Genetics: From Genes to Genomes. Boston: Mgraw-Hill, 1999.
- ↑ Guo, Min, and Paul Schimmel. "ScienceDirect.com - Trends in Biochemical Sciences - Homeostatic mechanisms by alternative forms of tRNA synthetases." ScienceDirect.com | Search through over 11 million science, health, medical journal full text articles and books.. N.p., n.d. Web. 7 Dec. 2012. <http://www.sciencedirect.com/science/article/pii/S0968000412001077>.
- ↑ G. Fu et al. tRNA-controlled nuclear import of a human tRNA synthetase J. Biol. Chem., 287 (2012), pp. 9330–9334