Structural Biochemistry/DNA Mutation
In general, there are two ways that mutations in DNA sequences could occur:
Environmental effects
[edit | edit source]Altering Nucleotide bases: File:Environmental agents damage DNA.jpgEnvironmental effects such as Ultraviolet light, radiation, or toxic chemicals could modify nucleotide bases to make them look like other nucleotide bases, resulting in damages of DNA. For example, certain environmental agent will change the structure of Guanine base so that it has the shape like Adenine. Thus, during the DNA replication, that “Guanine base” can no longer bind to Cytosine, instead, because it has a shape of Adenine, it will bind to Thymine.
Breaking the Phosphate backbone: Environmental agents can also break phosphodiester bonds between oxygen and phosphate groups. By breaking the phosphate backbone of DNA within a gene, a mutated form of that gene could form. And this mutated gene might results in a mutated protein that functions differently, and might cause protein misfolding diseases.
However, cells usually attempt to fix the broken dens of DNA by joining free ends to other DNA fragments in the cell. This creates “translocation,” another kind of mutation. If this translocation breakpoint happens inside or near a gene, that gene’s functions may be influenced.
DNA replication mistakes
[edit | edit source]During DNA replication, DNA helicase first separates the DNA double-strand into two single strands. Then, DNA polymerase helps add corresponding nucleotides to both template strands, creating two double-stranded DNA molecules. However, DNA polymerase could make mistakes during this process at the rate of once every 100,000,000 bases. The result is mutations of genes, which could lead to many malfunction of translated protein.
In fact, most of the mistakes can be repaired by a type of protein later in the replication process. This protein will replace incorrectly paired nucleotides with correct ones. So that the number of mutations of DNA is actually lower.
Frameshift mutations
[edit | edit source]Frameshift mutation is caused by insertion or deletion of a number of nucleotides that is not evenly divisible by three from a DNA sequence. It often occurs when the addition or loss of DNA bases disrupts a gene’s reading frame. A reading frame consists of groups of three DNA bases (codons), which each code represents one specific amino acid. The resulting protein is usually nonfunctional because the wrong reading frame of the gene translate a very different protein sequence from the normal reading frame.
For example, mRNA with sequence AUG CAG AUA AAC GCU UAA Normal amino acid sequence reading frame should be: MET GLN ILE ASN ALA STOP However, a wrong reading frame (a deletion of the first base 'A') could give a traslation of mRNA of: UGC AGA UAA ACG CUU AA an abnormal amino acid sequence translate would be: CYS ARG STOP
In the case above, a frameshift mutation causes the reading of all codons after the mutation to code for different amino acids. The stop codon ("UAA")cannot be read, which a stop codon could be created at an earlier or later site. The protein being created above is abnormally short, which contain the wrong amino acid; thus, it is nonfunctional.
Frameshift mutations can result in severe genetic diseases such as Tay-Sachs disease, which is caused by the missing enzyme due to genetic mutation that result in the accumulation of fatty substance (Gangliosides) in the nervous system. However, frameshift mutation can be beneficial. For example, a frameshift mutation was responsible for the creation of nylonase, which is capable to digest certain byproducts of nylon 6 manufacture.
Chromosomal Translocation
[edit | edit source]Another type of DNA mutation that can occur is chromosomal translocation, which is a chromosome abnormality that is caused by the rearrangement of parts of nonhomologous chromosomes. There are instances where the two separated genes are joined together, forming a fusion gene, which is common in cancer. This fusion gene can be detected on a karyotype of affected cells. There are two main types of chromosomal translocation that can occur: reciprocal (non-Robertsonian) and Robertsonian. Translocations can also be balanced, where there is an even exchange of genetic material with no information extra or missing, or unbalanced, where the exchange of chromosomal material is uneven resulting in extra or missing genes. Some diseases that result from translocation include cancer, infertility, and Down Syndrome.
Reciprocal Translocation
[edit | edit source]Reciprocal translocations are usually an exchange of material between nonhomologous chromosomes. These kinds of translocations are, for the most part, harmless because the amount of genetic material exchanged is the same. They can usually be detected through prenatal diagnosis. However, carriers of balanced reciprocal translocations have an increased risk of creating gametes that have unbalanced chromosome translocations that end up leading to miscarriages or even children with abnormalities.
Robertsonian Translocation
[edit | edit source]The Robertsonian translocation is most commonly found in children with Down Syndrome. The parents of children with Down syndrome are carriers of unbalanced gametes which lead to miscarriages and/or abnormal offspring. The case of translocation in children with Down syndrome is called trisomy.

Chromosomal Inversion
[edit | edit source]An inversion is a rearrangement of the chromosomes where a segment of the chromosome is reversed end to end. An inversion occurs when a single chromosome undergoes breakage and rearrangement within itself. There are two types of inversions: paracentric and pericentric. Paracentric inversions do not include the centromere and so both breaks occur in one arm of the chromosome. Pericentric inversions do include the centromere and so there is a break point in each arm.
Inversions usually do not cause any abnormalities in carriers so long as the arrangement is balanced. This means there are no extra or missing genetic information. However, those who are heterozygous for an inversion have an increased production of abnormal chromatids, which leads to lowered fertility due to the production of unbalanced gametes.
Point Mutations
[edit | edit source]A point mutation is a mutation where a single base nucleotide is replaced with another nucleotide of the genetic material, DNA or RNA. The term point mutation often includes insertion and/or deletions of a single base pair. Point mutations can be categorized as one of two types:
Transitions: the replacement of a purine base with another purine or the replacement of a pyrimidine with another pyrimidine
Transversions: the replacement of a purine with a pyrimidine or vice versa
Point mutations can also be categorized functionally:
Nonsense mutations: code for a stop, which can truncate the protein
Missense mutations: code for a different amino acid
Silent mutations: code for the same or a different amino acid but there is no functional change in the protein
An example of a missense mutation is sickle-cell disease. A missense mutation occurs in the beta-hemoglobin gene that converts a GAG codon into a GTG codon, which encodes for the amino acid valine rather than glutamic acid.

Insertion
[edit | edit source]Insertion is the addition of one or more nucleotide base pairs into a DNA sequence. This can happen often when DNA polymerase is slipping in microsatellite regions. Insertions can vary in size, some being only a single nucleotide base pair whereas others can be a section of another chromosome being inserted into the DNA sequence. On the chromosomal level, an insertion refers to the insertion of a larger sequence into a chromosome. This usually occurs due to unequal crossover during meiosis. There are a couple different kinds of insertions that can occur based on how and what is inserted. An N region addition is the addition of non-coded nucleotides during recombination by terminal deoxynucleotidyl transferase. A P nucleotide insertion is the insertion of a palindromic sequence encoded by the ends of the recombining gene segments.

Deletion
[edit | edit source]Deletion is a mutation in which a part of the chromosome or a sequence of DNA is missing. Deletion is the loss of genetic material. Any number of nucleotides can be deleted, ranging from a single base pair to an entire piece of the chromosome. Deletions are usually caused by errors in chromosomal crossover during meiosis. Some of the causes of deletions include losses from translocation, chromosomal crossovers within a chromosomal inversion, unequal crossing over, and breaking without rejoining. Some types of deletions are terminal deletion and interstitial deletion. Terminal deletion is a deletion that occurs near the end of a chromosome. Interstitial deletion is a deletion that occurs from the interior of the chromosome.
Small deletions are less likely to be fatal while large deletions can be more fatal because there are always variations based on what genes are lost. Some of the medium-sized deletions can lead to recognizable human disorders. Deletions are responsible for a variety of genetic disorders, such as male infertility and two thirds of cases of Duchenne muscular dystrophy.

Amplification
[edit | edit source]Amplification is the duplication of a region of DNA that contains a gene and can occur as an error in homologous recombination, a retrotransposition event, duplication of an entire chromosome. This duplication arises from unequal crossing-over that takes place during meiosis between misaligned homologous chromosomes. Amplification does not usually constitute a lasting change in a species' genome, not lasting longer than the initial host organism. Amplification is actually a way for a gene to be overexpressed. It can occur artificially via polymerase chain reaction or it can occur naturally, as was just explained.
Gene amplification is believed to play a major role in evolution and this belief has lasted for over 100 years in the scientific community. The duplication of a gene results in an additional copy that is free from selective pressure. The new copy of the gene is then allowed to mutate without deleterious consequences to the organism. With this freedom from these consequences, the mutation of novel genes can occur which could potentially increase the fitness of the organism or code for a brand new function. The two genes that are present after the gene duplication are paralogs and they usually code for proteins that have similar function and/or structure.
Deamination
[edit | edit source]
Deamination is removing the amino group from the amino acid and converting to ammonia. Since the bases cytosine, adenine and guanine have amino groups on them that can be deaminated, Deamination can cause mutation in DNA. For example, If a cytosine were to be deaminated to form uracil (uracil is an analog of thymine) in the template strand of DNA, then the polymerase would put in an adenine at the corresponding position on the nascent DNA strand instead of a guanine. The hydrolysis reaction (deamination) of cytosine into uracil is spontaneous.
In response to this mutation the cell has a repair process. In this process the cell utilizes the enzyme uracil-DNA glycosidase to recognize these uracils and removes them. This enzyme hydrolyzes the N-glycosidic bond between the deoxyribose ring and the uracil base. Therefore, the uracil base is removed.

Since this site on the DNA duplex is without either purine base or a pyrimidine base is called an AP site (either apurinic or apyrimidinic). Then the enzyme AP endonuclease cut the bond on the 3' side of the phosphodiester bond of the nucleotide. In this phase DNA polymerase I recognizes phosphodiester bond at the 3'end on the next nucleotide unit and cleaves the bond. After the ribose-phosphate unit is removed, DNA polymerase I analyzes the complementary strand and finds that the base that corresponds to the AP site is guanine. Then the enzyme inserts a cytosine unit at the AP site on the broken DNA strand. Finally, DNA ligase seals the inserted cytosine into the damaged strand. Spontaneously deamination of cytosine to form uracil can be repaired by the cell. [1]
Quickchange Method
[edit | edit source]Quickchange is a technique used to generate site-specific mutations with minimal hands-on manipulation. The sites of mutations are incorporated in the two complementary primers and the rest of the plasmid DNA is synthesized with a high fidelity DNA polymerase in a thermal cycler. Therefore the whole process is considered quick.
Although the reaction is done in a thermal cycler, it is not PCR. Since the template is circular, the newly synthesized single stranded DNA will terminate at the beginning of the primer on the same strand. This product will not overlap with the primer on the complementary strand. Therefore the newly made DNA cannot be used as a template for further DNA synthesis. Only the original template DNA can be used as templates. In each cycle, the amount of newly synthesized DNA is equal to the template. This is considered linear amplified, rather than exponential amplification in PCR.
Since the template DNA is isolated from bacteria, it contains methylated nucleotides. That makes it sensitive to methylation dependent nucleases, such as DpnI. For example, after 20 cycles of amplification, 10 ng plasmid will be amplified 20 fold and produce 200 ng new DNA. At this moment, restriction endonuclease DpnI will be used to eliminate the original plasmid DNA. The mixture of DNA is then put into bacteria and each DNA species will be separated in different bacteria cells. To see whether a cell contains the right mutation, a single cell needs to be picked, grown up, and the DNA it contains analyzed. Designing the primers is critical. A minimal annealing temperature of 78oC has to be met. Otherwise, the primer will not be attached to its template and the termination will not be stopped precisely.
Quickchange Protocol
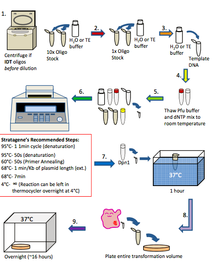
[edit | edit source]In general, the way in which quickchange if performed is in the steps below. This technique is similar to PCR. 1. If oligos are from IDT: Spin oligos down at high speed for 1 minute using a table top centrifuge and resuspend in H2O or TE buffer to a 10x stock (1250 ng/µl) If oligos are from Allele: Dilute oligos in H2O or TE buffer to a 10x stock (1250 ng/µl) 2. Further dilute oligos to 1x (125 ng/µl) in H2O or TE buffer 3. Dilute Template DNA to 20-50 ng/µl in H2O or TE buffer 4. Thaw 10x pfu ultra buffer and dNTP mix to room temperature 5. Set up quick change reaction in a 100µl thin walled PCR Tube: H2O: 40µl 10x PFU Ultra Buffer: 5µl dNTP mix (10mM): 1µl Template DNA (20-50 ng/µl): 1µl Forward Primer (125 ng/µl): 1µl Reverse Primer (125 ng/µl): 1µl PFU Ultra HF (2.5u/µl): 1µl Total: 50µl 6. Mix contents of PCR tube gently and place in the thermocycler. Program the thermocycler according to what is needed for the experiment 7. When the reaction has finished, remove the PCR tube from the thermocycler and add 1µl of Dpn1 directly to the contents of the PCR tube and incubate for 1 hour at 37°C 8. Perform a transformation (see transformation protocol) using 2-3µl of the Dpn1 treated PCR reaction into XL1 Blue competent cells and plate the entire volume onto LB agar plates that contain the antibiotic that corresponds to the template DNA. Incubate overnight at 37°C 9. Check plates for colonies the following day. If there are colonies, use them to inoculate 5 ml overnight cultures and perform mini preps the following day. Send 5µl of the mini prep DNA for sequencing and analyze the results
Reference
[edit | edit source]1. Campbell, Neil A. (2005). Biology. Pearson. ISBN 0-8053-7146-0. {{cite book}}: Check |isbn= value: checksum (help); Text "coauthors+ H.C. Van Ness, M.M. Abbott" ignored (help)
2. http://www.answers.com/topic/frameshift-mutation http://www.gmilburn.ca/2009/04/03/human-evolution-and-frameshift-mutations/ http://www-personal.ksu.edu/~bethmont/mutdes.html#types