Structural Biochemistry/Biophysics- Single Molecule techniques
Single-molecule techniques have recently become popular in the biophysical department in helping to discover or clarify and better understand some important biochemical properties such as protein-DNA interactions, protein folding, and the functions and capabilities of membrane proteins. Many of the single-molecule techniques were first revealed in the physics and biophysics department, and later were found to be of great assistance to research biological and biochemical molecules. In 1976, the technique called single ion-channel recordings was first discovered and that later became the gateway to recent techniques such as atomic force microscopy (AFM), optical and magnetic tweezers and single-molecule fluorescence spectroscopy. Many of these recent techniques have helped in fields such as protein folding, transcription, replication, translation, molecular motors, membrane proteins, and viral biology. These single-molecule methods have been able to provide information on problems that could not be solved before while also giving a more detailed view into subjects that have already been researched. Not only that, but single-molecule techniques have also been able to steer the biochemists away from the usual averaging of ensembles that results in the use of moles to a more detailed and unitary concept that involves the single molecules and particles.
Static vs. Dynamic Heterogeneity
[edit | edit source]A useful aspect of single-molecule techniques is that they give the distribution of values for a property instead of an average of the property which is averaged over a large molecule ensemble. This kind of feature allows for more complete and specific data that can speak more about the certain biomolecule that is being assessed than an average which gives a rather unspecific and broad view. The specificity of single-molecule methods also provides for information on the molecular heterogeneity, which is a fundamental aspect of complex biomolecules and their functions. Molecular heterogeneity can be classified as either static or dynamic. Static heterogeneity is when a collection of molecules that have several subpopulations are extremely stable and do not interconvert over the period of time that is being observed. An example of this is inactive molecules. When an ensemble method is being used, one is trying to determine the fraction of active molecules compared to the whole population. However, when using a single-molecule technique, the inactive molecule can be ignored or disregarded because they do not give an experimental signal. Therefore, static heterogeneity allows for the sole research of the species of interest (active molecule) because the inactive ones can be ignored and removed. On the other hand, dynamic heterogeneity is when a sample with subpopulations of molecules interconvert over the time period observed. An example of dynamic heterogeneity is an enzyme that interconverts between two catalytically active states, with each conformation yielding a different affinity for a substrate. When the interconversion of the enzyme is fast compared to the temporal resolution of the single-molecule technique that is being used, the value that comes out will show a weighted time-average mean of the affinity for each of the two states of the enzyme. On the other hand, if the interconversion is slower than the temporal resolution of the single-molecule method, then the observation of the interconversion between the two states can be directly seen. In addition, dynamic heterogeneity can also be observed in enzymes that operate on substrates in multiple steps, with each step having a different rate. With single-molecule analysis, the observation of each single step in the process of the enzyme can be carefully monitored as real time "movies" that show the kinetics of each step and the intermediate structures of the enzyme and substrate.
Kinetics of Single-Molecule Techniques
[edit | edit source]The type of information that is extracted from single-molecule techniques are also something that can be compared to the ensemble assays that are used frequently. One example is the kinetic information that is provided by the single-molecule methods. Kinetic observation of a reaction by single-molecule methods provide something called dwell times. These dwell times are of a single molecule at each of its certain states along the reaction pathway. Think of two structural states of a molecule called A and B that is being observed by a single-molecule observable. The single-molecule method with enough temporal resolution to look at the two states would be able to determine the dwell time for states A and B. Afterward, a frequency histogram can be plotted to provide information on how long each dwell time lasts. Dwell time distributions can be used to study enzyme-substrate interactions. This example can also be used to reveal the significant aspect of single-molecule technique in its method of identifying the structural states of a biological macromolecule.
Single-Molecule Methods
[edit | edit source]There is a wide variety of single-molecule techniques that are present right now. The commonly used methods are separated into two different attributes: force and fluorescence. These two groups were separated based upon the different time resolutions, span of observations, and different spatial resolutions. In the force-based detection methods, there are specific techniques called atomic force microscopy, optical tweezers, tethered partical motion, and magnetic tweezers. On the other hand, the fluorescence imaging includes confocal microscopy and total-internal-reflection fluorescence (TIRF).
Force-based Dectection
[edit | edit source]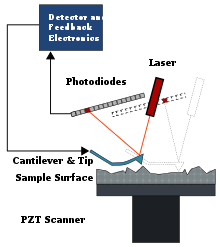
Atomic Force Microscopy (AFM)
[edit | edit source]Atomic force microscopy was first used for topographical imaging of molecules on a flat atomic surface. This form of imaging is done by scanning an extremely sharp tip along the sample surface and then measuring the deflection from the tip by using a laser and a quadrant photodetector. Dry and wet solutions can both be observed under this imaging, although the wet solution's temporal resolution is far worse than the dry solution's. The setup of AFM usually has the a small lever (AFM cantilever) that has a sharp nanometer scale tip which is attached to one of the ends of a biomolecule. By using piezoelectric positioning, the surface which the biomolecule is adsorbed can then be scanned in the x,y,z coordinate system (3D spatial) to an angstrom scale resolution. Then, the position of the tip could be measured by deflecting a laser beam off the surface of the tip and onto the photodetector, which will be position sensitive. The lever will act as a linear spring and when the biomolecule is moved relative to the tip, the lever will flex and apply a force on the molecule. Because the lever is very rigid, the force applied can be great and is therefore very useful in measuring the structural properties of folded proteins and chemical bonds.
Optical Tweezers
[edit | edit source]
For the method of optical tweezers, the setup involves a high-power infrared laser that is focused tightly because of a high-end microscope lens. The focused infrared laser beam would trap a micron-sized bead at its focal spot, which allows for the control of the position of the bead. Then, the trap acts like a spring, which would exert a higher force on the bead when it moves farther away from the laser axis. And by measuring the position of the bead in relation to the focal spot, the trap's applied force can be determined (this can be done with the quadrant photodetector, like that in AFM). There are two main modes of optical tweezers: constant force and constant position. When it is in the constant force mode, a constant trap force can be maintained by a feedback loop that will displace the optical trap or the sample coverslip surface, which will keep the position of the bead constant within the zone that it is trapped in. In the constant position mode, the trap position's center is held and the bead will experience a progressively growing force as it is being pulled out of the trap.
Magnetic Tweezers
[edit | edit source]The setup for magnetic tweezers has the small bead that is used in optical tweezers replaced by a small magnetic bead. The small magnetic bead will be controlled by a pair of magnets that is close to the sample. This setup allows for the tethered biomolecule to have a force applied on it and a rotation that will be easily imposed. Magnetic tweezers function in the constant force mode because the force is an important control parameter for the interaction of interest. In addition, compared to the optical tweezers, the magnetic tweezers cannot obtain the exquisite spatial sensitivity. However, it is easier to use and can be compatible with long term analysis of single biomolecules. Magnetic tweezers is usually used for studying the structural properties of DNA and protein-DNA transactions.
Tethered Particle Motion
[edit | edit source]This method is used for studying the interation of proteins and polymers. The main idea is one bead is bound to the surface while the other bead is attached to the end of the polymer. These are then put into an aqueous solution and it can be seen that the bead has restricted movement in which it moves in a Brownian motion. The position of this bead is then recorded by optical microscopes; the recordings can tell us information about the DNA during transcription. Different bead types have also improved ways of analyzing polymers. Firstly, metal beads are sometimes used due to their high intensity of gold light. Secondly, polystyrene beads are also used in conjunction with the optical tweezers because these beads are less intense than the metal ones.
Fluorescence-based Methods
[edit | edit source]Fluorescent biomolecules contain several properties that can be used by fluorescent microscopy and spectroscopy to look at their location, structure and dynamics. There are two main methods for single-molecule fluorescence: confocal microscopy for point detection and wide-field imaging for area detection. Confocal microscopy for point detection tries to collect fluorescence that is emitted by a diffraction-limited volume of ~1 femtoliter. On the other hand, for wide-field imaging, there is total-internal-reflection fluorescence (TIRF) microscopy. TIRF is capable of observing hundreds of molecules that are immobile on the surface for extended periods of time through using evanescent-wave excitation within a thin layer just above the surface and using a ultrasensitive camera above surface for imaging.
FIONA or fluorescence imaging with one nanometer accuracy, has enabled motion tracking with precision of 1 nanometer. This can be done with any single fluorophore or light-scattered particle (example: small gold particle) that is attached to the molecule of interest. With that set up, one can identify the presence of a molecule and track its movement on molecular tracks in vitro or its diffusion in vivo. This remarkable tracking ability has been able to reveal how molecular motors like kinesin and myosin move on their tracks. The combination of this precision in tracking and the founding of complicated methods of switching on and off the fluorescence of some probes allows for techniques such as PALM (photoactivated localization microscopy) and STORM (stochastic optical reconstruction microscopy). PALM and STORM have been characterized as "super-resolution microscopies" because of their ability to achieve spatial resolutions that are better than 50 nanometers.
With two probes attached to a molecule, there are several new capabilities that come forth. FRET (fluorescence resonance energy transfer) is distance dependent and can be exploited to measure nanometer distances and the change in distances within single molecules. For FRET, the first probe would act as the FRET donor while the second probe would be the FRET acceptor. The first probe would be fluorescent, and the second probe would quench the donor in a distance-dependent manner (it can also be fluorescent). Because of this setup, the movements that change the donor-acceptor's distance of separation would also change their fluorescence. These changes are used to observe the kinetics of conformational changes or molecular association/disassociation. However, single-molecule FRET is also affected by relative orientation and rotational freedom of the fluorophores, which can be mistaken for conformational changes. This difficulty can be examined by using MFD (multi-parameter fluorescence), which is a method that can tell the fluorescence properties of single molecules such as fluorescence intensity, anisotropy, and lifetime at different wavelengths. Another method that can help the complication is ALEX (alternating laser excitation), which uses two lasers to measure FRET and relative probe stoichiometries.
Combination of Force and Fluorescence Approaches
[edit | edit source]The combination of force and fluorescence techniques is being pursued right now by single-molecule groups because of it will allow for simultaneous manipulation and visualization of single molecules as they interact and react. This combination works because the force and fluorescence techniques are highly complementary due to the fact that force-based methods can achieve timescales of 50-100ms while fluorescence-based can be many times faster and are not restricted by an applied force. In addition, whereas force-based methods provide information more on the global structural and mechanical rearrangements in biomolecules, fluorescence-based methods reveals the local conformational changes. This complementary ability of the two types of single-molecule method can lead to great discoveries in the future.
References
[edit | edit source]Kapanidis, A.N. and Strick, T., 2009, Biology, one molecule at a time, Trends in Biochemical Sciences, v. 34, p. 234-243.
