Structural Biochemistry/Heat Shock Proteins
Heat Shock Proteins
[edit | edit source]Heat shock proteins, or HSP, are a class of proteins with related functions. Their expression increases when cells are exposed to elevated temperatures or other stress. Heat shock proteins help protect other proteins from heat stress.[1] This response to heat stress can also be seen in heat-stressed animals and microorganisms.[1] Some heat-shock proteins are called chaperone proteins because they function in unstressed cells as temporary scaffolds that help other proteins fold into their functional shapes.[1] The dramatic upregulation, or increase of cellular components, of heat shock proteins plays a key role in heat shock response and is induced primarily by heat shock factor. HSPs can be found in almost all living organisms, ranging from bacteria to humans.
Heat shock proteins are named in accordance to their molecular weight. Some widely-studied HSPs are Hsp60, Hsp70, and Hsp90, whose families consist of heat shock proteins weighing 60, 70, and 90 kilodaltons in size, respectively.
Classifications
[edit | edit source]Hsp 70
[edit | edit source]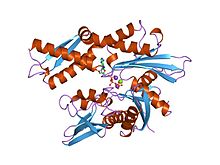
Heat shock proteins with molecular weight near 70,000, is one of the important part for protein folding to help protect heat stress. It is also a chaperone protein. It was discovered by FM Ritossa in 1960s when a “puffing pattern” –the “heat shock response”-elevated gene transcription was observed.
Structure
N-terminal domain-the one with ATPase activity( 44 kDa) – consists of two lobes, splited by a cleft and adenine nucleotide binds to it.
Substrate binding domain(18 kDa)-- made up of a groove. The groove has an attraction for neutral hydrophobic amino residues. It could also interact with peptides up to seven residues.
C-terminal domain—consists of beta-sandwich damaging and second region full of alpha helical structure for substrate binding domain. The beta-sandwish region is composted of two sheets with 4 anti-parallel beta strands. They form a “pocket” for peptide binds. It open like a lid and peptides bind and release relatively rapidly when Hsp 70 protein is ATP bound. When the lid is closed, and peptides are tightly bind to the substrate HSP 70 proteins is ADP bound.
Reaction cycle with Hsp 70
The ability of Hsp 70 to bind and release hydrophobic amino determines chaperone function, which ATP binding and hydrolysis is depend on this binding and releasing of substrate proteins. First, the N-terminal ATPase domain adjusts the attraction between Hsp 70 and substrate by altering the conformation of the C-terminal region. Once the ATP bind the Hsp 70 open the “lid” and binding and releasing of substrates happens rapidly. When ADP exist, Hsp 70 closed the “lid” and binding and releasing of substrates slow down. Due to ATP hydrolysis to ADP, the interaction is stabilized by converting Hsp 70 to a more active state. It is a repeating cycle when ADP transformed into ATP and followed with the substrate being released.
Activeness of Hsp 70
J-proteins and nucleotide exchange factors (NEFs) both affect the activity of Hsp 70. NEFs stimulate transformation of ADP to ATP, and consequently stimulate the chaperone cycle with Hsp 70. The rate of nucleotides exchange is 10-20 fold faster than the rate of hydrolysis if J-protein is nonexistent. When J-protein exist, hydrolysis is still stimulated but nucleotide release will become limited. Nef with Hsp 70 bond to ADP could excite the reaction of nucleotide exchange rate up to 5000-fold.

Hsp90
[edit | edit source]Heat shock protein 90, or Hsp90, is a molecule chaperone and is a member of the heat shock protein family. Like other heat shock proteins, Hsp90 is upregulated in response to stress and/or elevated temperatures. Hsp90 is found in bacteria and all branches of eukarya. It is one of the most abundant proteins expressed in cells. The functions of Hsp90 includes assisting in protein folding, cell signaling, and tumor repression.
The Structure and Function of the Sections of Hsp90
N-terminal Region: ~25kDa of the N-terminal region of Hsp90 was determined after proteolysis. It was found that there were various sections in the sequence that were homologous to MutL mismatch repair proteins and type II topoisomerases, proteins that alter DNA with the aid of ATP. This evidence pointed researchers in the direction that ADP/ATP had special significance to Hsp90.
The pocket in the N-terminal is capable of binding to adenine nucleotides. Furthermore, the binding site in the N-terminal is especially important for the binding to ATP/ADP, without which, Hsp90 is unable to perform its function properly.
Middle Region (Catalytic loop and binding site to client protein): A tight coil of many α-helices that are small in length connects the N-terminal region to the C-terminal region. A hydrophobic area around Trp 300 and amphiphatic characteristics of the residues between 327-340 promote interactions with client proteins. An arginine at residue 380 is required for the function of ATPase. The catalytic loop in the middle region is responsible for Hsp90's reaction with ATP/ADP.
C-terminal Region: This region is responsible for Hsp90 dimerization. A bundle of 4 helices in this region (2 from each protein) is the structure of this region taken in dimerization. This region of Hsp90 diverges most from similar proteins - evidence for this lies in a couple small deletions and a lower sequence similarity.
Although the full structure of a eukaryotic model does not currently exist, with the combination of the structures of the various regions from different model organisms, a good sense of the actual structure can be determined. The largest problem in determining the full structure is the link between the middle region and the n-terminal region because this area is very poorly conserved between organisms (its residues vary widely between various organisms).
The biomedical importance of Hsp90
The clientele of Hsp90 is restricted; unlike many other proteins which require the assistance of molecular chaperones to fold properly. Although not essential in bacteria, Hsp90 plays an important role in eukaryotes in which it maintains both cellular and organismal viability. Furthermore, scientists have discovered that Hsp90 seems to enable cancer cells to survive both hostile environments and chronic genetic instability within the host. Also, many viruses seem to require Hsp90 chaperone machinery to propagate successfully.
The Hsp90 chaperone cycle requires conformational flexibility
Like most chaperones, Hsp90 relies on conformational flexibility for its activity. Although the overall structure of bacterial Hsp90, or HtpG, and eukaryotic Hsp90 proteins are very similar, only the eukaryotic Hsp90 proteins interact with co-chaperones that helps with the stabilization of various Hsp90 conformational states as well as participate in Hsp90-dependent client protein binding, folding, and maturation. Unlike bacterial Hsp90, eukaryotic Hsp90s contain an unstructured flexible region of variable length that links the N-domain with the M-domain, which provides docking sites for client proteins and various co-chaperones. The well preserved N-, M-, and C-domains despite the large conformational rearrangements undergone by the Hsp90 dimer suggests that the conformational flexibility of Hsp90 results from the displacement of the domains with respect to each other.
By using X-ray crystallography and by analogy to other members of the GHKL superfamily of which Hsp90 is a member, scientists have determined that nucleotide binding to and hydrolysis by Hsp90 relays between two stable conformational states: an "open" apo state in which N-domains of nucleotide-free Hsp90 are not dimerized and a "closed" ATP-bound state in which N-domains are dimerized. Furthermore, the use of single particle electron microscopy has proven that bacterial Hsp90 can exist in three distinct conformational states: the "open" apo state that is nucleotide-free, the "closed" state in which both N-domains transiently dimerize in the presence of ATP, and a "compact" state in which the N-domains are no longer dimerized but instead make novel intermolecular contacts with their respective M-domain. These conformational changes are thought to impact client protein and co-chaperone binding and release.
Hsp90 chaperone cycle driven by ATP binding or hydrolysis?
Low Hsp90 ATPase activity makes it difficult to envisage ATP hydrolysis as the driving force behind the Hsp90 chaperone cycle. Researchers Southworth and Agard have brought up evidence that nucleotide binding does not drive conformational change in Hsp90. Instead, their data show that multiple Hsp90 conformations co-exist in a dynamic steady-state equilibrium in the absence of nucleotide and that this equilibrium is only moderately perturbed by nucleotide binding. In consideration of bacterial Hsp90 dynamics, the addition of either AMPPNP (a non-hydrolyzable ATP analog) or ADP can slightly skew the equilibrium to favor the closed or compact state. However, yeast Hsp90, under similar conditions, adopt only two distinct conformations, while human Hsp90 showed no obvious conformational changes in the absence of either nucleotide.
Conservation of the three-step Hsp90 chaperone cycle from bacteria to humans
Although the data collected from bacterial Hsp90 seems to fit the three-state conformational model, the data of both yeast and human Hsp90 seem to not support the model. Yeast and human Hsp90 seem to show different conformational responses to nucleotide. By using a cross-linking technique to trap rarely populated conformational states, Southworth and Agard showed that both yeast and human Hsp90 proteins conform to the three-state model. Southworth and Agard provided evidence that explained recent kinetic studies that suggest a conserved ATPase cycle among Hsp90 proteins from different species and provided support for transient N-domain dimerization in human Hsp90. An important conclusion is that the population occupancy of each conformation at equilibrium is unique for different species. This suggests that evolution have optimized the kinetics of the Hsp90 chaperone cycle to meet the distinct chaperoning requirements of each species.
References
[edit | edit source]- Trends in Biochemical Sciences Volume 34, Issue 5, May 2009, Pages 223-226
- Anatomy of gene regulation: a three dimensional structural analysis pg 249 By Panagiotis A. Tsonis
- Molecular chaperones and iron-sulfur cluster biogenesis in Saccharomyces ... By Amy J. Andrew, The University of Wisconsin - Madison
- Pearl L, Prodromou C. Structure and Mechanism of the Hsp90 Molecular Chaperone Machinery. Annual Review of Biochemistry. 75:271-294.
