Structural Biochemistry/DNA recombinant techniques/Mutagenesis
Oligonucleotide mutagenesis is a process in which a specific DNA sequence is incorporated into a vector (such as a plasmid or lambda phage) to introduce a new function into that molecule. Mutagenesis allows the insight into how proteins fold, catalyze reactions, act as substrates and process information. Cloned gene proteins can be used medically in a number of ways. For instance, insulin and interferon can be obtained from production of bacteria. DNA probes may be manufactured for discovery of genetic diseases, cancer, and bacterial infections. Two main forms are used to produce these modified proteins, Site directed mutagenesis and oligonucleotide-directed mutagenesis. The later is used if restriction sites are absent.
Mutagenesis is the process of changing or creating genetic information. This process can occur naturally or made by using different methods. There are more approaches involving mutagenesis methods. One method is a method of site-directed mutagenesis using mismatched oligonucleotides. Second method is cassette mutagenesis by annealing complementary oligonucleotides. Third method is PCR by generating a mutant fragment starting from a double-stranded DNA template using mismatched oligonucleotides. Mutagenesis is also used by experimenters to analyze DNA. Site mediated mutagenesis mutates one sequence of amino acids to determine that specific sequences overall purpose once it stops working. This is an example of experimenters using Mutagenesis.
Site-Directed Mutagenesis
[edit | edit source]In site-directed mutagenesis, individual amino acids in the primary structure of a protein are replaced by changing the DNA sequence of a cloned gene. The process involves removal of a DNA segment and replacement of that segment with a chemically synthesized segment identical to the original but with the desired change. The steps are as follows: A recombinant plasmid containing the gene of interest is treated with a restriction endonuclease to cleave the sequence of interest. The synthetic DNA fragment with the specific base pair change is inserted into the plasmid. This is done using a DNA ligase. The plasmid then contains the gene with the desired nucleotide base pair change.
Site directed mutagenesis is also useful for testing the validity of an enzymatic mechanism.
Oligonucleotide-Directed Mutagenesis
[edit | edit source]This form of mutagenesis creates a specific DNA sequence change. This method makes point mutations, differing in a single nucleotide. In this process, a DNA strand is synthesized with the specific nucleotide base pair change and annealed to the copy of the gene in a plasmid. Increasing the temperature allows for the mismatched base pair to be substituted in order for this reaction to occur. This annealed strand acts as a primer for the complementary strand to be synthesized in the plasmid. Oligonucleotide-directed mutagenesis uses DNA polymerase, dNTPs and DNA ligase. The result are two types of progeny: one with the original sequence and another with the new sequence.
Cassette Mutagenesis
[edit | edit source]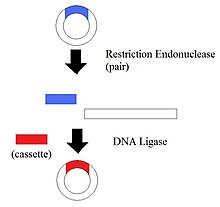
Cassette Mutagenesis is the replacement a region of DNA with new sequences. Any length and sequence can be replaced by this method. Therefore, more variations of proteins can be made. By taking advantage of this, new mutant sequences can be made. The Cassette Mutagenesis uses the original gene containing restriction sites. This site allows cleavage of the gene and produce the region where the new DNA is inserted. After appropriate restriction sites are identified in a vector, the vector is cleaved at the two sites by using a endonucleotide. The new sequence is inserted and ligated in the region produced. The new sequence allows a variety of studies on protein structures or nucleic acid sequences that have not been explored before.
Polymerase Chain Reaction
[edit | edit source]Polymerase chain reaction is used for recombination of sequences and mutagenesis. This method is very useful because it is a fast reaction and it is not limited by any restriction sites. In the PCR, oligonucleotide primers are incorporated into the ends of the product DNA. The 5' ends of these primers can contain any desired sequence. two PCR products are added by PCR. They are mixed, denatured, and reannealed. There are two different sequences produced. One is 5' overlapping strands which is not productive. 3' overllapping strand is produced and reactive. The polymerase extends the 3'strand. In this process, the 3' strand acts as primers. The whole process are repeated by producing synthetic sequences. This overlapping by joining two DNA fragments together can produce mutatation in PCR fragment.
Reference
[edit | edit source]M.J. McPHERSON, Directed Mutagenesis