Structural Biochemistry/Cell Organelles/Ribosome
Overview
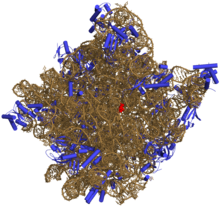
[edit | edit source]The purpose of the ribosome is to translate messenger RNA (mRNA) to proteins with the aid of tRNA. In eukaryotes, ribosomes can commonly be found in the cytosol of a cell, the endoplasmic reticulum or mRNA, as well as the matrix of the mitochondria. Proteins synthesized in each of these locations serve a different role in the cell. In prokaryotes ribosomes can be found in the cytosol as well. This protein-synthesizing organelle is the only organelle found in both prokaryotes and eukaryotes, asserting that the ribosome was a trait that evolved early on, most likely present in the common ancestor of eukaryotes and prokaryotes. Ribosomes are not membrane bound.
Ribosomes are also play a key role in the catalysis of two important and crucial biological processes. They are responsible for peptidyl transfer, in which they form peptide bonds during protein synthesis as well as peptidyl hydrolysis; in this process, the ribosome releases the completely formed protein from the peptidyl transfer RNA after completion of translation. Translation is the process of converting mRNA (from transcription) to functional proteins; The three steps involved are initiation, elongation, and termination. Proteins are translated one amino acid (three base pairs) at a time. During this process, tRNA assists the ribosome by bringing the complementary bases to the ribosome as translation proceeds.
Because ribosome synthesis is a major metabolic activity that involves hundreds of individual reactions in eukaryotes, errors will occur eventually during these processes. To deal with the error in ribosome synthesis if it occurs, eukaryotes will degrade the erroreous ribosomes. In addition, not only the erroneous ribosomes will be degraded but also the excess ribosomes. Recent researches show that eukaryote cells enhanced many strategies to recognize specific dysfunctional or functionally deficit ribosomes for degradation[1].
Translation: Eukaryotic and Bacterial
[edit | edit source]Along with having more complicated assembly regulations than bacterial ribosomes, eukaryotic ribosomes are triggered differently than bacterial ribosomes in translation initiation. Eukaryotic ribosomes use a scanning mechanism and require at least nine initiation factors. [2] In bacteria, ribosomes can identify the right reading frame and know where to attach to a mRNA stran, by finding the Shine Dalgarno sequence on the mRNA. This ribosome binding site is upstream from the AUG start codon on the mRNA. In the bacterium E. Coli, the Shine Dalgarno site is a purine-rich nucleotide sequence—5’ AGGAGGU 3’, on the mRNA that is located four to eight bases ahead of the start codon. This Shine-Dalgarno sequence is complementary to the sequence 5’ ACCUCCU 3’ on the 16S rRNA, which is found in the 30S subunit of the ribosome. [3]
Structure
[edit | edit source]Ribosomes usually comprise 2 protein subunits and some component of rRNA. The protein componenet of the ribosomes is first synthesized in the cytosol much like any other protein. The newly synthesized protein subunits are then transported from the nucleus into the nucleolus, the center of ribosomal RNA transcription. RNA polymerase are then used to polymerize the rRNA needed for the synthesis of the ribosome. Cells that have a high rate of protein synthesis have many ribosomes.
In the past, scientists studied ribosome to get a better understanding about its components, which is consists of 30S, 50S, 70S, and 100S. All are made of by the same particles but different concentration of Mg ions. 30S and 50S are small and large subunits of 70S while 100S is a collection of 70S. Ribosomal structure varies between eukaryotes and prokaryotes. Eukaryotes have 40S (small) and 60S (large) subunits, together the subunits form the 80S ribosome.[4] Prokaryotes have 30S (small) and 50S (large) subunits. These subunits work together during translation to synthesize proteins.
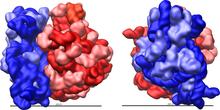
In prokaryotes, the small subunit (30S) and the large subunit (50S) together make the 70S ribosome. The 30S subunit is composed of 21 ribosomal proteins and 16S rRNA. Its function is to decode and decipher mRNA to determine the corresponding amino acid to the three bases in codons. The small subunit consists of the body (5’), the platform region (central domain), and the head (3’). Each component can be formed separately of each other. The small subunit also contains a 3’-minor domain consisting of the last two helices (44 and 45) and the end of the 3’ end of the rRNA. The structural components of the small subunit have been shown to be structurally independent from each other suggesting that during protein synthesis they move and conform relative to one another. The long 44 helix stretches across the small subunit from the body to the head functioning as a potential relay for information across the entire subunit. Unlike the small subunit, the large subunit is mostly rigid with mobility restricted its peripheral regions. The 50S subunit consists of 33 proteins and the 23S and 5S rRNA.
The mRNA binding site is located along the neck of the small subunit while the large subunit contains the peptidyl transferase center (PTC) where aminoacyl and peptidyl-tRNA attach. The three binding sites are A, P, and E. Binding site A attracts aminoacyl-tRNA, binding site P attracts peptidyl-tRNA, and binding site E, the exit site, attracts deacylated tRNA. The anticodon stem loop is attached to its complementary codon on the mRNA binding site of the small subunit while the amino acid end of the tRNA attaches to A and P sites both located in the PTC. The GTPase-associated center and the sarcin/ricin loop are located on the large subunit and help to stimulate GTP hydrolysis needed for elongation of the protein.
Ribosome assembly consists of transcription, translation, the folding of rRNA and ribosomal proteins, the binding of ribosomal proteins, and the binding and release of the assembly components to make the ribosome. There are two intermediates in ribosome assembly, in vitro and in vivo. One method of in vitro assembly of the 30S is to only use free rRNA and ribosomal proteins. Another is to combine 16S rRNA with purified and recombinant proteins and precursor 16S rRNA. With the different ways to assemble proteins, the assembly of ribosomes also varies with different temperatures. At low temperature of 0°C -15°C, the reconstitution intermediate (RI) particle, composed of 16S rRNA and 15 ribosomal proteins, is formed and settled to 21S-22S. Then the RI particle is rearranged after getting heated to 40°C and settled to 25S-26S. After completing these two processes, the RI particle together with the remaining proteins can form the 30S subunit.
In contrast, the in vitro assembly of the 50S subunit requires more steps and harsher conditions compared to the assembly mechanism of the 30S. There are three reconstitution intermediates each dependent on different temperatures and ionic conditions. RI50 (1), the first intermediate, yields the 33S particle. As RI50 (1) is heated, RI50 (2) forms and sediments at the 41S-43S. The remaining proteins form the RI50 (3) which create the inactive 48S particle. In order for the 48S particle to become the active 50S subunit, the temperature and concentration of magnesium needs to be increased along with the involvement of the 5S rRNA.

Like the in vitro assembly mechanism, the in vivo assembly of the 30S subunit has two intermediates (p130S and p230S) and the 50S subunit has three intermediates (p150S, p250S, and p350S). However, the reconstitution intermediates are not the same as in vitro. The intermediates of the 30S subunit yield 21S and 30S particles while the intermediates of the 50S subunit yield 32S, 43S, and 50S particles. The intermediates in the in vivo assembly are precursor rRNA which is different from in vitro which uses matured rRNA. To complete the mechanism of ribosome assembly, these precursor rRNA gets transformed in the polysomes.
Eukaryotic Ribsome
[edit | edit source]A eukaryotic ribosome is composed of a 40S and 60S subunits, they both have a solvent exposed side and a subunit interface. The solvent exposed side contains a higher concentration of protein and RNA elements that are only in eukaryotes compared to the subunit interface.
The eukaryotic specific proteins are responsible for 40S structure being stabilized by tertiary contacts. The majority of the eukaryotic specific proteins and extensions are to interconnect other proteins in the 40S and 60S subunit. The 40S subunit has 14 of these interconnected proteins located in the head. eukaryotic specific proteins and extensions play a bigger role in 60S subunits because their protein mediated connections are more extensive and reach across the subunit. The 60S specific extensions consist of beta sheets and long alpha helices which are crucial in long distance tertierary interaction, meaning they have the ability to interact across the subunit. This trait is unique for eukaryotes because protein-protein contact is very rare in the prokaryote ribosome.
The 40S and 60S subunits are joined by eukaryote specific inter-subunit bridges that connects them thus forming 80s. This is done by eukaryotic specific bridges forming at the 40S and the RPL19 interacts with ES6E and RPL24 interacts with rpS6. The other two eukaryotic specific bridges connect by 60S segment ES31 that connects to rpSA and another 60S segment, ES41 that connects with rp58 from the 40S subunit. [5]
Ribosomal Cofactors
[edit | edit source]Ribosomal cofactors help facilitate with improper folding during in vitro. The improper folding during in vitro is a cause which results in vitro being much slower than in vivo. Ribosomal factors can be used as a “check point” to make sure the assembly is correct. Examples of ribosomal cofactors are DEAD-box proteins, chaperones, and GTPases. DEAD-box proteins are factors which help with proper folding of the RNA. An over expression of DEAD-box proteins can suppress the deletion of another strain. DEAD-box proteins (DBPs) are characterized by a core of approximately 350 amino acids, each containing at least 9 conserved amino acid motifs. Studies have shown that these proteins use the energy from ATP hydrolysis to rearrange inter- or intra-molecular RNA structures or dissociate RNA-proteins interactions. In eukaryotes, DEAD-box proteins are fundamental in the life of a ribosome, but in E. coli, they are often unnecessary. To date, five different genes encoding DEAD-box proteins have been identified in E.coli – SrmB, CsdA, DbpA, RhlE, and RhIB – which are versatile cofactors that have helped further the understanding of cellular processes in many cells because of their roles in protein biosynthesis and ribosome assembly. For instance, CsdA and SrmB are essential for proper cellular growth; cells deficient in these proteins exhibit severe ribosomal effects.

Brief History
[edit | edit source]Ribosomes played a big role in our evolution. The origin of ribosomes are said to have roots in the RNA world, preceding the existence of proteins. Ribosomes evolved in structure and function, alongside the formation of complex and versatile peptides. A huge breakthrough in the scientific study of ribosomes began when Venkatraman Ramakrishnan and his group determined the crystal structure of the ribosome, and described it in atomic detail. Uncovering the structure of the ribosome allowed scientists a closer look at the mechanisms involved with ribosome function. More specifically for example, according to "The ribosome goes Nobel" article by Rodnina and Wintermeyer, in the "Trends in Biochemical Sciences" journal, knowledge of the structure of the ribosome revealed details of the translation process like "how the ribosome checks for proper codon–anticodon interaction by establishing specific contacts in the decoding site". More recently, ribosomes prove to have huge clinical potential as targets for antibiotic products. With the overuse of antibiotics and antibiotic resistance constantly on the rise, ribosomes could be the basis of new antimicrobials that would retaliate against the infamous antibiotic resistance.
Free Ribosomes
[edit | edit source]Ribosomes found in the cytosol that are not bound to any other organelle are known as free ribosomes. These ribosomes are known to produce non-hydrophobic proteins that ultimately function in the cytosol. An example would be the catalysts for the first steps of sugar breakdown.
Endoplasmic Reticulum Ribosomes
[edit | edit source]Ribosomes which are attached to the endoplasmic reticulum or ERare known as ER Ribosomes. These ribosomes are what give rough endoplasmic reticulum its "rough" appearance. Proteins that are produced from the ribosomes of the rough ER are sent through the lumen of the endoplasmic reticulum, modified within the ER, and then is exported in a vesicle to the cis face of the golgi apparatus, where the protein undergoes further modification.
The Peptidyl-Transfer Reaction Catalyzed by the Ribosome
[edit | edit source]
Enzymes transform their substrates through the active site residues. This transformation can allow for the direct participation in chemical catalysis, such as facilitation of proton transfer reactions and covalent chemistry. These active-site residues can form the covalent bond acting as general acid-base or as nucleophiles. The ribosomes undergo general acid-base or nucleophilic catalysis. The residues can also be structural complement to the transition state (TS), dissolve or reorganize water molecules, and decrease the delta S (entropy) by using the binding energy. In the elongation process, the ribosome PTC catalyzes the aminolysis of the ester bond: the alpha amino group of the A-site aminoacul tRNA acts as nucleophile and attacks the P-site peptidyl tRNA at the carbonyl carbon of the ester bond that connects the tRNA and the peptide. In order to efficiently form peptide bonds amines react with esters. This reaction is fast with the help of the ribosome because it makes the reaction rate increase by approximately 106 - 107 folds. Before the peptide bond is formed, the aminoacyl tRNA connects to the A site at a rate of 10 s-1 which is much slower than the rate of the tertiary complex forming a peptide bond. The slow rate of reaction allows for the minimal use of substrate analog which is short RNA fragments that is similar to the tRNA CCA 3’ end or the alpha amine substituted with a hydroxyl group. The minimal substrates consisted of puromycin (Pmn), C-puromycin (C-Pmn), and CC-puromycin (CC-Pmn). These substrates bind to the ribosome fast to react with the P-site substrate in order to allow for the direct analysis of their chemistry relationship. Recently, methods of analysis using kinetic, biochemical, genetic, and computation allow for more progress in ribosome crystallography. However, there are still many questions regarding the proton transfer mechanism. Peptidyl transfer by the ribosome accompanies a decrease in the entropy activation, which provides more understanding of ribosome catalysis involves defining the molecular mechanism that make entropy of activation lower by taking into consideration the substrate, position of the active site, desolvation, and electrostatic shielding.
1)Peptidyl Transfer 2)Peptide Release 3)Structures of transition state analogs used to study the ribosome
Degradation of erroneous and excess ribosomes in eukaryotes
[edit | edit source]When cells are in stress, disease condition, and normal condition, RNA damage occurs. Exposing to ultraviolet light, oxidation, chlorination, nitration and alkylation result in chemical modifications to nucleobases, generating the potential of triggering ribosome degradation pathways.
Types of error in ribosomes synthesis
[edit | edit source]There are several sources of errors in ribosome synthesis such as the alternation in rRNA sequences (occur in cis) and the failure to bind to or loss of an assembly factor or ribosomal protein (occur in trans). Beside these main sources, specific growth conditions can also cause certain effects in ribosome synthesis. For example, starvation requires excess ribosomes which are turned over efficiently[6].
Potential error-occurring process in Ribosome synthesis
[edit | edit source]mRNA decoding and peptidyl-transfer reaction are two subunites that have specialized function in ribosome translation[7]. Each eukaryote ribosome is composed of 4 rRNAs and ~80 ribosomal proteins. The processes of synthesizing, maturizing, transporting individual ribosomes and assembling into ribosomal subunits requires the contribution of ~200 proteins trans-acting cofactors and a large amount of small nucleolar RNAs (snoRNAs). These processes engage in hundreds of individual error-causing reactions[8][9].In addition, many ribosomal proteins also perform non-ribosomal functions, so the functions in these processes which are not directly connected to ribosome biogenesis are currently being assigned to ribosome synthesis factors (RSFs). RSFs include connections to cell cycle progression, pre-mRNA splicing, DNA damage response, nuclear organizing and telomere maintanance. Due to the large number of reactions that occur in ribosome assembling pathway, the possibility of error occurring is very high such as the potential harms for cell viability and human health. For example, inability to bind correctly to or loss of a synthesis factor can cause the production of erroneous ribosomes such as lacking part of ribosomal proteins or carrying misfolded rRNA which will affect on translation. As a result, cells develop control mechanism to avoid such problems. For example, instead of participating in later steps, synthetic factors that involved in late cytoplasmic ribosome assembly steps can first bind with pre-rRNAs at early stages, bringing pre-ribosomes to productive synthesis pathways[10].
Mutation in cis: non-funtional ribosomal RNA decay
[edit | edit source]There are many pathways that have been known to control the structural integrity of mature RNA molecules[11]. One of the pathways that control the decay of mRNAs is the "no-go" decay (NGD), which the mRNAs that cause the degradation of specific ribosomes[12]. LaRivie`re et al. introduced the way to test whether or not mutations in functional and non-functional ribosomal sites affect rRNA stability which is performing substitutions in the decoding site and peptidyl transferase centre at the position that are essential for ribosomal function in bacteria. This experiment gave result in the identification of non-functional rRNA decay, a pathway that detects and eliminates dysfunctional parts of mature ribosomes[13].
Mutations in trans: The TRAMP-exosome pathway and nucleolar surveillance
[edit | edit source]Aberrant nucleolar and nuclear pre-rRNAs are found to be degraded actively by a nuclear surveillance mechanism that involves the addition of unstructured oligoadenylate tails at the 3’-end of flawed pre-rRNAs. This is done with the TRAMP complexes’ polymerase activity, and after with their degradation by the RNA exosome. TRAMP conplexes consist of a poly(A) polymerase, a zonc-knuckle-containing and putative RNA-binding protein, and the DEVH helicase Mtr4. The pathway goes from having the addition of short poly(A) tails with the actual binding between TRAMP complex and RNA, and these two commit deviated molecules to degradation. This is done by separating them from the normal RNA and by stirring exosomal activity[14].
The decay of bulk ribosome and pre-ribosome by ribophagy and PMN
[edit | edit source]Growth-inhibiting conditions cause ribosome synthesis to shut down. The synthesis of complex molecules from simple molecules such as amino acids and sugars is obstructed and existing pre-ribosomes and mature ribosomes are directed towards bulk degradative pathways. To manage stress, and to adapt to a new environment, pre-ribosomes and mature ribosomes are passed on to be recycled for important cellular components. Ubiquitin, a small regulatory protein essential to 25S NRD (Nonfunctional RRNA Decay), is also responsible for bulk ribosome degradation.[15]
Autophagy is a catabolic mechanism that takes unnecessary or dysfunctional cellular components and performs cellular degradation through the lysosome, or the vacuole in yeast and plants.[16] Starvation conditions cause large portions of the cytosol, along with protein, to assemble and whole organelles, such as ribosomes and mitochondria, are recycled through two major types of autophagy: microautophagy and macroautophagy. Macroautophagy involves the development of a double membrane around the organelle, this together known as an autophagosome. [17] This isolates the organelle and delivers it to the lysosome, which engulfs the organelle through endocytosis for its breakdown and recycling. [18] In contrast, during microautophagy, the lysosome directly engulfs the cytoplasmic material, through the inward folding of the lysosomal membrane. [19] Because lysosomes and vacuoles contain non-specific enzymes for hydrolysis, autophagy can essentially degrade any biomolecule. Macroautophagy and microautophagy are both crucial to a cell's survival during starvation. Both mechanisms associate common trans-acting factors, and both are essentially bulk degradative processes. However, studies show that selective types of both channels exist. [20]
Ribophagy is the process in yeasts in which both small and large ribosomal subunits are are delivered to the vacuole through selective macroautophagy. This process is caused by prolonged nitrogen starvation and specific only to ribosomal subunits. [21] Small and large ribosomal subunit ribophagy depends on different and distinct pathways. The increased turnover kinetics of large subunits are dependent upon the Ubp3 ubiquitin protease and its activator Bre5. Increased turnover rates of small subunit are scantly affected when mutations arise in UBE3 or BRE5, indicating independent pathways of ribophagy. However, for small ribosomal subunit ribophagy, the effector remains unknown.[22] As Upb3 deubiquitylation necessary for 60S ribophagy, cells deficient of Upb3 have an increased ubiquitylation level of many ribosome-associated proteins which have yet to be identified. It is believed that the deubiquitylation of Upb3-Bre5 targets aids autophagosomes with the packing of ribosomes, or allows the maturation of autophagosomes and/or its merging with the vacuole. [23]
Ribosome surveillance in human health
[edit | edit source]Alterations in RNA can cause ribosome degradation, some of which can lead to serious diseases. RNA damage are not only results of stress and disease situations, as they occur during normal cell growth as well. Exposure to ultraviolet light, chlorination, oxidation, alkylation and nitration can all result in the introduction of chemical modifications to nucleobases, as well as RNA-RNA and RNA-protein crosslinks, into RNA and RNPs. [24] Many believe that RNA oxidation could be associated with disease progression, and that many factors, including the extent of association with proteins or iron, may be causes to leaving RNA susceptible to oxidative damage. Some cases of ribosomal RNA damage inflicted by UV radiation and oxidation have lead to human neurodegenerative diseases, such as Alzheimer's, Parkinson's, and atherosclerotic plaques. [25] Reactive oxygen species (ROS) is developed by chronological aging, oxidative stress and other apoptotic stimuli. Yeast cells that are exposed to high levels of ROS fragment Mature 5.8S and 25S rRNA considerably. [26] Yeast cells that are given anti-metabolites and the chemotherapeutic agent 5-fluorouracil (5-FU) accrue polyadenylated pre-rRNAs. In exosome complexes that are hypersensitive to the drug, these accumulations are distressed, proposing that nucleolar surveillance is what is responsible for the degradation of 5-FU. Many nucleolar stresses, such as drug-based interference of pre-RNA processing and rRNA synthesis, have lead to nucleolar breakdown, p53 stabilization and cell cycle advancement defects and/or apoptosis. [27]
Ribosomal Functional Specialization
[edit | edit source]In the core of every ribosome, which is the protein factory, there are ribosomal proteins. Recent research indicates that these core proteins can differ between ribosomes under different growth conditions and that these differences confer the ability to more efficiently translate specific subpopulations of mRNA. Ribosomes exist in every living organism for the same purpose, to synthesize proteins from mRNA. Previously, it was thought that ribosomes were homogenous and lacked any structural diversity. Because function follows structure, these differences should be expected to alter a given ribosome’s affinity for more complementary mRNA strands. In addition to core variants, post-transcriptional modification of the ribosomal RNAs leads to additional structural differentiation, and therefore functional specialization. For ribosomes to be functionally specialized it requires that the producing cell be sensitive to changing environmental conditions such that the ribosomes being synthesized exhibit biochemical differences that affect translation. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3056915/
References
[edit | edit source]Edward Ki Yun Leung, Nikolai Suslov, Nicole Tuttle, Raghuvir Sengupta, and Joseph Anthony Piccirilli. "The Mechanism of Peptidyl Transfer Catalysis by the Ribosome." http://www.ncbi.nlm.nih.gov/pubmed/21548786
Sebastian Klinge, Felix Voigts-Hoffmann, Marc Leibundgut and Nenad Ban. "Atomic structures of the eukaryotic ribosome." Zahra Shajani, Michael T. Sykes, and James R. Williamson. "Assembly of Bacterial Ribosomes." http://www.ncbi.nlm.nih.gov/pubmed/21529161
Gilbert WV. "Functional Specialization of Ribosomes?" Trends Biochem Sci. 2011 Mar;36(3):127-32. Epub 2011 Jan 16. PMID: 21242088 [PubMed - indexed for MEDLINE] http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3056915/
- ↑ Lafontaine, Denis. "Elsevier: Article Locator." ScienceDirect.com | Search through over 10 million science, health, medical journal full text articles and books.. N.p., n.d. Web. 20 Nov. 2012. <http://www.sciencedirect.com/science/article/pii/S096800040900245X>.
- ↑ Sebastian Klinge, Felix Voigts-Hoffmann, Marc Leibundgut and Nenad Ban. "Atomic structures of the eukaryotic ribosome."
- ↑ Joan L. Slonczewski, John W. Foster. "Microbiology: An Evolving Science."
- ↑ Sebastian Klinge, Felix Voigts-Hoffmann, Marc Leibundgut and Nenad Ban. "Atomic structures of the eukaryotic ribosome."
- ↑ Sebastian Klinge, Felix Voigts-Hoffmann, Marc Leibundgut and Nenad Ban. "Atomic structures of the eukaryotic ribosome."
- ↑ Lafontaine, Denis. "Elsevier: Article Locator." ScienceDirect.com | Search through over 10 million science, health, medical journal full text articles and books.. N.p., n.d. Web. 20 Nov. 2012. <http://www.sciencedirect.com/science/article/pii/S096800040900245X>.
- ↑ Steitz, T.A. (2008) A structural understanding of the dynamic ribosome machine. Nat. Rev. Mol. Cell Biol. 9, 242–253
- ↑ Henras, A.K. et al. (2008) The post-transcriptional steps of eukaryotic ribosome biogenesis. Cell. Mol. Life Sci. 65, 2334–2359
- ↑ Strunk, B.S. and Karbstein, K. (2009) Powering through ribosome assembly. RNA 15, 2083–2104
- ↑ Lafontaine, D. et al. (1995) The 18S rRNA dimethylase Dim1p is required for pre-ribosomal RNA processing in yeast. Genes Dev. 9, 2470–2481
- ↑ Doma, M.K. and Parker, R. (2007) RNA quality control in eukaryotes. Cell 131, 660–668
- ↑ Doma, M.K. and Parker, R. (2006) Endonucleolytic cleavage of eukaryotic mRNAs with stalls in translation elongation. Nature 440, 561–564
- ↑ LaRiviere, F.J. et al. (2006) A late-acting quality control process for mature eukaryotic rRNAs. Mol. Cell 24, 619–626
- ↑ Lafontaine, Denis. "Elsevier: Article Locator." ScienceDirect.com | Search through over 10 million science, health, medical journal full text articles and books.. N.p., n.d. Web. 20 Nov. 2012.
- ↑ Lafontaine, Denis L.J. "A ‘garbage Can’ for Ribosomes: How Eukaryotes Degrade Their Ribosomes." Trends in Biochemical Sciences 35.5 (2010): 267-77.
- ↑ Lin NY, Beyer C, Gießl A, et al. (September 2012). "Autophagy regulates TNFα-mediated joint destruction in experimental arthritis". Ann. Rheum. Dis.. doi:10.1136/annrheumdis-2012-201671. PMID 22975756
- ↑ Česen MH, Pegan K, Spes A, Turk B (July 2012). "Lysosomal pathways to cell death and their therapeutic applications". Exp. Cell Res. 318 (11): 1245–51. doi:10.1016/j.yexcr.2012.03.005. PMID 22465226.
- ↑ Mizushima N, Ohsumi Y, Yoshimori T (December 2002). "Autophagosome formation in mammalian cells". Cell Struct. Funct. 27 (6): 421–9. PMID 12576635.
- ↑ Česen MH, Pegan K, Spes A, Turk B (July 2012). "Lysosomal pathways to cell death and their therapeutic applications". Exp. Cell Res. 318 (11): 1245–51. doi:10.1016/j.yexcr.2012.03.005. PMID 22465226.
- ↑ Lafontaine, Denis L.J. "A ‘garbage Can’ for Ribosomes: How Eukaryotes Degrade Their Ribosomes." Trends in Biochemical Sciences 35.5 (2010): 267-77.
- ↑ Kraft, Claudine, Anna Deplazes, Marc Sohrmann, and Matthias Peter. "Mature Ribosomes Are Selectively Degraded upon Starvation by an Autophagy Pathway Requiring the Ubp3p/Bre5p Ubiquitin Protease." Nature Cell Biology 10.5 (2008): 602-10.
- ↑ Lafontaine, Denis L.J. "A ‘garbage Can’ for Ribosomes: How Eukaryotes Degrade Their Ribosomes." Trends in Biochemical Sciences 35.5 (2010): 267-77.
- ↑ Kraft, Claudine, Anna Deplazes, Marc Sohrmann, and Matthias Peter. "Mature Ribosomes Are Selectively Degraded upon Starvation by an Autophagy Pathway Requiring the Ubp3p/Bre5p Ubiquitin Protease." Nature Cell Biology 10.5 (2008): 602-10.
- ↑ Wurtmann, Elisabeth J., and Sandra L. Wolin. "RNA under Attack: Cellular Handling of RNA Damage." Critical Reviews in Biochemistry and Molecular Biology 44.1 (2009): 34-49.
- ↑ Nunomura, Akihiko, Tim Hofer, Paula I. Moreira, Rudy J. Castellani, Mark A. Smith, and George Perry. "RNA Oxidation in Alzheimer Disease and Related Neurodegenerative Disorders." Acta Neuropathologica 118.1 (2009): 151-66.
- ↑ Mroczek, S., and J. Kufel. "Apoptotic Signals Induce Specific Degradation of Ribosomal RNA in Yeast." Nucleic Acids Research 36.9 (2008): 2874-888.
- ↑ Lafontaine, Denis L.J. "A ‘garbage Can’ for Ribosomes: How Eukaryotes Degrade Their Ribosomes." Trends in Biochemical Sciences 35.5 (2010): 267-77.
