Methods and Concepts in the Life Sciences/Cloning Methods
Cloning Methods
[edit | edit source]Molecular cloning is a set of experimental methods that are used to assemble recombinant DNA molecules and to direct their replication within host organisms. The use of the word cloning refers to the fact that the method involves the replication of one molecule to produce a population of cells with identical DNA molecules. Molecular cloning generally uses DNA sequences from two different organisms: the species that is the source of the DNA to be cloned, and the species that will serve as the living host for replication of the recombinant DNA.
Here the whole process is outlined using the example of cloning by restriction and ligation, which is the traditional method. Following this, special strategies and alternative methods are described.
Cloning by restriction and ligation
[edit | edit source]Prior to the 1970s, our understanding of genetics and molecular biology was severely hampered by an inability to isolate and study individual genes from complex organisms. This changed dramatically when restriction enzymes and DNA ligase were found to be valuable tools for molecular biology. The first recombinant DNA molecules were generated and studied in 1972. Cloning by restriction and ligation soon became a standard procedure in molecular biology. The whole process typically involves the following seven steps.
Choice of host organism and cloning vector
[edit | edit source]Although a very large number of host organisms and molecular cloning vectors are in use, the great majority of molecular cloning experiments begin with a laboratory strain of E. coli and a plasmid cloning vector. E. coli and plasmid vectors are in common use because they are technically sophisticated, versatile, widely available, and offer rapid growth of recombinant organisms with minimal equipment. If the DNA to be cloned is exceptionally large (hundreds of thousands to millions of base pairs), then a bacterial artificial chromosome or yeast artificial chromosome vector is often chosen.
Specialized applications may call for specialized host-vector systems. For example, if the experimentalists wish to harvest a particular protein from the recombinant organism, then an expression vector is chosen that contains appropriate signals for transcription and translation in the desired host organism. Alternatively, if replication of the DNA in different species is desired (for example transfer of DNA from bacteria to plants), then a multiple host range vector (also termed shuttle vector) may be selected. In practice, however, specialized molecular cloning experiments usually begin with cloning into a bacterial plasmid, followed by subcloning into a specialized vector.
Preparation of vector DNA
[edit | edit source]The cloning vector is treated with a restriction endonuclease to cleave the DNA at the site where foreign DNA will be inserted. The restriction enzyme is chosen to generate a configuration at the cleavage site that is compatible with that at the ends of the foreign DNA. Typically, this is done by cleaving the vector DNA and foreign DNA with the same restriction enzyme. It is important that this restriction digest works well, otherwise there will be many colonies which contain the original vector without an insert. Religation of the vector can lead to this problem as well. When the ends of the linear vector are compatible, they can be linked in the ligation step and cause false-positive colonies. To prevent this from happening, the vector may be treated with a phosphatase. DNA ligase requires phosphate groups to create phosphodiester bonds, therefore it cannot connect dephosphorylated vector ends. In this case it is necessary that the insert DNA carries 5′ phosphate groups, which might not be the case if it was obtained from a PCR.
Preparation of insert DNA
[edit | edit source]For cloning of genomic DNA, the DNA to be cloned is extracted from the organism of interest. Virtually any tissue source can be used, as long as the DNA is not extensively degraded. The DNA is then purified to remove contaminating proteins, RNA and smaller molecules. PCR methods are often used for amplification of specific DNA or RNA (RT-PCR) sequences prior to molecular cloning.
DNA for cloning experiments may also be obtained from RNA using reverse transcriptase (cDNA cloning), or in the form of synthetic DNA (artificial gene synthesis). cDNA cloning is usually used to obtain clones representative of the mRNA population of the cells of interest, while synthetic DNA is used to obtain any precise sequence defined by the designer.
The purified DNA is then treated with a restriction enzyme to generate fragments with ends capable of being linked to those of the vector. If necessary, the desired restriction sites may be added to the insert in the PCR step.
Ligation
[edit | edit source]In this step, vector and insert are mixed together at appropriate concentrations and exposed to DNA ligase, which covalently links the ends together. The resulting DNA mixture containing randomly joined ends is then ready for introduction into the host organism.
DNA ligase only recognizes and acts on the ends of linear DNA molecules, usually resulting a complex mixture of DNA molecules with randomly joined ends. The desired products (vector DNA covalently linked to foreign DNA) will be present, but other sequences (e.g. foreign DNA linked to itself, vector DNA linked to itself and higher-order combinations of vector and foreign DNA) are also usually present. This complex mixture is sorted out in subsequent steps of the cloning process, after the DNA mixture is introduced into cells.
Transformation
[edit | edit source]Following the in vitro manipulation of the DNA, the product is typically introduced into E. coli in a process called transformation. For transformation to happen, bacteria must be in a state of competence. Some bacteria show natural competence, in others this state must be induced by laboratory procedures. Typically, the cells are incubated in a solution containing divalent cations (often calcium chloride) under cold conditions, before being exposed to a heat shock.
The surface of bacteria such as E. coli is negatively charged due to phospholipids and lipopolysaccharides on its cell surface, and the DNA is negatively charged as well. One function of the divalent cations therefore would be to shield the charges by coordinating the phosphate groups and other negative charges, thereby allowing a DNA molecule to adhere to the cell surface. It is suggested that exposing the cells to divalent cations in cold condition may also change or weaken the cell surface structure of the cells, making it more permeable to DNA. The heat-pulse is thought to create a thermal imbalance on either side of the cell membrane, which forces the DNA to enter the cells through either cell pores or the damaged cell wall.
Electroporation is another transformation method. In this method the cells are briefly shocked with an electric field of 10-20 kV/cm which is thought to create holes in the cell membrane through which the plasmid DNA may enter. After the electric shock the holes are rapidly closed by the cell's membrane-repair mechanisms.
Selection
[edit | edit source]The introduction of recombinant DNA into the chosen host organism is usually a low efficiency process; that is, only a small fraction of the cells will actually take up DNA. This makes a selection step necessary, in which only cells that have taken up the DNA with the selectable marker will survive.
When bacterial cells are used as host organisms, the selectable marker is usually a gene that confers resistance to an antibiotic, typically ampicillin. The step is carried out by plating the solution containing the transformants on agar plates with the antibiotic and incubating them at 37 °C overnight. Cells harboring the plasmid will survive and form colonies, while those that have failed to take up plasmid sequences will die.
Screening for correct clones
[edit | edit source]Modern bacterial cloning vectors (e.g. pUC19 and later derivatives including the pGEM vectors) use the blue-white screening system to distinguish colonies of transgenic cells from those that contain the parental vector (i.e. vector DNA with no recombinant sequence inserted). Therefore, it is easy to identify transgenic bacterial clones, while ignoring those that do not contain recombinant DNA.
If the vector contains no such system, a colony PCR can be performed to find out which colonies carry plasmids with inserts. A restriction digest and subsequent gel electrophoresis is another option. This can only be carried out after colonies have been picked, cultured and the plasmids have been isolated.
Even if these methods show that DNA was integrated into the vector, one cannot be sure that it is the desired sequence. For this reason, DNA sequencing needs to be performed as the final validation step.
Golden Gate cloning
[edit | edit source]Golden Gate Cloning is a method that allows a researcher to simultaneously and directionally assemble multiple DNA fragments into a single piece using Type IIs restriction enzymes and DNA ligase. This assembly is performed in vitro. Most commonly used Type IIs enzymes include BsaI, BsmBI and BbsI.
Unlike standard Type II restriction enzymes like EcoRI and BamHI, these enzymes cut DNA outside of their recognition sites and therefore can create non-palindromic overhangs. Since 256 potential overhang sequences are possible, multiple fragments of DNA can be assembled by using combinations of overhang sequences.
Gibson assembly
[edit | edit source]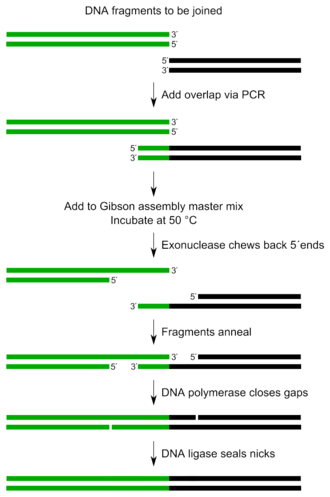
Gibson assembly is a DNA assembly method which allows for the joining of multiple DNA fragments in a single, isothermal reaction. It was invented in 2009 by Daniel Gibson while he was at the J. Craig Venter Institute (JCVI).
The entire Gibson assembly reaction requires a small number of components with very few manipulations.
The method can simultaneously combine numerous (>10) DNA fragments based on sequence identity. It requires that the DNA fragments contain ~20-40 base pair overlap with adjacent DNA fragments. These DNA fragments are mixed with a cocktail of three enzymes, along with other buffer components.
The three required enzyme activities are: exonuclease, DNA polymerase, and DNA ligase.
- The exonuclease chews back DNA from the 5' end. The resulting single-stranded regions on adjacent DNA fragments can anneal.
- The DNA polymerase incorporates nucleotides to fill in any gaps.
- The DNA ligase covalently joins the DNA of adjacent segments, thereby removing any nicks in the DNA.
The entire mixture is incubated at 50°C for up to one hour. The resulting product is different DNA fragments joined into one.
This DNA assembly method has many advantages compared to conventional restriction enzyme/ligation cloning of recombinant DNA.
- No restriction digest of the DNA fragments after PCR is necessary. The backbone vector can be digested, or synthesized by PCR.
- It is far simpler than conventional cloning schemes, as it requires fewer steps and fewer reagents. The process also takes less time.
- No restriction site scar remains between two DNA fragments (a.k.a, "scarless").
- Multiple DNA fragments can be combined simultaneously in a single-tube reaction.
TA and TOPO cloning
[edit | edit source]TA cloning
[edit | edit source]The TA cloning method takes advantage of the terminal transferase activity of Taq polymerase. This enzyme adds a single adenosine (A) to the 3'-end of PCR products. It is best if the PCR primers have guanines at the 5' end as this maximizes probability of Taq DNA polymerase adding the terminal adenosine overhang. Polymerases containing extensive 3´ to 5´ exonuclease activity should not be used as they do not leave the 3´ adenine-overhangs. The target vector is linearized and cut with a blunt-end restriction enzyme. This vector is then tailed with dideoxythymidine triphosphate (ddTTP) using terminal transferase. It is important to use ddTTP to ensure the addition of only one T residue. This tailing leaves the vector with a single 3'-overhanging thymine residue on each blunt end. This makes it possible to clone the PCR product directly into the linearized vector.
TOPO TA cloning
[edit | edit source]TOPO TA cloning combines the advantages of TA cloning with the ligation activity of DNA topoisomerase I. The biological role of topoisomerase is to cleave and rejoin supercoiled DNA ends to facilitate replication. Vaccinia virus topoisomerase I specifically recognizes the DNA sequence 5´-(C/T)CCTT-3'. During replication, the enzyme digests DNA specifically at this sequence, unwinds the DNA and re-ligates it again at the 3' phosphate group of the thymidine base.
TOPO vectors are designed in such a way that they carry this specific sequence 5´-(C/T)CCTT-3' at the two linear ends. The linear vector DNA already has the topoisomerase enzyme covalently attached to both of its strands' free 3' ends. This is then mixed with PCR products. When the free 5' ends of the PCR product strands attack the topoisomerase/3' end of each vector strand, the strands are covalently linked by the already bound topoisomerase. This reaction proceeds efficiently when the solution is incubated at room temperature with required salt.
TOPO blunt-end cloning
[edit | edit source]TOPO blunt-end cloning makes use of linearized vectors with topoisomerases attached to the ends as well but does not rely on A/T overhangs. This means that polymerases with proofreading activity can be used for PCR.
Gateway cloning
[edit | edit source]The Gateway cloning System, invented and commercialized by Invitrogen since the late 1990s, enables researchers to efficiently transfer DNA-fragments between plasmids using a proprietary set of recombination sequences, the "Gateway att" sites, and two proprietary enzyme mixes, called "LR Clonase", and "BP Clonase". The method is based on the site-specific recombination system used by phage l to integrate its DNA in the E. coli chromosome.
Gateway Cloning Technique allows transfer of DNA fragments between different cloning vectors while maintaining the reading frame. Using Gateway, one can clone or subclone DNA segments for functional analysis. The system requires the initial insertion of a DNA fragment into a plasmid with two flanking recombination sequences called attL1 and attL2, to develop a “Gateway Entry clone”.
Large archives of Gateway Entry clones, containing the vast majority of human, mouse and rat ORFs have been cloned from human cDNA libraries or chemically synthesized to support the research community using NIH (National Institutes of Health) funding. The availability of these gene cassettes in a standard Gateway cloning plasmids helps researchers quickly transfer these cassettes into plasmids that facilitate the analysis of gene function. Gateway cloning takes more time for initial set-up, and is more expensive than traditional restriction enzyme and ligase-based cloning methods, but it saves time, and offers simpler and highly efficient cloning for down-stream applications.
The technology has been widely adopted by the life science research community especially for applications that require the transfer of thousands of DNA fragments into one type of plasmid (e.g., one containing a CMV promoter for protein expression in mammalian cells), or for the transfer of one DNA fragment into many different types of plasmids (e.g., for bacterial, insect and mammalian protein expression).
The first step in Gateway cloning is the preparation of a Gateway Entry clone. Entry clones are often made in two steps:
1) Gateway attB1, and attB2 sequences are added to the 5’ and 3’ end of a gene fragment, respectively, using gene specific PCR primers and PCR-amplification;
2) The PCR amplification products are then mixed with special plasmids called Gateway “Donor vectors” and the proprietary “BP Clonase” enzyme mix. The enzyme mix catalyzes the recombination and insertion of the attB-sequence-containing PCR product into the attP recombination sites in the Gateway Donor vector. Once the cassette is part of the target plasmid, it is called an "Entry clone" in the Gateway nomenclature, and recombination sequences are referred to as the Gateway “attL” type.
The gene cassette in the Gateway Entry clone can then be simply and efficiently transferred into any Gateway Destination vector (Invitrogen nomenclature for any Gateway plasmid that contains Gateway “attR” recombination sequences and elements such as promoters and epitope tags, but not ORFs) using the proprietary enzyme mix “LR Clonase”. Thousands of Gateway Destination plasmids have been made and are freely shared amongst researchers across the world. Gateway Destination vectors are similar to classical expression vectors containing multiple cloning sites. Gateway Destination vectors are commercially available from Invitrogen, EMD (Novagen) and Covalys.
Since Gateway cloning uses patented recombination sequences, and proprietary enzyme mixes available only from Invitrogen, the technology does not allow researchers to switch vendors and contributes to the lock-in effect of all such patented procedures.
To summarize the different steps involved in Gateway cloning:
- Gateway BP reaction: PCR-product with flanking attB sites (e.g., amplified from cDNA library) + Donor vector containing attP sites + BP clonase => Gateway Entry clone, containing attL sites, flanking gene of interest
- this step can be replaced by other cloning methods
- Gateway LR reaction: Entry clone containing attL sites + Destination vector containing attR sites, and promoters and tags + LR clonase => Expression clone containing attB sites, flanking gene of interest, ready for gene expression.
Ligation independent cloning (LIC)
[edit | edit source]Ligation is often the most problematic step in traditional cloning methods. When using restriction enzymes, this step is necessary because the short sticky ends generated by the enzymes cannot hold vector and insert together permanently. Ligation independent cloning eliminates the need for ligation by creating longer overhangs. When the complementary overhangs of vector and insert anneal, the hydrogen bonds are strong enough to hold them together and the construct can be transformed directly. The nicks are thereupon repaired by host cell enzymes.
To create the overhangs, T4 DNA polymerase is used. This enzyme has a 5' to 3' polymerase activity and a 3' to 5' exonuclease activity. The exonuclease can remove nucleotides from 3' ends, but as longs as all nucleotides are present in the solution they are immediately replaced by the polymerase. In LIC, however, only dGTP is added to the mixture, which means that the exonuclease progresses until it reaches the first G in the sequence. This means that the overlap region must be designed in a way that the resulting overhangs are long enough.
To perform the reaction, the vector is linearized by restriction digest or PCR, if the use of restriction enzymes is to be avoided entirely. The insert is amplified by PCR and sequences which are homologous to the ends of the vector are added to the 5' ends.
Sequence and ligation independent cloning (SLIC)
[edit | edit source]In this variation of ligation independent cloning, all dNTPs are excluded from the reaction at first. After an appropriate time, in which the 3' to 5' exonuclease activity of T4 DNA polymerase creates 5' overhangs, the reaction is stopped by adding dCTP. As soon as the enzyme reaches a C in the 3' to 5' strand, the exonuclease activity can be balanced by the polymerase. However, because only one nucleotide was added, the overhangs are retained and can be used to connect vector and insert.
In-Fusion Cloning
[edit | edit source]In-Fusion® Cloning is a proprietary assembly methodology developed by Clontech which is similar to SLIC and Gibson assembly. In contrast to these methods, the recommended overlap length is only 15 bp and it uses the Vaccinia virus DNA polymerase to create overhangs.
Seamless Ligation Cloning Extract (SLiCE)
[edit | edit source]SLiCE is similar to cloning methods such as SLIC or Gibson assembly in that it requires the same types of DNA starting molecules and results in the same product. The major difference is that it uses bacterial cell extract, which is easy to generate and potentially very cost-effective.
Phosphorothioate-based ligase-independent gene cloning (PLICing)
[edit | edit source]PLICing is an enzyme-free and sequence-independent cloning method which is based on a chemical cleavage reaction of phosphorothioate bonds.
In the first step, both vector and insert are amplified by PCR using primers with complementary phosphorothioated nucleotides at the 5' end. These nucleotides form phosphorothiodiester bonds instead of phosphodiester bonds. After PCR, these bonds are cleaved in an iodine/ethanol solution, which results in single-stranded overhangs. The ends of vector and insert hybridize and the construct can be transformed.
