Structural Biochemistry/Polynucleotide Kinase/Phosphatase enzyme
Polynucleotide Kinase/Phosphatase
[edit | edit source]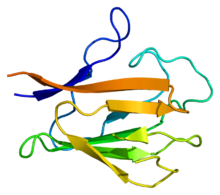
One of the most common problems that could occur in a living organisms is damage to cellular DNA, or mutations. Mutations are so common that it is involved with a variety of important factors in life, such as cancer treatment, neurological disorders, and aging. The basic of idea of DNA damage is that internal and external agents cause the loss of bases, causing the DNA strand to break. Before the strand breaks can be repaired, the termini of breaks require processing before the missing bases can be properly replaced. This is where Polynucleotide Kinase/Phosphatase (PNKP) comes into play by catalyzing the restoration of 5'-phosphate and 3'-hydroxyl termini. PNKP interacts with various other proteins, especially XRCC1 and XRCC4, and uses the different pathways to repair DNA. The 5' kinase and 3'phosphate activities of PNKP processes, or repairs, both single and double stranded termini in DNA. Understanding its mechanism has lead to an opportunity to treat diseases and cancer. PNKP inhibitors are also known to sensitize cells towards IR and chemotherapeutic agents since they prevent PNKP from processing DNA repair. PNKP’s role in the restoration of DNA strand breaks follows three major DNA repair pathways: single-stranded break repair (SSBR), base excision repair (BER), and double-stranded break repair (DSBR). These three mechanisms can provide useful information on how other proteins bind and react to the PNKP enzyme.[1]
Molecular Structure of PNKP
[edit | edit source]PNKP is a multi-domain enzyme that is made up of two main domains: N-terminal forkhead-associated (FHA) domain and the C-terminal catalytic domain. Additionally, C-terminal catalytic domain is composed of fused phosphate and kinase sub-domains. The two main domains, FHA and catalytic domain, are binded through a flexible polypeptide segment. It is at this connection where the binding to CK2-phosphorylated regions of other proteins. The polynucleotide kinase phosphatase and the phage T4 polynucleotide kinase (a cloning enzyme) differ such that the T4 enzyme does not contain the FHA domain whereas PNKP has the FHA domain in the N-terminal. However, the kinase subdomain of the T4 polynucleotide kinase is in the N-terminal instead of the C-terminal catalytic domain.[1] The two specific proteins PNKP interact with are called XRCC1 and XRCC4. The major distinction between the proteins XRCC1 and XRCC4 is that XRCC1 repairs DNA single-stranded breaks while the XRCC4 protein fixes the DNA double-stranded breaks.[1] Only the FHA domain binds to a the region of these proteins that are specifically phosphorylated by CK2. For XRCC1, there are clustered regions of CK2, typically between residues 515 and 526, that are required for it to bind to FHA and repair DNA. In contrast, only a primary CK2 site is required for XRCC2. In addition, aprataxin and aprataxin and PNKP-like factor (APLF) contribute to DNA repair; these two DNA repair factors have FHA domains as well.[1]
PNKP also contains two catalytic active sits that reside on the same side of the enzyme, as well as separate ATP and DNA binding sites. The different DNA binding sites significantly differ between phage and mammalian enzymes. For phage enzymes, the binding site occurs through a narrow channel that leads to catalytic aspartic acid residues, which only aids in single-stranded repair. As for mammalian enzymes, they phosphorylate the 5'-hydroxyl terminal repairing double-stranded more efficiently than single-stranded.
PNKP in Single-Stranded Break Repair (SSBR)
[edit | edit source]Specifically for IR induced strand breaks, the loss of nucleotides is repaired with a process that is carried out with poly(ADP-ribose) polymerase (PARP), XRCC1, and AP endonuclease I (APE1). This process may be done with a short patch, using DNA polymerase and DNA ligase III. It can also be done with a long patch, which uses DNA polymerase, ligase I, and FEN1 endonuclease. The role of APE1 is to remove 3'-phosphoglycolates. PNKP itself performs hydrolysis in 3'-phosphate groups while keeping 5'-OH phosphorylated, which is a much stronger activity than APE1. The overall phosphate activity is significantly more active than the kinase activity, causing an over-expression of phosphatase-detective PNKP.
The basic mechanism of SSBR is:
- Breaks are recognized by PARP
- Recognition attracts only XRCC1, which is typically binded with DNA ligase
- XRCC1 recruits PNKP/APE1
- Recruited enzymes restore termini, allowing DNA polymerase to add missing nucleotides and DNA ligase to bind the broken strands.
Stressed or damaged cells really need the CK2-phosphorylated XRCC1 to bind to FHA domain of PNKP so that repair can occur. Unstressed cells are able to cope with non-phosphorylated proteins because repair is not in urgent need.
PNKP in Double-Stranded Break Repair (DSBR)
[edit | edit source]The double-stranded break repair pathway depends on the protein XRCC4, which when phosphorylated will not stimulate the enzyme PNKP; reversely, unphosphorylated XRCC4 will activate PNKP. However, the combination of phosphorylated XRCC4 with DNA ligase IV can trigger PNKP by promoting the binding between the PNKP’s FHA domain and the phosphorylated XRCC4 protein.[1] PNKP is only involved with the nonhomologous end joining (NHEJ) pathway of DSBR, but not with homologous recombination. The mechanism is similar to SSBR, except that kinase activity of PNKP is required for ligation to occur. Also, phosphorylation is actually dependent on XRCC4, not XRCC1, in order to bind PNKP to DNA ligase, which stimulates the DNA ligase. It is also noted that XRCC4 is necessary for cell survival after IR or chemo treatment. The most important and distinct fact about DSBR is that phosphorylated XRCC4 actually fails to bind to PNKP through the FHA domain, inhibiting cell repair. However, adding DNA ligase reverses the inhibition and therefore allows successful DNA repair.
Non-phosphorylated XRCC4 for DSBR works in a manner similar to XRCC1 in Single-Stranded Break Repair (SSBR) because they both encourage enzymatic cell turnover for Polynucleotide Kinase/Phosphotase (PNKP). Although non-phosphorylated XRCC4 has an attraction to the T-Terminal Fork-Head Associated (FHA) Domain of PNKP, the phosphorylated XRCC4 promotes even greater attraction, which is then better for DNA double-strand repair. Together, the non-phosphorylated and phosphorylated XRCC4 both works in a complex way with DNA ligase and PNKP in order to repair DNA ends.
PNKP in Base Excision Repair (BER)
[edit | edit source]The repairs of most minor base modification that are caused by ionizing radiation, reactive oxygen species and alkylating agents are repaired through the process known as base excision repair (BER). The first step of this process involves DNA glycosylase and its removal of the modified base. This step is then followed by the cleavage of the DNA at the newly formed apurinic/apyrimidinic (AP) site by AP endonuclease I (APE1). The existence of PNKP and its function in the BER pathway became known when nei endonuclease VIII-like-1 (NEIL1) and NEIL2 mammalian DNA glycosylases were discovered, which are proteins that help induc 3’-phosphate termini. NEIL 1 and NEIL 2 (nei endonucleases VII-like 1 and 2 respectively) can form complexes containing PNKP or other BER components. NEIL 1 and NEIL 2 repair DNA by excising, or cutting out, the damaged DNA base and then removing the errors. The competition between the NEIL glycosylases may lead to a base excision repair pathway that does not depend on the enzyme APE1 (AP endonuclease 1).[1] Both of these newly found proteins, even though were found to not bind directly to PNKP, were found to be associated with larger components of the BER process, among which is PNKP. These glycosylases are able to cleave non-basic pair sites and cause sensitive of PNKP depleted cells to the methyl methanesulfonate (MMS), an alkylating agent. It is still unclear exactly what PNKP’s function is in BER, but in studying NEIL1 and NEIL2 more in depth, these proteins could possibly provide explanations regarding PNKP’s role and function in this repair pathway.
Clinical Role of PNKP
[edit | edit source]Through PKNP's elaborate structure, we can find out about the complex function that allows it to help mend the loose DNA strands that have been damaged. One of the innovative purposes that PNKP can serve in clinical research within the human race is that it could help defend cells from radiation damage, which could be beneficial in the clinical field of cancer chemotherapy. Because of PNKP's role in the rebuilding of tumor cells, many of PNKP's inhibitors, as well as other inhibitors of other DNA Repair Proteins, are of interest recently in order to make tumor and cancer cells more vulnerable to attack from radiation. When these malignant cells are subjected to more weakness, the radiation therapy would be rendered more potent and the cancer in the patient has a higher chance of being in remission.Some examples of the cancers that the inhibitors may fight include ovarian and colon cancers, which are more of the common cancers. While the role of PNKP to repair DNA is very beneficial to normal cells, the opposite job of the DNA Repair Inhibitor Enzymes is also helpful when trying to eradicate cancer and tumor cells.
When there is a mutation of disruption with the function of one of more of the pathways of PNKP (Non-Homologous End Joining, Single Strand Break Repair, and Base Excision Repair), there is an association with the result of severe neurological disorders within humans, which is detrimental to normal development. For example, mutations with enzyme LIG4 is associated with microcephaly, which is a condition in which a child is born with a much smaller brain circumference than what is considered natural or normal. In mice, when enzyme XRCC1 (which must be phosphorylated and then binded to the FHA domain of PNKP)is deleted contributes to seizures. A recent study found that Autosomal recessive microcephaly, infantile-onset seizures, and developmental delay (MCSZ) is caused by PNKP mutations in both domains of the enzyme. Through the many combinations of the mutations could come from either or both phosphatase and kinase domains, many symptoms of the affected individuals also vary. Through the varying symptoms of MCSZ, it is shown that PNKP can function through multiple enzymatic repair pathways (DSBR, SSBR, and BER).
Mutations in PNKP can lead to autosomal recessive neurological disorder so increased levels of PKNP can mend the effects of reactive oxygen species (ROS), which are free-radicals that contain oxygen molecules which cause DNA, protein, and lipid oxidation inside the body. The increase in cadmium and copper levels can be damaging and neurotoxic and thus leads to PNKP inhibition, which increases the probability of cancer since PNKP is the enzyme used to repair DNA strand breaks in human cells.[1] Two other findings showed that PNKP can be denatured by natural quantities of cadmium and copper. Cadmium and copper actually have harmful carcinogenic and neurological effects in a physiological sense, which seems contradictory to the function of PNKP. The accumulation of cadmium and copper will prevent the PNKP from properly functioning so when cells cannot repair the breaks in DNA strands or fix the errors, often times the result is cancer.[1]