Structural Biochemistry/Chemical Bonding/ Disulfide bonds
Introduction
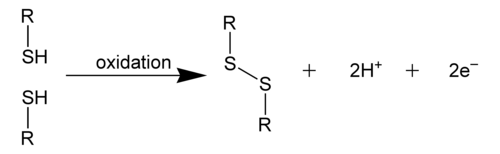
[edit | edit source]A disulfide bond, also called an S-S bond, or disulfide bridge, is a covalent bond derived from two thiol groups. In biochemistry, the terminology R-S-S-R connectivity is commonly used to describe the overall linkages. The most common way of creating this bond is by the oxidation of sulfhydryl groups. (2 RSH → RS-SR + 2 H+ + 2 e-) This process of oxidation can produce stable protein dimers, polymers, or complexes, in which the sulfide bonds can help in protein folding. The process mostly occurs with the thiol groups in cysteine. [1]
Disulfide bonds can occur in two ways: intramolecularly and intermolecularly. Intermolecular disulfide bonds occur between polypeptide chains while intramolecular disulfide bonds occur within a polypeptide chain and are usually responsible for stabilizing tertiary structures of proteins. On the other hand, intermolecular disulfide bonds are attributed to stabilizing quaternary protein structures. [1]
Disulfide Bonds in Proteins
[edit | edit source]
Disulfide bonds in protein membranes are found in both bacteria and eukaryotes. Extracellular proteins often have several disulfide bonds, whereas intracellular proteins usually lack them. In proteins, these bonds form between the thiol groups of two cysteine amino acids. Cross-linkage between multiple linear polypeptide chains is not uncommon in proteins. Most of the cross-linkages are from disulfide bonds formed by the oxidation of two cysteine amino acids. The result is a disulfide bond called cystine connecting the polypeptide chains. The cysteine amino acid group is the only amino acid capable of forming disulfide bonds, and thus can only do so with other cysteine groups. These bonds are responsible for the stabilizing the globular structure and are the strongest type of bond that a protein can possess and are one of the major forces responsible for holding proteins in their respective conformations, and therefore have an important role in protein folding and stability.
The typical bond dissociation energy of a disulfide bond ranks at 60 kcal/mole and has a bond length of 2.05 Å. Fairly low energy is required to produce rotations about the S-S bonds, thus these rotations are common. At dihedral angles near 90°, the bonds tend to be more stable. However, the bonds become significantly better oxidants at angles approaching 0° and 180°. Disulfide bonds have been identified in the protein folding in E. Coli. They are used in many processes, including DNA replication.
Disulfide Bonds in cyclic peptides
[edit | edit source]Most cyclic peptide bonds are formed between disulfide bonds. As a result, the denaturation of cyclic peptides can often be attributed to the stability of disulfide bonds. In the study with the peptide 1 (cyclo(1,4)-Cys-Gly-Phe-Cys-Gly-OH), where it was conducted in buffer solutions between pH 1-11 at 70 degrees C. It was found that the most stability came from pH ~ 3 and a Vshape between pH ~1-~5. As the pH goes from neutral to basic, degradation was found between Gly2-Phe3, which is due to the breaking of disulfide bonds.

Making disulfides
[edit | edit source]Multiple ways to make disulfides
In the journal article “Multiple ways to make disulfides” by Neil J. Bulleid and Lars Ellgaard, they discuss how disulfides can be formed in the endoplasmic reticulum (ER), and the different enzymes that catalyze the pathways of formation. Disulfides increase the stability of the protein and also “regulate redox-dependent functions,” and over the years, our ideas of how disulfides form in proteins have drastically changed. [2]
Disulfides are created in the presence of enzymes in the protein disulfide isomerase (PDI) family. They act as a oxidizing agent, oxidizing the thiol group on a protein. If the protein's amino acid residues, specifically cysteines, are close to one another they will form a disulfide bond even if it is not properly folded. If a disulfide bond forms when the protein is not properly folded, they call this a non-native disulfide. This could be a misfolded protein, or it could be one of the intermediates before the protein folds into its native state. PDIs help non-native disulfides become native disulfides by acting as a catalyst to the isomerization process (they have to help brake the non-native disulfide bond so that the protein can finish folding properly before they can form the native disulfide bond) (Figure 1). [2]
Figure 1: http://imgur.com/sEvem
Ero1
Bulleid and Ellgaard studied an enzyme in yeast to understand how disulfides were formed de novo (Latin for “in the beginning”). The enzyme they studied was ER oxidoreduct in (Ero1p). Experiments showed that “Ero1p and the mammalian homologues ERO1(alpha) and ERO1(beta) are able to catalyze oxidation by coupling de novo disulfide formation to the reduction of oxygen to hydrogen peroxide (H2O2).” Ero1p was shown to oxidize PDI, which allowed PDI to exchange disulfides on the protein. Using knockout experiments they were able to show that while in yeast, knockout of Erop1 interrupted disulfide formation, in higher eukaryotes (for example, mice and humans) knockout of ERO1(beta) only caused misfolding in proinsulin and the double knockout of ERO1(alpha) and ERO1(beta) did not do much worse than just the knockout of ERO1(beta). In fact, after some time, “double knockout cells re-established normal ER redox conditions after a strong reductive challenge, albeit at a slower rate than in wild-type cells.” This tells us that ERO1 is not as necessary in higher eukaryotes as it is in yeast and implies that there are other pathways to forming disulfide bonds. [2]
PRDX4
Because hydrogen peroxide is produced when disulfides are formed via ERO1 catalysis, and H2O2 can cause damage biomolecules, Bulleid and Ellgaard believed there had to be other proteins in order to remove the H2O¬2¬. This is where peroxiredoxin (PRDX4) comes in. PRDX4 is a group of enzymes located in the ER that both removes H2O2 and also forms disulfides. In this process the peroxidatic cysteine in PRDX4 takes an oxygen from H2O2 to make water and a –SOH group, this then reacts with the adjacent –SH group to form a disulfide bond (Figure 2). This can now be exchanged with the –SH groups on some PDI proteins so it can then exchange with substrate proteins (Figure 1). [2]
Figure 2: http://imgur.com/oyzmc
References
[edit | edit source]1. He, HT. "Synthesis and chemical stability of a disulfide bond in a model cyclic pentapeptide: cyclo(1,4)-Cys-Gly-Phe-Cys-Gly-OH." The University of Kansas, Lawrence, Kansas 66047, USA.
2. Neil J. Bulleid, Lars Ellgaard, Multiple ways to make disulfides, Trends in Biochemical Sciences, Volume 36, Issue 9, September 2011, Pages 485-492, ISSN 0968-0004, 10.1016/j.tibs.2011.05.004.(http://www.sciencedirect.com/science/article/pii/S096800041100082X)