Structural Biochemistry/Proteins/Signaling Control of Splicing Proteins
Overview
[edit | edit source]Technology advances in sequencing and microarrays allow for us to better understand pre-mRNA splicing patterns in different cells. For example, cellular splicing changes when it is stimulated by factors such as DNA damage, neuron depolarization and or metabolic changes in cells. In the last few years, there have been more studies regarding patterns in mechanisms that relate cellular stimuli to downstream alternative splicing control. Some of these splicing events include degradation of splicing factors, altered nuclear translocation, and regulated synthesis of splicing factors.
What is alternative splicing and how does it work?
[edit | edit source]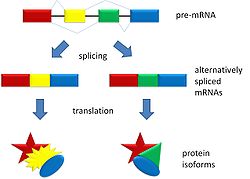
Alternative splicing is a process that occurs during gene expression and allows for the production of multiple proteins (protein isoforms) from a single gene coding. Alternative splicing can occur due to the different ways in which an exon can be excluded or included from the messenger RNA. It can also occur if portions on an exon are exclude/included or if there is an inclusion of introns. For example, if a pre-mRNA has four exons (A, B, C, and D) these fours exons can be spliced and translated in a number of different combinations. Exons A, B, and C can be translated together or Exons A, C, and D can be translated. This is what results in alternative splicing.
The pattern of splicing and production of alternatively spliced messenger RNA is controlled by the binding of regulatory proteins (trans-acting proteins that contain the genes) to cis-acting sites that are found on the pre RNA. Some of these regulatory proteins include splicing activators (proteins that promote certain splicing sites) and splicing repressors (proteins that reduce the use of certain sites). Some common splicing repressors include: heterogeneous nuclear ribonucleoprotein (hnRNP) and polypyrimidine tract binding protein (PTB). Proteins that are translated from alternatively spliced messenger RNAs differ in the sequence of their amino acids and this results in altered function of the protein. This is the reason why the human genome can encode a wide diversity of proteins. Alternative splicing is a common process that occurs in eukaryotes; most of the multi-exonic genes in humans are spliced alternatively. Unfortunately, abnormal variations in splicing are also the reason why there are many genetic diseases and disorders.

Spliceosome
[edit | edit source]The splicing of messenger RNA is accomplished and catalyzed by a macro-molecule complex known as the spliceosome. The areas for ligation and cleavage are determined by the many sub-units of the spliceosome. These sub-units include the branch site (A) and the 5' and 3' splice sites. Interactions between these sub-units and the small nuclear ribonucleoproteins (snRNP) found in the spliceosome create a spliceosome A complex which helps determine which introns to leave out and which exons to keep and bind together. Once the introns are cleaved and removed, the exons are joined together by a phosphodiester bond.
Regulatory Proteins
[edit | edit source]As noted above, splicing is regulated by repressor proteins and activator proteins, which are are also known as trans-acting proteins. Equally as important are the silencers and enhancers that are found on the messngerRNAs, also known as cis-acting sites. These regulatory functions work together in order to create splicing code that determines alternative splicing. The cis-acting sites will be discussed here.

Splicing silencers are regulatory sites that are found in pre-messengerRNA's and are where the splicing repressor proteins bind to. When the repressor binds to the silencer site, it reduces the chance that a site close-by will be chosen as a splicing junction. These silencer sites can be found on introns or on exons. When found on introns, these sites are known as intronic splicing silencers and on exons they are called exonic splicing silencers. The sequences found on these sites are numerous and that allows for different kinds of proteins to bind.

On the other hand, splicing enhancers are regulatory sites where splicing activator proteins can bind to. When the activator protein binds to the enhancer site, it increases the chance that a site close-by will be chosen as a splicing junction. Just like the splicing silencers, these sites can also be found in introns and exons. In introns they are called intronic splicing enhancers and in exons they are called exonic splicing enhancers. However, unlike their silencer counterparts, enhancer sites usually allow the binding of activator proteins that belong to the family of SR proteins. These proteins are rich in arginine and serine.
How is alternative splicing regulated by some specific signals?
Alternative splicing has been recently revealed to occur in nearly all human genes. Most typically, a specific exon may be either included or excluded in different cell types or growth conditions when alternative splicing occurs. In each case, the pattern of splicing, the binding of regulatory proteins to cis-acting auxiliary sequences generally determines the pattern of splicing and these sequences in turn control where the binding occurs and/or how the enzymatic complex reacts at neighboring splice sites. (Combinatorial Regulation of Alternative Splicing)
Importantly, the open reading frame of the resultant mRNA or the presence of cis-regulatory elements that control mRNA stability or translation can be altered by any of these above differential patterns. Therefore, shaping the proteome of any given cell requires the precise control of alternative splicing , and how the cellular function responses to changing environmental conditions can also be significantly altered by changes in splicing patterns.

Combinatorial Regulation of Alternative Splicing The spliceosome is a macromolecular complex that catalyzes the removal of introns and the basic joining of extrons. The binding of various subunits of the spliceosome in order to sequence elements at the intron and extron boundaries in a pre-mRNA determines the precise sites of ligation and cleavage. Those subunits are the 5 splice site, the branch point sequence, a pyrimidine-rich track, and the 3 splice site. However, for mammals, the splice sites are poorly conserved; hence, they are typically not sufficient to bind the spliceosome with high affinity. The efficiency of spliceosomal binding via mechanisms can be impacted by proteins bound to non-splice site sequences within the exon or intron. Exonic or intronic splicing enhancers are the sequences that help promote spliceosomal recognition of an exon, while the splicing silencers are needed to inhibit recognition of the exon. Exon inclusion (green ovals) is promoted by the binding of the enhancers of members of the ubiquitously expressed SRSF protein family, while the exon usage is repressed by members of the hnRNP family of proteins via silencer elements (red ovals). FOX, CELF, neuro-oncological ventral antigen (NOVA) and muscleblind-like (MBNL) proteins are some other splicing regulators that are more tissue restricted and these regulators function equally as enhancers and repressors of splicing through mechanisms that are still largely undefined. Therefore, the ratio of mRNA isoform expression can frequently be altered by the binding of single regulatory proteins or the subtle changes in the balance of expression.
-
Neuro-oncological ventral antigen 1
-
Muscleblind-like splicing regulator 1
Post-Translational Modification of Splicing Proteins
Phosphorylation, acetylation, methylation, sumolylation and hydroxylation are involved in the modification of splicing regulatory proteins in many cases. The phosphorylation of the extensive Arg-Ser dipeptides found within SR proteins is the best characterized modification. The extensive post-translational modifications also includes the HnRNP proteins, along with other non-SR splicing factors.
Alternative Splicing and its Signals
[edit | edit source]
Recently, technical tools such as deep sequencing and sensitive microarrays have opened up for more knowledge of alternative splicing events. Almost all human genes go through some sort of alternative splicing, which includes differential exclusion or inclusion of a specific exon, exclusion of a part of an exon, and inclusion of introns and exons. These differential trends can change the reading frame of the processed mRNA or alter any cis-regulatory factors that monitor mRNA translation or stability. For that reason, the regulation of alternative splicing is crucial in shaping the proteome of cells; alterations in splicing patterns can change functions in cells in response to environmental changes. Observations in heart tissue in its development stage, pre and post depolarization of neurons and cells before and after apoptosis have showed that alternative splicing events play a large role in the functional outcome of the signaling and developmental processes.
Since alternative splicing is generally determined by binding regulatory proteins to auxiliary sequences that control the location of binding and activity of the enzymatic complex at neighboring sites of splicing, it is used in response to DNA damage and T cell activation. One case for DNA damage includes the alternative splicing of the E3 ubiquitin ligase murine double minute-2 (MDM2). MDM2 specifically controls levels of p53, a tumor-suppressing gene, by targeting it for proteasomal degradation. Once DNA damage is perceived, Mdm2 exons are skipped to reduce the functioning of MDM2, thus allowing p53 to accumulate. This induced regulation of MDM2 provides an example of how splicing that is coupled with transcription as the exon skipping mimics the damaged DNA. In this case, cells show a "tight control of alternative splicing that helps regulate protein expression due to changing conditions in the cell."[3]
Altering the interactions of proteins is another method in which alternative splicing can be achieved. One demonstration of this is T cell activation. In T cell activation, alternative splicing is used similarly in DNA damage where the altered protein interaction with other proteins regulate the splicing of, specifically, the CD45 gene during T cell activation. In resting T cells, PSF, a RNA binding protein, is phosphorylated by the enzyme GSK3 and this causes the phosphorylated PSF to form a complex with TRAP150. As a result, the PSF cannot bind to the CD45 RNA. This prevents any possible exon exclusion and results in no participation in splicing. However, in an activated T cell, there is little to none GSK3 due to an inhibiting phosphorylation because an antigen binds to the T cell receptor and causes GSK3 activity to drop. Without the GSK3, PSF is not bound to the TRAP150 and is free to bind to the RNA. This is a major example of how splicing is controlled by signal-induced changes in transcription.
RNA-binding Proteins Regulate Splicing
[edit | edit source]Altering the level of expression of a regulatory protein is the most simple way that can affect alternative splicing. A small change in the expression of one splicing factor can change the elements that determine exon exclusion or inclusion, due to the complex influences on a given transcript. The control of transcriptional activators such as nuclear factor-kappa B and nuclear factor of T-cells have been proven to be altered by signaling pathways. Therefore, signaling induce transcription of genes encoding SR proteins or other splicing regulators that can change the splicing of genes that respond to these factors. In one instance, it is proposed that stimulation of T cells trigger the splicing signal of the gene that encodes tyrosine phosphatase CD45. Furthermore, the proteins PTB-associated splicing factor and hnRNP L-like activate the elimination of CD45 exons 4 and 6. Interestingly enough, inducible changes in protein expression do not only result from transcription. As shown in the splicing regulatory protein CELF1, its increased protein levels is due to an increase in the stability and phosphorylation of CELF1, which then leads to the overall up-regulated steady-state levels. This increase in phosphorylation is also responsible for the protein kinase C activity in DM cells. Not surprisingly, the increase in protein stability also has other regulations; it is also controlled by miRNAs during heart development. The two coupled- mechanisms highlight the idea that regulating regulatory protein expression is important to keeping a proper splicing pattern required for functions in cells. [3]
Localization of RNA-binding proteins
[edit | edit source]In addition to the method of protein expression and stability mentioned above, alternative splicing can occur when signals are changed due to the localization of regulatory proteins. Many of the regulatory proteins, such as SR proteins and hnRNP mentioned above, have to travel to and from the nucleus and cytoplasm. As a result, the relative distribution of these regulatory proteins in the nucleus versus the cytoplasm can alter signaling pathways. These altered pathways will lead to splicing differences. Two regulatory proteins that have their distributions regulated include SRPK1 and hnRNP proteins (hnRNP A1 specifically). In the case of SRPK1, this regulatory protein is normally found in the cytoplasm due to interactions with heat shock proteins. However, when the cell undergoes osmotic shock the SRPK1 proteins move to the nucleus and cause phosphorylation of SR proteins. This phosphorylation results in different interactions between the proteins and their target genes and produces varying splicing patterns. In the case of hnRNP, osmotic shock actually has an opposite effect on the localization of this protein in relation to SRPK1. hnRNP is also normally found in the cytoplasm but as opposed to SRPK1, osmotic shock does not cause it to move to the nucleus. In fact, phosphorylation of hnRNP prevents it from entering the nucleus.
Feedback Loops in Alternative Splicing
[edit | edit source]
As all living things go through homeostasis, cells do the same. In order for cells to practice homeostasis, they must therefore turn off induced splicing signals once conditions are normal again. For example, these regulations can include getting rid of antigen, DNA repair and neurons repolarization. One way to reset gene expression is to deactivate signals by removing the initial receptors or signaling factors themselves. Of course, receptors such as phosphatases and kinases undergo autoinhibitory signal-induced alternative splicing. For instance, in response to T cell activation, alternative splicing of CD45 will reduce the sensitivity of the cell to receive antigen stimulation signals. In another example, molecules that encode kinases responsible for T-cell signaling activation such as the FYN proto-oncogene, signal-regulated kinase-1, and tyrosine kinase 2 beta protein, all go through alternative splicing due to T cell activation to lessen expression or change localization patterns.
Inducing expression of an opposing regulatory factor can help in resetting the induced splicing signals. Neuron chronic depolarization is an example of this, which results in increased skipping of exons controlled by CaRREs. Some of these CaRRE-reduced exons appear again in prolonged depolarization. This splicing pattern is related to CaMK-induced alternative splicing of FOX1 that encodes RNA-binding proteins. FOX1 regulates the splicing patterns of genes involved with synaptic activity. In addition, many genes controlled by CaRREs also have a FOX1 binding site which can have an antagonistic effect on exon inclusion like that of the CaRRE sequence. Since most studies only regulate a few genes, many further studies are needed to have a fuller grasp of alternative splicing that occurs in the downstream of a given pathway. [3]
What is next for protein splicing?
[edit | edit source]Despite the stimulated factors mentioned above, the overall picture of how signaling pathways regulate alternative splicing is far from being complete. The study of these signaling pathways is still very much in progress. The methods introduced here usually correspond to the alternative splicing of only a few genes. As a result, more progress needs to be made in order to understand the alternative splicing of an entire pathway.
References
[edit | edit source]1. Black, Douglas L. (2003). "Mechanisms of alternative pre-messenger RNA splicing". Annual Reviews of Biochemistry 72 (1): 291–336.
2. Clark, David (2005). Molecular biology. Amsterdam: Elsevier Academic Press.
3. Heyd, Florian, and Kristen W. Lynch. DEGRADE, MOVE, REGROUP: signaling control of splicing proteins Philadelphia: Trends in Biochemical Sciences, 2011. Print.
4. Matlin, AJ; Clark F, Smith, CWJ (May 2005). "Understanding alternative splicing: towards a cellular code". Nature Reviews 6 (5): 386–398.
5. Nilsen, T.W. and Graveley, B.R. (2010) Expansion of the Eukaryotic Proteome by Alternative Splicing. Nature 463, 457-463.
6. Pan, Q; Shai O, Lee LJ, Frey BJ, Blencowe BJ (Dec 2008). "Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing". Nature Genetics 40 (12): 1413–1415.
7. Images: Wiki-Media Commons
8. Heyd F, Lynch KW. Trends Biochem Sci. 2011 Aug;36(8):397-404. Epub 2011 May 17. Review. 2. PMID: 21596569 [PubMed - indexed for MEDLINE]
9. Barash, Y; et al (2010). "Deciphering the splicing code". Nature 465 (7294): 53–59.
10. Wang, Z; Burge, Cb (2008). "Splicing regulation: from a parts list of regulatory elements to an integrated splicing code"