Biochemistry/Proteins/Introduction
Protein role and importance[edit | edit source]
Proteins are among the fundamental molecules of biology. They are common to all life present on Earth today, and are responsible for most of the complex functions that make life possible. They are also the major structural constituent of living beings. According to the Central Dogma of Molecular Biology (proposed by Francis Crick in 1958), information is transferred from DNA to RNA to proteins. DNA functions as a storage medium for the information necessary to synthesize proteins, and RNA is responsible for (among other things) the translation of this information into protein molecules, as part of the ribosome.
Virtually all the complex chemical functions of the living cell are performed by protein-based catalysts called enzymes. Specifically, enzymes either make or break chemical bonds. Protein enzymes should not be confused with RNA-based enzymes (also called ribozymes), a group of macromolecules that perform functions similar to protein enzymes. Further, most of the scaffolding that holds cells and organelles together is made of proteins. In addition to their catalytic functions, proteins can transmit and commute signals from the extracellular environment, duplicate genetic information, assist in transforming the energy in light and chemicals with astonishing efficiency, convert chemical energy into mechanical work, and carry molecules between cell compartments.
Functions not performed by proteins[edit | edit source]
Proteins do so much that it's important to note what proteins don't do. Currently there are no known proteins that can directly replicate themselves. Prions are no exception to this rule. It is theorized that prions may be able to act as a structural template for other chemically (but not structurally) identical proteins, but they can't function as a native template for protein synthesis de novo. Proteins don't act as fundamental energy reserves in most organisms, as their metabolism is slower and inefficient compared to sugars or lipids. They are, on the other hand, a fundamental nitrogen and amino acid reserve for many organisms. Proteins do not directly function as a membrane in most organisms, except viruses; however, they are often important components of these structures, lending both stability and structural support.
Proteins as polymers of amino acids[edit | edit source]
Composition and Features[edit | edit source]
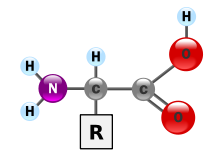
Proteins are composed of a linear (not branched and not forming rings) polymer of amino acids. The twenty genetically encoded amino acids are molecules that share a central core: The α-carbon is bonded to a primary amino (-NH2) terminus, a carboxylic acid (-COOH) terminus, a hydrogen atom, and the amino acid side chain, also called the "R-group". The R-group determines the identity of the amino acid. In an aqueous solution, at physiological pH (~6.8), the amino group will be in the protonated -NH3+ form, and the carboxylic acid will be in the deprotonated -COO- form, forming a zwitterion. Most amino acids that make up proteins are L-isomers, although a few exotic creatures use D-isomers in their proteins. It is important to note that levorotatory (L) and dextrorotatory (D) are not specific to rectus (R) and sinister (S) configurations. A levorotatory form of a protein can be either R or S configuration. Levorotatory and dextrorotatory refer to how the proteins bend light in a polarimeter.
Amino acids polymerize via peptide bonds, which is a type of amide bond. A peptide bond is formed upon the dehydration of the carboxy-terminus of one amino acid with the amine terminus of a second amino acid. The resulting carbonyl group's carbon atom is directly bound to the nitrogen atom of a secondary amine. A peptide chain will have an unbound amino group free at one end (called the N-terminus) and a single free carboxylate group at the other end (called the C-terminus).
The written list of the amino acids linked together in a protein, in order, is called its primary structure. By convention, peptide sequences are written from N-terminus to C-terminus. This convention mimics the way polypeptides are synthesized by the ribosome in the cell. Small polymers of less than 20 amino acids long are more often called peptides or polypeptides. Proteins can have sequences as short as 20-30 amino acids to gigantic molecules of more than 3,000 amino acids (like Titin,a human muscle protein).
Genetically-Encoded Amino Acids[edit | edit source]
While there are theoretically billions of possible amino acids, most proteins are formed of only 20 amino acids: the genetically-encoded (or more precisely, proteogenic) amino acids. Note that all amino acids except glycine have a chiral center at their α-carbons. (Glycine has two hydrogens on its α-carbon, and therefore it is achiral.) Besides glycine, all proteogenic amino acids are L-amino acids, meaning they have the same absolute configuration as L-glyceraldehyde. This is the same as the S-configuration, with the exception of cysteine, which contains a sulfur atom in its side chain, and so the naming priority changes. D-amino acids are sometimes found in nature, as in the cell walls of certain bacteria, but they are rarely incorporated into protein chains.
The side-chains of proteogenic amino acids are quite varied: they range from a single hydrogen atom (as for glycine, the simplest amino acid) to the indole heterocycle, as found in tryptophan. There are polar, charged and hydrophobic amino acids. The chemical richness of amino acids is at the base of the complexity and versatility of proteins.
Post-Translational Modification[edit | edit source]
There are two major types of post-translational modifications to protein: those that cleave the bonds of the peptide backbone and those that add or remove functional groups to the sidechains of individual amino acids. In the first type of post-translational modification, specialized enzymes called proteases recognize specific amino acids of a protein and break the associated peptide bond, thereby irreversibly modifying the primary structure. In the second major type of modification, the amino acid side chains of a given protein are chemically modified by enzymatic reactions or are spontaneously formed (non-enzymatic). Examples of sidechain modifications are quite numerous, but common ones include oxidation, acylation, glycosylation (addition of a glycan, or sugar), methylation, and phosphorylation. Both types of post-translational modifications are capable of exerting positive and negative control over a given protein or enzyme.
The importance of protein structure[edit | edit source]
Generally speaking, the function of a protein is completely determined by its structure. Molecules like DNA, which perform a fairly small set of functions, have an almost fixed structure that's fairly independent from sequence. By contrast, protein molecules perform functions as different as digesting sugars or moving muscles. To perform so many different functions, proteins come in many different structures. The protein function is almost completely dependent on protein structure. Enzymes must recognize and react with their substrates with precise positioning of critical chemical groups in the three-dimensional space. Scaffold proteins must be able to precisely dock other proteins or components and position them in space in the correct fashion. Structural proteins like Collagen must face mechanical stresses and be able to build a regular matrix where cells can adhere and proliferate. Motor proteins must reversibly convert chemical energy in movement, in a precise fashion.
Protein folding depends on sequence[edit | edit source]
As Anfinsen demonstrated in the 1960's, proteins acquire their structure by spontaneous folding of the polypeptide chain into the minimal energy configuration. Most proteins require no external factors in order to fold (although specialized protein exist in cells, called chaperones, that help other, misfolded, proteins acquire their correct structure) — the protein sequence itself uniquely determines the structure. Often the whole process takes place in milliseconds. Despite the apparent chemical simplicity of proteins, the vast number of permutations of twenty amino acids in a linear sequence leads to an amazing number of different protein folds. Nevertheless, protein structures share several characteristics in common: they are almost all built of a few secondary structure elements (short-range structural patterns that are recurrent in protein structures) and even the way these elements combine is often repeated in common motifs. Nonetheless, it is still impossible to know what structure a given protein sequence will yield in solution. This is known as the protein folding problem, and it is one of the most important open problems in modern molecular biology.
Protein denaturation[edit | edit source]
Proteins can lose their structure if put in unsuitable chemical (e.g. high or low pH; high salt concentrations; hydrophobic environment) or physical (e.g. high temperature; high pressure) conditions. This process is called denaturation. Denatured proteins have no defined structure and, especially if concentrated, tend to aggregate into insoluble masses. Protein denaturation is by no means an exotic event: a boiled egg becomes solid just because of denaturation and subsequent aggregation of its proteins. Denatured proteins can sometimes refold when put again in the correct environment, but sometimes the process is irreversible (especially after aggregation: the boiled egg is again an example). It is finally the proteins which are responsible for susceptibility or resistance to a pathogen or parasite.
Proteins can fold into domains[edit | edit source]
A significant number of proteins, especially large proteins, have a structure divided into several independent domains. These domains can often perform specific functions in a protein. For example, a cell membrane receptor might have an extracellular domain to bind a target molecule and an intracellular domain that binds other proteins inside the cell, thereby transducing a signal across the cell membrane.
The domain of a protein is determined by the secondary structure of a protein there are four main types of domain structures: alpha-helix, beta-sheet, beta-turns, and random coil.
The alpha-helix is when the polypeptide chain forms a helix shape with the amino acids side chains sticking out, usually about 10 amino acids long. The alpha-helix gets its strength by forming internal hydrogen bonds, that occur between amino acid 1 and 4 along the length of the helix. A high concentration of Glycine's in a row tend towards the alpha-helix conformation.
The beta-sheet structure is composed of poly-peptide chains stacking forming hydrogen bonds between the sheets. You can form parallel sheets by stacking in the same direction N-C terminal on top of N-C terminal or form anti-parallel by stacking in opposite directions N-C terminal on top of C-N terminal. Beta-turns link two anti-parallel beta strands by a 4 amino acid loop in a defined conformation.
A random coil is a portion of the protein that has no defined secondary structure.
Domains of a protein then come from unique portions of the peptide that are made up of these types of secondary structure.