Structural Biochemistry/DNA Gel Electrophoresis
Electrophoresis is the movement and subsequent separation of ions and charge macromolecules through a medium when an electric potential is applied. It is a powerful tool used in fundamental research and diagnostic settings for analysis of biomolecules. It is commonly used for purity and molecular weight determination for both DNA and protein solutions.
Theory
[edit | edit source]The idea behind DNA gel electrophoresis is to separate the fragments of a DNA sample and compare the results to a positive and negative control to determine if a DNA sample contains a particular gene. The size of the DNA fragments in the sample are determined by the specific primers used to fragment the DNA. The primers used will vary depending on the gene that is in question. For example, a DNA gel searching for the MPI (mannose phosphate isomerase) will use different primers than a Vil-Cre gel. Electrophoresis is a very powerful method to separate proteins and other macromolecules such as DNA and RNA. The speed of migration of a protein, DNA, or RNA molecule in an electric field is contingent on the strength of the electric field, the net charge of the protein, and the frictional coefficient. These turns can be summarized in the equation v = Ez/f, where v is the speed of migration, E is the electric field strength, and f is the frictional coefficient. Electrophoresis experiments are generally carried out in gels because the gel can act as a molecular sieve that amplifies the effects of separation. Small proteins move very quickly through the gel, while large proteins do not move as rapidly, staying near the point of application.
Agarose Gel
[edit | edit source]The structure of an agarose polymer is as follows:
In DNA electrophoresis, agarose gel is the common support material used. Agarose is a linear polymer of D-galactose and 3,6-anhydrogalactose that is isolated from seaweed. It forms a macroporous matrix which allows rapid diffusion of high molecular weight macromolecules without significant restriction by the gel. The concentration of agarose in the gel determines the average pore size. The size of a gel controls the mobility and resolution of components because of the sieving effect of the pores on macromolecular species.
Preparing the sample
[edit | edit source]Typically, the DNA sample is prepared by adding the specific primers to fragment the DNA and a loading dye which allows the DNA bands to show after the image is captured.
Loading the sample
[edit | edit source]After the agarose gel sets, the samples are loaded into the wells. Along with the samples in question, two control samples, a positive and negative control, are loaded as well. The positive sample of DNA will already be confirmed to contain the gene in question. The negative control should not contain any genetic material whatsoever and should therefore appear as a blank slot in the gel.
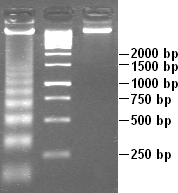
Additionally, a DNA ladder is loaded into one of the wells. The ladder contains fragments of known sizes and essentially serves as the ruler in determining the size of the sample bands. Various ladders can be used, depending on the sample being tested. Two common ladder sizes are 100 base pairs and 1000 base pairs. Each of these have very different ranges.
Running the gel
[edit | edit source]In order to run the sample DNA through the gel, a running buffer (PBS) is added to the gel box until it submerges the gel. Then, the sample is exposed to an electrical current, allowing the DNA to move through the gel and separating the various fragments in the process. After the voltage is applied, the sieving ability of the gel separates the proteins based on their size, with the smallest proteins moving most rapidly. The distance of mobility of polypeptide chains under these conditions can be measured by looking at the inverse logarithm of their mass. The mobility of the polypeptide chains is linearly proportional to this value.

After the DNA sample runs through a significant portion of the gel, an image of the various bands can be collected. By comparing the sample bands to the control bands, a researcher is able to determine if a DNA sample contains the specific gene being tested for. The strength of separation by electrophoresis can be seen through examining fractions produced during the purification scheme. The fractions that are produced initially will show many different proteins. However, as the separation runs for longer time, the number of bands will shrink, and the intensity of one of the bands should increase. The band with the highest intensity should reveal the protein of interest.
