Structural Biochemistry/Gene Expression
Gene Expression
[edit | edit source]Gene expression is the process where information from a gene is used in the synthesis of a functional gene product. These products are often proteins, but in non-protein coding genes such as rRNA genes or tRNA genes, the product is a functional RNA. The process of gene expression is used by all known life – eukaryotes (including multi-cellular organisms), prokaryotes (bacteria and archaea) and viruses – to generate the macromolecular machinery for life.
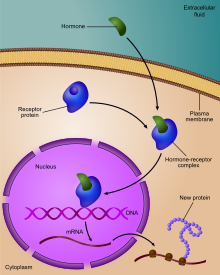

Genes are expressed when they translated into proteins after transcription into RNA. Some genes are expressed at all times, these genes are subject to constitutive expression. Many genes are not expressed at all times but only for certain situations, either in a specific cell or under a particular condition. These genes are subjected to regulated expression.
A video by Genome British Columbia demonstrating gene expression
Control of Gene Expression
[edit | edit source]Gene expression is controlled during transcription. Whether or not a gene is transcribed is mainly determined by the interplay between specific DNA sequences and the specific proteins that bind to these sequences. Genes are located in specific positions within the genome, but these sites do not have any distinguishing features. Gene regulation is controlled by regulatory sites that can be close or far away to the region of DNA that is being transcribed. To initiate gene transcription, regulatory proteins recruit RNA Polymerase to sites within the genome, this happening in both prokaryotes and eukaryotes.
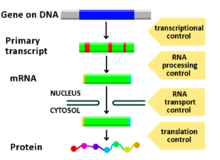
Regulatory Sites
[edit | edit source]Genes, which are located in a specific position in a genome, do not possess any distinguishing feature that allows regulatory system to recognize them, instead, other sequences in the genome contributes to its regulation. These sites, binding sites for specific DNA- binding proteins, are often close to the DNA being transcribed but its proximity it's not necessary. These proteins, when successfully bound to the sites, control whether a protein is expresses or not.
Regulatory sites were first discovered in E. coli bacteria. In the presence of sugar lactose, E. coli express a gene encoding for and enzyme that can process lactose into the use of an energy source. The nucleotide sequence for this enzyme is a nearly perfect inverted repeat.
5'-...TGTGTGGAATTGTGAGCGGATAACAATTTCACACA...3'
3'-...ACACACCTTAACACTCGCCTATTCTTAAAGTGTGT...5'
Symmetry regulatory sites like this usually correspond to the symmetry in the protein that binds the site.
Regulatory Sites in Operons
[edit | edit source]The Operon Model was proposed by Francois Jacob and Jacques Monod after a series of experiments as a common mechanism of expression of genes. The model consists of a regulator gene, the operator site which functions as a regulatory DNA sequence, and a set of structural genes. The Regulator gene encodes a repressor protein that will bind to the operator site preventing transcription. The operator site and the set of structural genes both form what is called the Operon. The Operon is a common regulatory unit in prokaryotes.
Regulation of Transcription
[edit | edit source]The main method of Gene control is regulating transcription. Transcription can be regulated by various strategies.
- Specificity Factors- alter the specificity of RNA polymerase for a promoter by changing the likelihood of binding.
- Repressors- bind to the non-coding sequences on that are close to or overlapping the promoter region of the DNA. It impedes RNA polymerase's progress, which blocks the expression of the gene.
- General transcription factors- These factors place RNA polymerase at the start of a protein-coding sequence and release the polymerase to transcribe the mRNA.
- Activator- speed up the rate of gene expression
- Enhancers- bring a specific promoter to start the complex
Expression System
[edit | edit source]An expression system is a system where the production of a gene product of choice is specifically designed. Such a system consists of a gene, normally encoded by DNA, and the molecular machinery required to transcribe the DNA into mRNA and translate the mRNA into protein using provided reagents. Although this process is utilized in every single cell, the term is more commonly used in reference to laboratory work. On the other hand, expression systems are fundamentally a natural process.
Living without 30 nm chromatin fibers
[edit | edit source]Firstly, chromatin fibers refer to strings of nucleosomes which often deal with coiling for compaction and the fitting of the genome into a cell's nucleus. According to various researchers, these strings of nucleosomes, also known as 10 nm chromatin fibers, make up the template of material that are transcriptionally active. Nowadays, with newly developed experiments such as with the chromatin conformation capture and the cryo-electron microscopy, the question of whether the in situ evidence for the 30 nm chromatin fiber emerges.
The Hierarchy of DNA Organization
[edit | edit source]A researcher named Roger Kornberg has been able to isolate the chromatin subunit which contains a histone-DNA complex, which later became known as the nucleosome. These same chromatin subunits signified the "beads-on-a-string" of the 10 nm chromatin fiber. Some experiments, such as X-ray diffraction, neutron scattering, and energy loss electron microscopy have allowed researchers to observe that 10 nm fibers illustrated the wrapping of DNA 1.75 turns around the nucleosome core. Some important observations to consider regarding the 10 nm chromatin fiber are that: a) the histones possess a positive charge which cause the core of the DNA to be positive, balancing out the charge with the DNA's negative backbone and b) Compaction occurs due to the wrapping of DNA around the histone core, which allows for the genome to reside within.
Additionally, another researcher named Aaron Klug brought up that a huge amount of the genome possess 30 nm fibers, but produced from the coiling of 10 nm fiber into a solenoid through compaction. This compaction often depends on various proteins and divalent metal ions such as magnesium.
30 nm chromatin fiber models
[edit | edit source]There are various models that allow us to observe these chromatin fibers. These models are the one-start solenoid, the two-start helix zigzag, the cross-linker, and the supranucleosome model.
1) The solenoid model = Deals with the coiling of the 10 nm fiber around an axis of symmetry with nucleosomes closely packed to one another.
2) The two-start zigzag model = Deals with the zigzagging of two nucleosomes, subsequently causing them to form and col into a helical conformation.
3) The cross-linker model = Deals with the DNA crisscrossing back and forth across the helical axis with the presence of closely packed nucleosomes located across from each other. This model is very similar to the two-start zigzag model.
4) The supranucleosome model = Deals with the clumping of nucleosomes separated by linker sequences.
--Out of these various models, only the solenoid model and the two-start zigzag model have gained wide acceptance and consistency.
Various Techniques and Experiments for Observation
[edit | edit source]One of the first experiments to observe the 30 nm fiber was done with starfish sperm and this was practiced in vivo through Electron Microscopy and Electron Spectroscopic Imaging. These studies have suggested that the 10 nm fiber dealt with folding and twisting into a width of 30 nm.
1) Cryo-EM: Another technique done was cryo-EM which allowed researchers to observe mitotic chromosomes and their lack of 30 nm fibers in a fully condensed form of the chromatin. With this particular technique, there are actually advantages such as the fact that the samples are completely hydrated and do not require chemical fixation with cross-linking agents. This makes it easier for researchers to observe the chromatin and it also infers that the 10 nm chromatin may be more energetically favorable since they are present in a fully condensed state.
2) Model Tweezers Experiment: With this, single force spectroscopy is utilized to measure the force required to convert a 30 nm chromatin fiber, which contains of 25 nucleosome repeat arrays, into an extended 10 nm fiber. This experiment provides sub-pico Newton force resolution which allows for the precise mathematical modeling of transitions between chromatin fiber configurations.
3) EM studies in vitro: These experiments had to do with the reconstitution of nucleosome arrays. This supports the two-start and the solenoid models. The downfall with this particular model though is that it is a bit unclear about how the regular spacing of the 30 nm solenoid chromatin fiber could accommodate the irregular nucleosome spacing that is present in vivo.
4) Electron Spectroscopic Imaging (ESI): This is a high-contrast technique that does not depend on heavy metal contrast agents. It is based only on the electrons that have interacted with specific elements in the specimen, producing a dark field image of high contrast, which can subsequently be utilized to observe nucleosomes.
5) Chromosome conformation capture (3C) technique: Illustrated that the subchromosomal domain within the yeast genome did not exist as a compact 30 nm fiber, but instead as an extended fiber. The linear mass density found with this technique was less than in the FISH experiment, which was inconsistent with a 30 nm chromatin fiber. This technique has only the possibility to figure out the long-range chromatin interactions from a locus of interest, causing it to have technical limitations.
6) DNA fluorescence in situ hybridization (FISH): In this technique, it has been assumed that the 30 nm fiber is the lowest order of chromatin folding in vivo. This is usually done in yeast in which you can observe a flexible 30 nm-like polymer. However, it is argued that this technique may lack sufficient resolution to conclude a highly bent and kinked 10 nm fiber in comparison to a less bent or kinked 30 nm fiber.
7) Hi-C technique: This technique was done by Lieberman-Aiden and was an adaptation of the 3C method that allows for the unbiased identification of long-range chromatin interactions, to a human lymphoblastoid cell line. In this technique, there is the digestion of cross-linked DNA with a restriction enzyme to create 5' overhangs filled with nucleotides.
Sources
[edit | edit source]1. Trends in Biochemical Sciences; Jan2011, Vol. 36 Issue 1, p1-6, 6p 2. Berg, Jeremy M., John L. Tymoczko, and Lubert Stryer. "The Control of Gene Expression." Biochemistry. Sixth ed. New York: W. H. Freeman, 2007. 892-99. Print.