Medical Physiology/Cellular Physiology/DNA and Reproduction
Introduction
[edit | edit source]All cells contain the complete genetic code of the entire organism, but there are several different kinds of cells. The differences depend on which genes get expressed by the cell. During the development process different cells are 'guided' into different functions and arranged into different patterns. In the mature organism, some of the cell types such as the hemopoietic cells, most connective tissue and epithelial cells continue dividing on a regular basis, others such as the liver cells are capable of division, but rarely do, and others such as the muscle cells and neurons do not divide, or if they do they divide rarely. Cell division is regulated by several regulators called proto-oncogenes which, because of they importance in cancer studies, are the subject of intensive investigation. For the same reason natural cell death - apoptosis- is also the subject of intensive study.
Cell division in eukaryotes has important differences to cell division in prokaryotes, and these differences are taken advantage of by antibiotics. For this reason, and the fact that much of the research on DNA replication has be carried out on E Coli, we will be looking briefly at cell division in prokaryocytes.
Forming a DNA strand
[edit | edit source]As we saw in the biochemistry section (Nucleic Acids) DNA consists of two anti-parrellel strands of Nucleic acids arranged as a Double Helix(fig.3). The following is a recap of the formation and replication of DNA. See the Biochemistry section for the full chemical formulars.
The spine of the strand consists of Deoxyribose and phosphate molecules linked by a phosphodiester bonds (Fig.2).
The strands are linked together by hydrogen bonds (Fig.3). The links are always between thymine and adenine nucleic acid molecules and guanine and cytosine molecules.
In eukaryotic cells the enzyme DNA polymerase catalyses the reaction. First the nucleic acid is matched to the parental chain, and then a phosphodiester bond is formed. Note that there are three hydrogen bonds between guanine and cytosine, and only two bonds between thymine and adenine.
Transcribing DNA
[edit | edit source]During cell division a new strand of DNA is transcribed from the parental strand. The enzyme δ-DNA polymerase catalyses the reaction by attaching a matching nucleic acid to the exposed 3' hydroxyl group. The Enzyme does not disengage and reattach with each nucleic acid: it is said to be highly processive.
Chromosomes and Genes
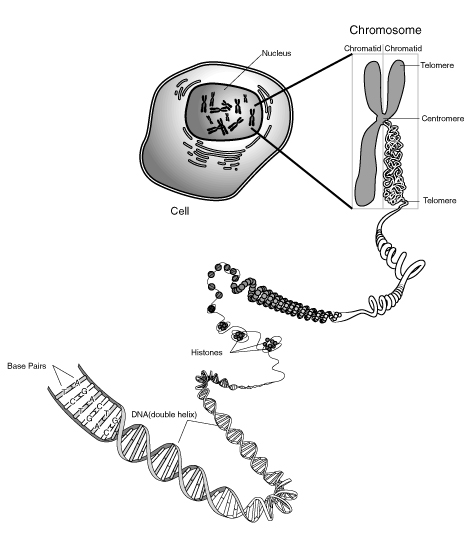
[edit | edit source]The human Diploid cell has 22 matching chromosomes and 2 sex chromosomes (Fig.4). One set is inherited from the father and one from the mother. Genes are arranged in a linear fashion along each chromosome. A haploid cell - either a sperm or an ovum contains 23 chromosomes. Fusion of the two cells in fertilization produce the stem cell from which all the diploid cells are made. In genetics a gene is a fundamental unit of hereditary, in biochemical or structural terms it is a sequence of DNA which can manufacture a single protein or RNA sequence.
The cell contains about 6 Billion nucleic acid base pairs divided into 46 chromosomes. The DNA in each chromosome is a single molecule between about 3 and 7 cms long. The total length of the DNA in a cell is about 2 meters. The question is how is all this packed into a single nucleus.
Chromosome Structure
[edit | edit source]The Chromosome is formed by wrapping DNA round a a double complex of 4 proteins (8 in all) called histones.
Fig. 6. DNA wrapping round histones
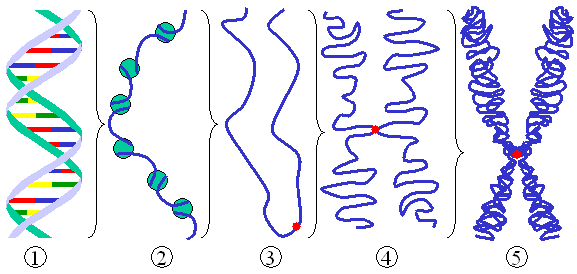
This results in super coiled strands of DNA:
In the interphase of cell production the strands exist as chromatin; in the prophase the strand and its transcribed copy will join at the centromere; and during metaphase the chromatin will become will become coiled into its chromosomal form
The following illustration summarises the process in graphic form:
Genes
[edit | edit source]We will consider genes in some detail when we look at gene expression. For now note that the human genome contains about 35,000 genes contained in different chromosomes. Note that there are really two definitions of a gene. The Mendelian definition - coined before it was even understood that DNA was the mechanism by which inheritence took place - which considered a gene to carry a unique in heritable trait, and the definition that is used in biochemistry - a stretch of DNA which is capable of creating a functioning protein or a functioning strand of RNA.
The Cell Replication Cycle
[edit | edit source]Briefly cell division consists of four stages:
- The cell manufactures the material necessary to duplicate its DNA
- It duplicates its DNA
- It manufactures the materials necessary to build a new cell
- The cell divides (mitosis)
This is illustrated Graphically below:
Fig. 10. Eukaryotic Replication Cycle
Note that the most visible part of the replication cycle - mitosis - is in fact only a small part of a much larger process. The longest part is the G1 phase where the cell manufactures all the proteins and RNA to produce a new cell and a duplicate set of DNA. The S phase, the duplication of the DNA takes 6-8 hours, and the final phase before mitosis, the G2 phase manufactures all the structural elements that the duplicate cell will need.
DNA replication
[edit | edit source]Replication of the chromosome is accomplished by the two strands splitting apart, and each strand acting as a template for a new DNA strand. Each parental strand thus acts as a template for a new strand running in the opposite direction to create a new chromosome. This replication is said to be semi-conservative, i.e. the parental strands are conserved, and each one is paired with a newly synthesised strand. The backbone of each strand consists of Phopho-Ribose part of the nucleic acid being joined to each other by phosphodiester bonds. The phosphate molecule, attached to the 5' carbon ribose atom attaches to the hydroxy molecule at the 3' ribose carbon molecule. (Fig.5). These carbon atoms are also used to name the orientation of the DNA strand, being 5' at one end, and 3' at the other.
The replication process starts by numerous 'bubbles' forming in the DNA as the strands split apart. (Fig.6). The formation of the bubbles allows replication to procede at numerous sites in the DNA molecule, thus speeding up the replication process. At each end of the bubble a 'replication fork' is formed. One strand of the fork has is nearest the 5' end of its DNA strand and the other is nearest to the 3' end of its strand. The DNA strand that runs from 3' to 5' - the 5' being closest to the replication fork being is called the leading strand, and the DNA strand that runs the other way is called the lagging strand.
Fig 11. Bubbles forming in the DNA molecule.
Unwinding of parental strands
[edit | edit source]The formation of the fork is under the control of two enzyme groups, the topoisomerases and the helicases. Helicase splits hydrogen bond and unwinds the DNA; topoisomerases takes up the stress of the unwinding by breaking then rejoining the phosphodiester bonds. At each nucleic acid of an unpaired base 'single strand binding proteins' are formed to prevent the strand being split up, or the exposed nucleic acids from being degraded. (Fig.7)
Fig 12. Formation of a replication fork
Lengthening the Transcribed DNA strand
[edit | edit source]δ-DNA polymerase can only add to an existing chain, it cannot initiate transcription. Initiation is accomplished using a strand of RNA known as RNA primer. This is attached to the parental strand at the initiation point. Two enzymes are involved - primase and α-DNA polymerase. Primase attaches a primer of about 7-8 nucleic acids, and α-DNA polymerase then begins the process of attaching nucleic acids. α-DNA polymerase is not nearly as processive as δ-DNA polymerase, so once the initiation has been made δ-DNA polymerase takes over and simply adds nucleic acids to the exposed 3' hydroxyl group of transcribed chain.
The leading parental strand, that which goes from 3' to 5', can be transcribed continuously from the initiation point until it meets the next 'bubble' along the chain - marked by the existence of a strand of primer. Once the RNA primer has been applied it is just a matter of adding nucleic acids to the exposed 3' hydroxyl group of the transcribed chain. The δ-DNA polymerase does not need to dis- and re-engage for each nucleic acid, but stays attached until it reaches a stretch of RNA primer. The δ-DNA polymerase is said to be highly processive and this speeds up the rate of duplication
A primer strand of RNA is added to the parental strand catalyzed by the enzymes primase and α-DNA polymerase. Nucleic acids are then continuously added catalyzed by δ-DNA polymerase. The single stranded binding proteins are removed as transcription proceeds. The transcription proceeds towards the replication fork.
The lagging strand cannot be transcribed continuously, but must be done in stages. As soon as a suitable length of the parental lagging strand has been exposed, initiation of transcription begins as described above using an RNA primer. Transcription is then carried out in the usual manner, away from the replication fork, and continues until it meets the next fragment. Thus the lagging strand is made in fragments - called Okazaki Fragments after scientist who first described them.
Okazaki fragments are formed by adding a primer strand of RNA catalyzed by the enzymes primase and α-DNA polymerase. Nucleic acids are then added catalyzed by δ-DNA polymerase until a previously added Okazaki is reached. The single stranded binding proteins are removed as transcription proceeds. Not the transcription proceeds away from the replication fork.
Removing the RNA Primer
[edit | edit source]When another Okazaki fragment is reached, the RNA primer is removed by δ-DNA polymerase assisted by the enzyme flap endonuclease; the gap left by its removal is filled by repair DNA polymerase; and the strands are ligated by DNA Ligase. This is summarized in the illustration below.
The RNA primer is removed, the gap between Okazaki framents is filled using repair DNA polymerases, and the fragments are joined using DNA ligase.
The 3' overhang problem
[edit | edit source]As the replication approaches the end of the chromosome, a problem occurs in the lagging strand. DNA polymerase can only add nucleic acids to the 3' end of the strand, so when the last Okazaki fragment is formed in the lagging, there is no way that than the RNA primer can be reconstituted in the normal manner. When the RNA primer breaks down we are left with an 'overhang' at the 3' end of the parental chain and shortening of the transcribed chain. This is illustrated below:
Without a remedy, this state of affairs can lead to chromosome shortening with each cell division, and the possible loss of essential genes. How can this state of affairs be remedied. For the answer we must look to telomers.
Telomeres and Chromosome shortening
[edit | edit source](There is much current research into telomeres because of their important role in aging and cancer, and the following description is an outline only of the subject. ["See wiki article telomeres"])
Telomeres are the ends of the eucaryotic chromosome. In humans they consist of numerous (in young humans this can number several thousand bases)repeats of the nucleic acid sequence TTAGGG and its complements. This is the first protection against degradation of the chromosome by overhang. The DNA being shed is merely the non-functional telomere, the TTAGGG sequence. However sooner or later the telomeres will be used up, and the chromosome will start shedding important genes. When this happens, the cell will usually die.
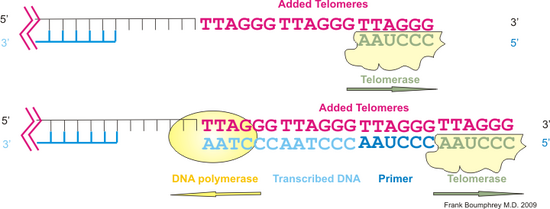
To overcome this problem, certain cells exhibit telomerase activity. Telomerase is a combination of a reverse transcriptase type enzyme, and an RNA strand, complementary to the telomere sequence. Telomerase will induce formation of the telomere DNA at the overhanging 3' end, adding nucleic acids to the 3' hydroxyl group. When enough telomeres have been added, primer is added, and DNA polymerase will transcribe the telomere strand similar to the action already described for Okazaki fragments. The fragments are joined by ligase. This is shown in the following illustration.
1. The enzyme induces telomeres to be added to the 3' overhang using its own complementary RNA strand.
2. When sufficient telomeres have been added, an RNA primer is attached, and DNA polymerase transcribes the existing DNA strand consisting mainly of telomeres.
3. The fragments are joined by Ligase in the usual manner.
Telomerase is not active in most cells although it is active in stem cells, germ cells, hair follicles and in 90 percent of cancer cells.
Telomeres and Aging
[edit | edit source]Telomere length is related to aging - older subjects by and large have shorter telomeres - and short telomeres appear to be particularly associated with 'cardia age'. Other factors also affect telomere length and this is the subject of much ongoing research.
Telomeres and Cancer
[edit | edit source]Telomerase is active in 90% of cancer cells, and must be active to confer 'immortality' on the cancer cell. Those cancer cells that do not have telomerase activity probably employ an alternative telomere lengthening process called alternative lengthening of telomeres (ALT), a non-conservative telomere lengthening pathway involving the transfer of telomere tandem repeats between sister-chromatids.
DNA Polymerases in Eucaryotes
[edit | edit source]At least 9 DNA polymerases have been discovered (Together with 3 for procaryotes). These all have different functions, and they all assist in DNA repair.
| Pol α | Initiation of replication in a complex with primase. Aids in starting the primer. DNA repair |
| Pol β | DNA repair exclusively |
| Pol γ | DNA replication in mitochondria |
| Pol δ | Replication by processive DNA synthesis on the leading and lagging (Okazaki fragments) strand. DNA repair |
| Pol ε | DNA replication. In some tissues takes the place of δ-DNA polymerase. DNA repair |
| Pol κ, η, ζ, ι | DNA repair (bypass polymerase) |
Table 1. Eukaryotic DNA Polymerases.
DNA Repair
[edit | edit source]DNA transcription is a complex process and is subject to error. As well as this there are numerous mutagenic effects that also operate to degrade DNA. Without an effective repair mechanism, these would accumulate, and the cell would soon be in trouble!
The δ-DNA polymerase enzyme has a 'repair' mechanism. As the δ-DNA polymerase adds nucleic acids, it audits them to make sure that
- The match is correct
- There is not a mutation to the nucleic acid.
If there is it pauses, and corrects the problem. Thus at the end of duplication of the strand there is only about one error in 106 base pairs. If the repair mechanism of δ-DNA polymerase is blocked, there is an error rate of about 103
However other DNA polymerases are acting to to audit and repair the DNA, and these reduce the error rare to between 109 and 1012. A remarkable fidelity!
There are several repair mechanisms, but they all follow the same basic pattern:
- The damaged portion of the chain is identified.
- The damaged portion is excised using endo- or exonucleases
- The removed portion is replaced using repair DNA polymerase
- The remaining 'nick' is repaired using DNA ligase.
This is shown in the following illustration.
Fig 18. Repair of a damaged strand of DNA
DNA Mutations
[edit | edit source]There are numerous environmental and other factors that can cause mutations. The following just mentions a few of the most common ones. Chemicals that cause mutations are called Mutagens. The sub-group that causes cancer are called Carcinogens.
"(wikipedia article on carcinogens)"
One of the first chemical carcinogens to be identified was Benzopyrene, a product of tobacco smoke. Benzopyrene is oxidized in the cell, and then can combine with nucleic acids causing a disruption in the DNA molecule.
Fig 19. Oxidized Benzopyrene molecule attached to Guanine
Fig 20. DNA model of Benzopyrene molecule attached to nucleic acid
Other common causes are radiation and ultraviolet light. These can generate hydroxyl radicals in the cell which can react with DNA, either altering the nucleic acids, or cleaving the DNA strand.
These mutations represent point mutations. Other mutations are caused by rearrangement of the chromosomal material. (See 'Genetic Rearrangement') below.
Fig 22. Five types of chromosmal mutations
More on mutations can be found at the wikipedia article "mutation"
Mitosis
[edit | edit source]Mitosis is the visible part of cell division. (See fig. 10). By the time mitosis begins all the 'heavy lifting' in the form of DNA replication and production of elements necessary for division has already been done.
At the start of mitosis, the duplicated DNA exists as chromatin joined at its centromere (See fig. 8:4), but has not yet been packaged into its chromosomal form.
Early Prophase. The Centiole divides, and the chromatin starts to condense. The centrioles are pushed to the poles of the cell by growth of microtubles and actin strands which appear as the 'spindles' under light microscope. The star like configuration, the asta that surrounds the centrioles is also made up of microtubules and actin strands.
Late Prophase. The Chromatin condenses into chromosomes (Se figs. 8 & ( above) The Nuclear membrane dissipates, and the chromosomes attach them selves to the spindles
Metaphase. The chromosomes align along the equator.
Anaphase. The chromosomes divide at their centromere, and start to 'walk' along the spindles to their respective poles.
Telophase. The nuclear membranes reform round the chromosomes; the chromosomes unwind to form chromatin (See fig 8:2); the cell constricts; and two new cells are formed.
The cell now either renters the G0 or G1 phase of cell replication.
Further details can be found at the "Wiki entry on Mitosis"
Meiosis
[edit | edit source]Meiosis only occurs in Germ cells, and produces ova in females and sperm in males. In the process of meiosis, the genetic material is reshuffled, and the chromosomes are reduced to their haploid number. During fertilisation when the ovum and sperm unite, the diploid number is restored.
We will revisit miosis when we look at reproductive physiology, but will also look at it here to see how genetic rearrangements occur in prophase I of meiosis.
Genetic Rearrangement & Gene reshuffling
[edit | edit source]Proto-oncogens and Cancer
[edit | edit source]Cell Death - Apoptosis
[edit | edit source](pronounced a-po-TOS-is from Greek for a petal dropping)