AP Biology/Membranes
Contents | Preface | Diagrams | Resources | Contributors | Edit Navigation Bar
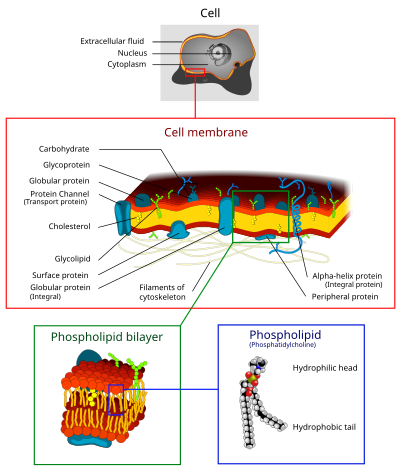
Introduction
[edit | edit source]The cell membrane is a semi-permeable membrane composed of all four types of macromolecules, with lipids and proteins being the most prevalent in dry weight. The membrane is present in all cells and functions to regulate incoming and outgoing materials, maintain intracellular homeostasis, and participate in signal transduction.
Models
[edit | edit source]Various models have been created to help visualize the complex structure of the cellular membrane, the most famous being the fluid mosaic model, which describes the membrane as a two-dimensional "sea" of molecular units, and the liquid crystal model, which describes the membrane as a sea of crystals with liquid properties.
Fluid Mosaic
[edit | edit source]The fluid mosaic model was developed in 1972 by S.J. Singer and Garth Nicolson. In their report, they proposed that biological membranes can be considered as a two-dimensional fluid composed of a mosaic of molecules, the majority of which had hydrophobic and hydrophilic properties (phospholipids). Thus, the molecular interactions create a membrane that is not solid, but which behaves like a fluid as molecules change positions and has the ability to stretch and contract vertically.
Liquid Crystal Model
[edit | edit source]The liquid crystal model was developed in 1970 by Freye and Edinin. It proposes that the phospholipid bilayer behaves like a series of crystals moving in an fluid, yet ordered manner.
Elements of the Membrane
[edit | edit source]The phospholipid bilayer is composed of the following in dry weight:
- 50% proteins
- 30% lipids
- 5% glycolipids
- 15% other
Each element of the membrane performs a specific function in sensory transduction, transport, etc.
Proteins
[edit | edit source]
Glycoproteins
Glycoproteins are transmembrane proteins with oligosaccharides covalently bonded to their amino acid chains. Glycoproteins generally play an important role in sensory transduction, particularly with regard to immune system responses.

The membrane is represented in light brown.
Transmembrane Proteins
Transmembrane proteins are proteins that extend through the membrane, having both an exterior and an interior domain. They are integral proteins, meaning they are permanently bonded to the PLB. Many function as hormone/signal acceptors, ion channels, and transport proteins. As a rule, they act as channels between the intra- and extracellular spaces.
There are two structural categories of transmembrane protein:
- Alpha-helical, those composed of repeating hydrogen bonds that make up alpha helices on the secondary level of structure
- Beta barrel, those which have repeating hydrogen bonds that form pleated sheets
Alpha-helical make up the largest group, with beta barrel proteins found primarily in mitochondrial and other prokaryotic gram-positive membranes.
Peripheral Proteins
Peripheral proteins are proteins that are bonded by weaker interactions to either the interior or the exterior of the membrane. Their loose bonds allow them to detach and function as secondary messengers or extracellular messengers upon receipt of a signal from the cell. Some prominent functions are in metabolism and in sensory transduction.
Lipids
[edit | edit source]Phospholipids
Phospholipids are the "glue" that gives the membrane its distinct structure. Their amphipathic nature and tendency to create micelles in water were likely the origin of the first cells.
Glycolipids
Glycolipids are lipids with attached carbohydrates. Their most prominent function is providing extracellular support to form tissues in multicellular organisms.
Cholesterols
Cholesterol molecules are lipids distinguished by their four-ring structure. In the membrane, they act to resist as a type of "anti-freeze" to prevent freezing and overheating.
Transmembrane Transport
[edit | edit source]Passive
[edit | edit source]Osmosis and Diffusion

Porin Proteins
Facilitated Diffusion
[edit | edit source]Active Transport
[edit | edit source]phospholipid bilayer (plasma membrane)
- proteins (peripheral, integral, transmembrane, adhesion, recepto, channel, recognition and adhesion)
- cholesterol
simple diffusion osmosis channel proteins facilitated transport active transport pumps (h+ and na/k) endo-, pino-, phago- cytosis receptor-mediated endocytosis bulk flow exocytosis
