Structural Biochemistry/Proteins
Protein Structure and Function
[edit | edit source]A protein is a functional biological molecule that is made up of one or more polypeptides that are folded/coiled into a specific structure [1]. Proteins are important macromolecules that serve as structural elements, transportation channels, signal receptors and transmitters, and enzymes. Proteins are linear polymer that are built up of the monomer units called amino acids. There are 20 different amino acids and they are connected by a peptide bond between the carboxyl group and the amino group in a linear chain called a polypeptide. Each protein has different side chains or the "R" groups. Proteins have many different active functional groups attached to them to help define their properties and functions. Proteins cover a wide range of functions, ranging from very rigid structural elements to transmitting information between cells. Each person has several hundred thousands of different proteins in their body. Proteins fold into secondary, tertiary, and quaternary structures based on intra-molecular bonding between functional groups or intermolecular bonding (quaternary only) and can obtain on a variety of three-dimensional shapes depending on the amino acid sequence. All proteins have primary, secondary and tertiary structures but quaternary structures only arise when a protein is made up of two or more polypeptide chains [1]. The folding of proteins is also driven and reinforced by the formation of many bonds between different parts of the chain. The formation of these bonds depends on the amino acid sequence. The study of their structures is important because proteins are essential for every activity in the human body as well as they are the key components of biological materials. Primary structure is when amino acids are linked together by peptide bonds to form polypeptide chains. Secondary structure is when the polypeptide chains fold into regular structures like the beta sheets, alpha helix, turns, or loops. A functional protein is much more than just a polypeptide, it is one or more polypeptides that have been precisely folded into a molecule with a very specific, unique shape which is critical to its function [1].
Proteins are usually portrayed in 3D structures and categorized into four different characteristics and levels:

Primary: The primary structure of a protein is the level of protein structure which refers to the specific sequence of amino acids [1]. When two amino acids are in such a position that the carboxyl groups of each amino acid are adjacent to each other, they can be combined by undergoing a dehydration reaction which results in the formation of a peptide bond [1]. Amino acids in a polypeptide (protein) are linked by peptide bonds that begin with the N-terminal with a free amino group and ends at C-terminal with a free carboxyl group. rts . The peptide bond is planar and cannot rotate freely due to a partial double bond character. While there is a restricted rotation about peptide bond, there are two free rotations on (N-C) bond and (C-C) bond, which are called torsion angles, or more specifically the phi and psi angles. The freedoms of rotation of these two bonds are also limited due to steric hindrance. Genes carry the information to make polypeptides with a defined amino acid sequence. An average polypeptide is about 300 amino acids in length, and some genes encode polypeptides that are a few thousand amino acids long. It's important to know the primary structure of the protein because the primary structure encodes motifs that are of functional importance in their biological function; structure and function are correlated at all levels of biological organization [1].
Secondary: The amino acid sequence of a polypeptide, together with the laws of chemistry and physics, cause a polypeptide to fold into a more compact structure. Amino acids can rotate around bonds within a protein. This is the reason proteins are flexible and can fold into a variety of shapes. Folding can be irregular or certain regions can have a repeating folding pattern. The coils and folds that result from the hydrogen bonds between the repeating segments of the polypeptide backbone are called secondary structures [1]. Although the individual hydrogen bonds are weak, they are able to support a specific shape for that part of the protein due to the fact that they are repeated many times over a long part of the chain [1]. Secondary structures of a protein are proposed by Pauling and Corey. Its structures are formed by amino acids that are located within short distances of each other. Because of the planar nature of the peptide bonds, only certain types of secondary structure exist. The three important secondary structures are α-helix, β-sheets, and β-turns. Also, the beta sheets can be parallel, antiparallel, or mixed. Antiparallel beta sheets are more stable because the hydrogen bonds are at a ninety degree angles. The a-helix is a coiled structure stabilized by intrachain hydrogen bonds.
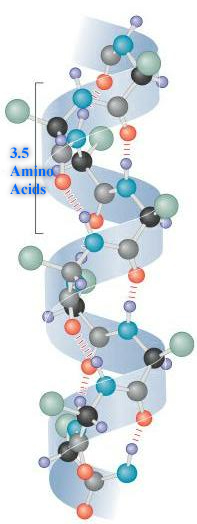
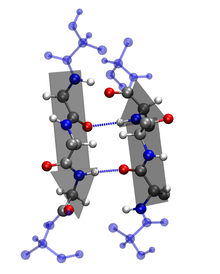
Characteristics of the Secondary Structures:
1. α-helix: In an α-helix, the polypeptide backbone forms a repeating helical structure that is stabilized by hydrogen bonds between a carbonyl oxygen and an amine hydrogen. These hydrogen bonds occur at regular intervals of one hydrogen bond every fourth amino acid and cause the polypeptide backbone to form a helix [1]. The most common helical structure is a right-handed helix with its hydrogen bonds parallel to its axis. The hydrogen bonds are formed between carbonyl oxygen and amine hydrogen groups of four amino acid residues away. Each amino acid advances the helix, along its axis, by 1.5 Å. Each turn of the helix is composed of 3.6 amino acids; therefore the pitch of the helix is 5.4 Å. There is an average of ten amino acid residues per helix with its side chains orientated outside of the helix. Different amino acids have different propensities for forming x-helix, however proline is a helix breaker because proline does not have a free amino group. Amino acids that prefer to adopt helical conformations in proteins include methionine, alanine, leucine, glutamate and lysine (malek).
2. β-sheet: ß-sheets are stabilized by hydrogen bonding between peptide strands. In a β-sheet, regions of the polypeptide backbone come to lie parallel to each other and are connected by hydrogen bonds [1]. The hydrogen bonds are formed between the carbonyl oxygen and the amine hydrogen of amino acid in adjacent strands in a polypeptide, which means that the hydrogen bonds are inter-stand. β-sheet regions are more extended than an α-helix, and the distance between adjacent amino acids is 3.5 Å. Hydrogen bonding in β-strand can occur as parallel, anti- parallel, or a mixture. Amino acid residues in β- parallel configuration runs in the same orientation. Pleated sheets makes up the core of many globular proteins and also are dominant in some fibrous proteins such as a spiders web [1]. The large aromatics such as: tryptophan, tyrosine and phenylalanine, and beta-branched amino acids like: isoleucine, valine, and threonine prefer to adopt β-strand conformations.This orientation is energetically less favorable because of its slanted, non-vertical hydrogen bonds. Trytophan, tyrosine, and phenylalanine are hydrophobics while the other amino acids are hydrophilics.
3. β-turns: Poly peptide chains can change direction by making reverse turns and loops. Loop regions that connect two anti-parallel β-strands are known as reverse turns or β-turns. These loop regions have irregular lengths and shapes and are usually found on the surface of the protein. The turn is stabilized by hydrogen bond between the backbone of carbonyl oxygen and amine hydrogen. The CO group of the residue, in many reverse turns, which is bonded to the NH group of residue i + 3 . The interaction stabilizes abrupt changes in direction of the polypeptide chain. Unlike the alpha-helices and ß-strands, loops do not have regular periodic structures. However, they are usually rigid and well defined. Since they loops lie on the surface of the proteins, they are able to participate in interactions between proteins and other molecules. Ramachandran plot is a plot that shows the available torsion angles of where proteins can be found. However, in the plot, if there are many dots that locate all over the place, it means that there exists a loop.
Tertiary: As the secondary structure becomes established due to the primary structure, a polypeptide folds and refolds upon itself to assume a complex three-dimensional shape called the protein tertiary structure. Tertiary structure is the overall shape of a polypeptide.[1] Tertiary structure results from the interactions between the side chains (R groups) of the various amino acids [1]. This three dimensional structure is due to intramolecular interactions between the side groups along the polypeptide chain. Its domain typically contains 300 – 400 amino acids, and it adopts a stable tertiary structure when it is isolated from their parent protein. As a polypeptide folds into its functional shape, amino acids that have hydrophobic side chains tend to end up clustered at the core of the protein so that they are out of contact with water [2]. Covalent bonds called disulfide bridges can also affect the shape of a protein [1]. Disulfide Bridges form where two amino acids containing sulfhydryl groups on their side chains are brought close together by how the protein is folding [1]. For some proteins, such as ribonuclease, the tertiary structure is the final structure of a functional protein. Other proteins are composed of two or more polypeptides and adopt a quaternary structure.
Quaternary:
While all proteins contain primary, secondary and tertiary structures, quaternary structures are reserved for proteins composed of two or more polypeptide chains [1]. Proteins that have quaternary structures contain more than one polypeptide and each adopt a tertiary structure and then assemble with each other via intermolecular interactions. The quaternary structure of a protein is the overall structure that is the result of the addition of these polypeptide subunits [1]. The individual polypeptides are called protein subunits, which means different polypeptides folded separately. Subunits may be identical polypeptides or they may be different. When proteins consist of more than one polypeptide chain, they are said to have quaternary structure and are also known as multimeric proteins, meaning proteins consisting of many parts. Quaternary structures can also defined as when more than one protein come together to create either a dimer, trimer, tetramer, etc... [2]. Hemoglobin is an example of a quaternary structure that is composed of two alpha subunits and two beta subunits.

Globular and Fibrous Proteins:
[edit | edit source]Fibrous proteins: Fibrous proteins also known as Schleroprotein are long protein chains shaped liked rodwires. Unlike Globular Protein, they do not denature as easily, and contain many repeats of secondary structures. They are mostly structural proteins that are responsible for organisms in support and protection such as forming connective tissue, muscle fibers, bones, and tendons . The two examples of fibrous proteins are:
1. α –keratin: α –keratin (essential in hair, hooves, horn, fingernails, and etc.) is a coiled-coil protein composed of two intertwining α-helices. Coiled-coil structures are found in other structural proteins, for example, the myosin of skeletal muscle; it has heptads repeats correspond to 3.5 amino acids per turn. Residues in the position of a, d, a’ and d’ in the helices of these proteins are usually hydrophobic. The two strands in a coiled-coil are held together by hydrophobic interaction as well as ionic interactions and disulfide bonds.
2. Collagen: Collagen (of tendon, cartilage, blood vessel walls) is the most abundant protein in human’s body. Collagen is a triple helix that is unlike α-helix, it has 3.3 amino acids and 10 Å per turn. Collagen is stabilized by hydrogen bonds, which is formed between the carbonyl oxygen and the amine hydrogen of amino acids situated on neighboring chains and is perpendicular to the fiber axis. It is abundant in proline, and contains hydroxyproline and hydroxlysine. However, due to the abundance of proline, there are no intrachain of hydrogen bonds, and the hydroxylation of proline and lysine requires Vitamin C. Vitamin C deficiency causes scurvy. One third of amino acids of collagen are glycine because of overcrowding; only glycines are found in the center of the collagen molecules. Collagen molecules can be cross-linked by covalent bonds to from larger fibers and sheets.
Globular Protein: Globular proteins are folded to bury the hydrophobic side chains. All globular proteins have an inside where the hydrophobic core is arranged. It has an outside toward which the hydrophilic groups are directed. The uncharged polar amino acid residues are usually found on the protein surfaces but it can also occur in the interior. In the latter case, it will hydrogen bonded to other groups, i.e. ser, thr, tyr are all polar, uncharged.
Factors that influence protein structure:
[edit | edit source]Several factors determine the way that polypeptides adopt their secondary, tertiary and quaternary structures. The amino acid sequences of polypeptides are the defining features that distinguish the structure of one protein from another. As polypeptides are synthesized in a cell, they fold into secondary and tertiary structures, which assemble into quaternary structures for most proteins. As mentioned, the laws of chemistry and physics, together with amino acid sequence, govern this process. Five factors are critical for protein folding and stability:
1. Hydrogen bonds: Hydrogen bonds are formed between a hydrogen bond donor and hydrogen bond acceptor. For amino acids, hydrogen bonding would occur between the backbone of the amine group and the oxygen of the carbonyl group.
2. Ionic bonds: Electrostatic interactions occur between two oppositely charged molecules. Ionic interactions are weaker in water than in vacuum, this is due to a different dielectric constant faced in water between opposing charges within the protein's structure.
3. Hydrophobic effect: The hydrophobic interaction originates from the tendency of non-polar molecules to minimize their interactions with water. When non-polar molecules interact with water, these molecules tend to cluster together in the center to form a micelle.
4. Van der waals forces: Van der waals forces exist between non-polar molecules at close range. Of the three van der waals interactions, interactions between permanent dipoles is the strongest, dipole-induced dipole interactions are weaker than permanent dipole and the London dispersion forces are the weakest. While van der waals forces between individual atoms are weak, the sum of van der waals forces resulting from interactions between many atoms in large macromolecules can be substantial. The strength of van der waals interactions varies with the distance between the atoms and is maximal at the van der waals contact distance.
5. Disulfide bridges: A disulfide bond can be form between two cysteines through oxidation. These are also the strongest covalent bonds within a protein's tertiary structure.
Protein denaturation:
[edit | edit source]Protein denaturation: is the loss of native conformations of tertiary structure. Denaturing proteins experience either the destruction of disruption of internal tertiary or secondary structure. Denaturation however, does not break the peptide bond between adjacent amino acids, thus not affecting the primary structure of the protein. Denaturation however, will interfere the normal alpha-helix and beta sheets in a protein which ultimately distort its 3D shape.
Denaturation causes the disruption of hydrogen bonding between close proximity amino acids, thus interfering a protein's secondary and tertiary structure. In tertiary structure there are four types of bonding interactions between "side chains" including: hydrogen bonding, ionic bridges, disulfide bonds, and hydrophobic intermolecular interactions. In other words, there are several different conditions to denature the conformation of a protein.
Conditions that denature proteins:
1. Extreme pH (pH < 4 or pH > 9) : alters H-bonding
2. Heat (temp >70oC): thermal effect, disrupts weak forces of non-covalent bonds
3. Detergents or organic solvents : disrupts hydrophobic interaction
4. Chaotropic agents (high concentrations) : e.g., urea and guanidinium chloride
Activation of specific proteins require positioning at their specific sites
[edit | edit source]As scientists started discovering more aspects of chemistry, they've actually found the magnitude of complexity in cell chemistry/biology. Although scientists found out that protein had an imperative role in the body, they've also discovered that the proteins assemble themselves at a specific site in the cell, being activated only when necessary. Using the GFP- tagged proteins (fluorescence) in animate cells, the positioning and repositioning of proteins were observed in response to the specific signals. When extracellular signal molecules bind to the receptor proteins, it reels in different proteins towards the inner area of the plasma membrane to create protein apparatus that will pass on the signal.

Humans have 10 PKC enzymes that differ both in their regulation and in their functions. When the PKC gets activated it will move from the cytoplasm to various intracellular locations and will eventually form specific complexes with other proteins thus allowing them to phosphorylate different protein substrates. Various ligases express this kind of behavior such as SCF ubiquitin ligases. These mechanisms involve the collaboration between protein phosphorylation and scaffold proteins that link specific activating, inhibiting, adaptor, and substrate proteins to a discrete part of a cell.

This occurrence is called induced proximity, which describes the reason why minute different forms of the enzymes with the same reaction sites can have different functions. This can be done by covalently modifying the protein’s location in various ways. These alterations construct binding sites on the proteins so that it would bind to scaffold proteins, making them cluster together so that different reactions can take place within a specific location of a cell. Scaffolds therefore allow the cells to group reactions without the need of membranes.
Scaffold proteins were thought to hold the proteins in specific locations relative to each other but in reality, unstructured regions of polypeptide chains connect the proteins that are interacting. This allows the proteins to frequently clash with each other in random orientations, some leading to successful reactions. The tethering of the proteins allows faster reaction rates to occur. Scaffold proteins therefore provide flexible methods of controlling the Cell Chemistry.
DEAD Box Proteins
[edit | edit source]
DEAD box proteins consist of RNA helicases, they are involved in RNA metabolism processes, and they are conserved in nine domains found in bacteria and viruses to humans. They are 350 amino acids in length. DEAD box proteins are involved in pre-mRNA processing, splicesosome formation, and rearranging of ribonucleoprotein (RNP) complexes. DEAD box proteins are required in the pre-mRNA splicing and the in vivo splicing process. During the pre-mRNA processing, the DEAD box proteins unwind to provide energy to rearrange the five snRNPs (U1, U2, U4, U5, and U6) required in pre-mRNA splicing. In the in vivo splicing, three DEAD box proteins, Sub2, Prp28, and Prp5, are needed. Prp5 helps rearrange the conformation of U2, which allows the U2 sequence bind to the branch point sequence. Prp28 helps the recognition of the 5’ splicing location.
The first DEAD box protein, the ElF4A translation initiation factor, are dependent on RNA ATPase activity. This protein helps unwind the secondary structure, which stops the scanning
References
[edit | edit source]Molecular biology of the cell 5th ed. alberts
Biochemistry 6th ed. Berg, Tymoczko and Stryer
Campbell Biology, 9th Edition, Neil Campbell and Jane Reece, 2010
