An Introduction to Molecular Biology/Replication of DNA and its repair

As we know Cell division is essential for an organism to grow, but, when a cell divides, it must replicate the DNA (DNA replication take place during S phase) in its genome so that the two daughter cells have the same genetic information as their parent. The double-stranded structure of DNA provides a simple mechanism for DNA replication. Here, the two strands are separated and then each strand's complementary DNA sequence is recreated by an enzyme called DNA polymerase. This enzyme makes the complementary strand by finding the correct base through complementary base pairing, and bonding it onto the original strand. As DNA polymerases can only extend a DNA strand in a 5′ to 3′ direction, different mechanisms are used to copy the antiparallel strands of the double helix. In this way, the base on the old strand dictates which base appears on the new strand, and the cell ends up with a perfect copy of its DNA.
Replication
[edit | edit source]
a: template, b: leading strand, c: lagging strand, d: replication fork, e: primer, f: Okazaki fragments
In a cell, DNA replication begins at specific locations in the genome, called "origins". Unwinding of DNA at the origin, and synthesis of new strands, forms a replication fork. In addition to DNA polymerase, the enzyme that synthesizes the new DNA by adding nucleotides matched to the template strand, a number of other proteins are associated with the fork and assist in the initiation and continuation of DNA synthesis. DNA replication can also be performed in vitro (outside a cell). DNA polymerases, isolated from cells, and artificial DNA primers are used to initiate DNA synthesis at known sequences in a template molecule. The polymerase chain reaction (PCR), a common laboratory technique, employs such artificial synthesis in a cyclic manner to amplify a specific target DNA fragment from a pool of DNA.[1]
Leading strand
[edit | edit source]The leading strand template is the template strand of the DNA double helix that is oriented in a 3' to 5' manner. All DNA synthesis occurs 5'-3'. The original DNA strand must be read 3'-5' to produce a 5'-3' nascent strand. The leading strand is formed along the leading strand template as a polymerase "reads" the template DNA and continuously adds nucleotides to the 3' end of the elongating strand. This polymerase is DNA polymerase III (DNA Pol III) in prokaryotes and presumably Pol ε in eukaryotes.
Lagging strand
[edit | edit source]
The lagging strand template is the coding strand of the DNA double helix that is oriented in a 5' to 3' manner. The newly made lagging strand still is synthesized 5'-3'. However, since the DNA is oriented in a manner that does not allow continual synthesis, only small sections can be read at a time. An RNA primer is placed on the DNA strand 3' to the origin of replication. Just as before, DNA Polymerase reads 3'-5' on the original DNA to produce a 5'-3' nascent strand. Polymerase reaches the origin of replication and stops replication until a new RNA primer is placed 3' to the last RNA primer. These fragments of DNA produced on the lagging strand are called Okazaki fragments. The orientation of the original DNA on the lagging strand prevents continual synthesis. As a result, replication of the lagging strand is more complicated than of the leading strand. On the lagging strand template, primase "reads" the DNA and adds RNA to it in short, separated segments. In eukaryotes, primase is intrinsic to Pol α. DNA polymerase III or Pol δ lengthens the primed segments, forming Okazaki fragments. Primer removal in eukaryotes is also performed by Pol δ. In prokaryotes, DNA polymerase I "reads" the fragments, removes the RNA using its flap endonuclease domain, and replaces the RNA nucleotides with DNA nucleotides (this is necessary because RNA and DNA use slightly different kinds of nucleotides). DNA ligase joins the fragments together.
Okazaki fragment
[edit | edit source]An Okazaki fragment is a relatively short fragment of DNA (with no RNA primer at the 5' terminus) created on the lagging strand during DNA replication. The lengths of Okazaki fragments are between 1,000 and 2,000 nucleotides long in E. coli and are generally between 100 and 200 nucleotides long in eukaryotes. It was originally discovered in 1968 by Reiji Okazaki, Tsuneko Okazaki, and their colleagues while studying replication of bacteriophage DNA in Escherichia coli.[2][3]

Rate of replication
[edit | edit source]The rate of DNA replication in a living cell was first measured as the rate of phage T4 DNA elongation in phage-infected E. coli.[4] During the period of exponential DNA increase at 37 °C, the rate was 749 nucleotides per second. The mutation rate per base pair per replication during phage T4 DNA synthesis is 1.7 per 108.[5] Thus semiconservative DNA replication is both rapid and accurate.
Classification of DNA polymerase
[edit | edit source]Based on sequence homology, DNA polymerases are subdivided into seven different families: A, B, C, D, X, Y, and RT.
1.Family A Polymerases contain both replicative and repair polymerases. Replicative members from this family include the extensively-studied T7 DNA polymerase, as well as the eukaryotic mitochondrial DNA Polymerase γ. Among the repair polymerases are Escherichia coli DNA pol I, Thermus aquaticus pol I, and Bacillus stearothermophilus pol I. These repair polymerases are involved in excision repair and processing of Okazaki fragments generated during lagging strand synthesis.
2.Family B In XPV patients, alternative error-prone polymerases, e.g., Pol ζ (zeta) (polymerase ζ is a B Family polymerase a complex of the catalytic subunit REV3L with Rev7, which associates with Rev1), are thought to be involved in mistakes that result in the cancer predisposition of these patients. The DNA polymerase which belongs to B family contain DTDS motif. The other members are Pol ε, Pol α, Pol δ.
3.Family C Polymerases are the primary bacterial chromosomal replicative enzymes. DNA Polymerase III alpha subunit from E. coli is the catalytic subunit and possesses no known nuclease activity. A separate subunit, the epsilon subunit, possesses the 3'-5' exonuclease activity used for editing during chromosomal replication. Recent research has classified Family C polymerases as a subcategory of Family X.
4.Family D Polymerases are still not very well characterized. All known examples are found in the Euryarchaeota subdomain of Archaea and are thought to be replicative polymerases.
5.Family X Contains the well-known eukaryotic polymerase pol β, as well as other eukaryotic polymerases such as pol σ, pol λ, pol μ, and terminal deoxynucleotidyl transferase (TdT). Pol β is required for short-patch base excision repair, a DNA repair pathway that is essential for repairing abasic sites. Pol λ and Pol μ are involved in non-homologous end-joining, a mechanism for rejoining DNA double-strand breaks. TdT is expressed only in lymphoid tissue, and adds "n nucleotides" to double-strand breaks formed during V(D)J recombination to promote immunological diversity. The yeast Saccharomyces cerevisiae has only one Pol X polymerase, Pol IV, which is involved in non-homologous end-joining.
6.Family Y Y Polymerases differ from others in having a low fidelity on undamaged templates and in their ability to replicate through damaged DNA. Members of this family are hence called translesion synthesis (TLS) polymerases. Depending on the lesion, TLS polymerases can bypass the damage in an error-free or error-prone fashion, the latter resulting in elevated mutagenesis. Xeroderma pigmentosum variant (XPV) patients for instance have mutations in the gene encoding Pol η (eta), which is error-free for UV-lesions. Other members in humans are Pol ι (iota), Pol κ (kappa), and Rev1 (terminal deoxycytidyl transferase). In E. coli, two TLS polymerases, Pol IV (DINB) and Pol V (UmuD'2C), are known.
7.Family RT (reverse transcriptase) The reverse transcriptase family contains examples from both retroviruses and eukaryotic polymerases. The eukaryotic polymerases are usually restricted to telomerases. These polymerases use an RNA template to synthesize the DNA strand.
Replication is semiconservative
[edit | edit source]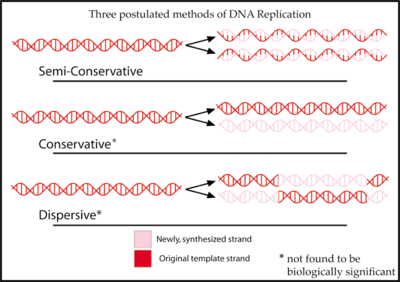
The Meselson and Stahl experiment was an experiment by Matthew Meselson and Franklin Stahl in 1958 which supported the hypothesis that DNA replication was semiconservative. Semiconservative replication means that when the double stranded DNA helix was replicated, each of the two double stranded DNA helices consisted of one strand coming from the original helix and one newly synthesized. It has been called "the most beautiful experiment in biology.[6]"
Three hypotheses had been previously proposed for the method of replication of DNA.
In the semiconservative hypothesis, proposed by Watson and Crick, the two strands of a DNA molecule separate during replication. Each strand then acts as a template for synthesis of a new strand.[7]
The conservative hypothesis proposed that the entire DNA molecule acted as a template for synthesis of an entirely new one. According to this model, histone proteins bound to the DNA, distorting it in such a way as to expose both strands' bases for hydrogen bonding.[8]
The dispersive hypothesis is exemplified by a model proposed by Max Delbrück, which attempts to solve the problem of unwinding the two strands of the double helix by a mechanism that breaks the DNA backbone every 10 nucleotides or so, untwists the molecule, and attaches the old strand to the end of the newly synthesized one. This would synthesize the DNA in short pieces alternating from one strand to the other.[9]
Each of these three models makes a different prediction about the distribution of the "old" DNA in molecules formed after replication. In the conservative hypothesis, after replication, one molecule is the entirely conserved "old" molecule, and the other is all newly synthesized DNA. The semiconservative hypothesis predicts that each molecule after replication will contain one old and one new strand. The dispersive model predicts that each strand of each new molecule will contain a mixture of old and new DNA.[10]
The semi-conservative theory can be confirmed by making use of the fact that DNA is made up of nitrogen bases. Nitrogen has an isotope N15 (N14 is the most common isotope) called heavy nitrogen. The experiment that confirms the predictions of the semi-conservative theory[11][12] makes use of this isotope and runs as follows: Bacterial (E coli) DNA is placed in a media containing heavy nitrogen(N15), which binds to the DNA, making it identifiable. Bacteria containing this DNA are then placed in a media with the presence of N14 and left to replicate only once. The new bases will contain nitrogen 14 while the originals will contain N15 The DNA is placed in test tubes containing caesium chloride (heavy compound) and centrifuged at 40,000 revolutions per minute. The caesium chloride molecules sink to the bottom of the test tubes creating a density gradient. The DNA molecules will position at their corresponding level of density (taking into account that N15 is more dense than N14) These test tubes are observed under ultraviolet rays. DNA appears as a fine layer in the test tubes at different heights according to their density. According to the semi-conservative theory, after one replication of DNA, we should obtain 2 hybrid (part N14 part N15) molecules from each original strand of DNA. This would appear as a single line in the test tube. This result would be the same for the dispersive theory. On the other hand, according to the conservative theory, we should obtain one original DNA duplex and a completely new one i.e. two fine lines in the test tube placed separately one from the other. Up to this point, either the semi-conservative or the dispersive theories could be truthful, as experimental evidence confirmed that only one line appeared after one replication. In order to conclude between those two, DNA had to be left to replicate again, still in a media containing N14. In the dispersive theory, after 2 divisions we should obtain a single line, but further up in the test tube, as the DNA molecules become less dense as N14 becomes more abundant in the molecule According to the semi-conservative theory, 2 hybrid molecules and 2 fully N14 molecules should be produced, so two fine lines at different heights in the test tubes should be observed. Experimental evidence confirmed that two lines were observed therefore offering compelling evidence for the semi-conservative theory.
Genetic evidence
An independent 'genetic' evidence for the semi-conservative theory was provided more recently by high throughput genomic sequencing of individual mutagenized bacteria. E. coli were treated with Ethyl methanesulfonate (EMS), known to induce G:C → A:T transitions due to generation of abnormal base O-6-ethylguanine, which is further misrecognized during DNA replication and paired with T instead of C. The sequenced DNA from individual colonies of EMS-mutagenized bacteria exhibited long stretches of solely G → A or C → T transitions, which in some cases were spanning entire bacterial genome. The elementary explanation of this observation is based on semi-conservative mechanism: one should expect the segregation between daughter strands into different cells after replication, which leads to each descendant cell having exclusively G → A or C → T conversions.
{{quotation|

Nitrogen is a major constituent of DNA. 14N is by far the most abundant isotope of nitrogen, but DNA with the heavier (but non-radioactive) 15N isotope is also functional.
E. coli were grown for several generations in a medium with 15N. When DNA is extracted from these cells and centrifuged on a salt density gradient, the DNA separates out at the point at which its density equals that of the salt solution. The DNA of the cells grown in 15N medium had a higher density than cells grown in normal 14N medium. After that, E. coli cells with only 15N in their DNA were transferred to a 14N medium and were allowed to divide; the progress of cell division was monitored by measuring the optical density of the cell suspension.
DNA was extracted periodically and was compared to pure 14N DNA and 15N DNA. After one replication, the DNA was found to have close to the intermediate density. Since conservative replication would result in equal amounts of DNA of the higher and lower densities (but no DNA of an intermediate density), conservative replication was excluded. However, this result was consistent with both semiconservative and dispersive replication. Semiconservative replication would result in double-stranded DNA with one strand of 15N DNA, and one of 14N DNA, while dispersive replication would result in double-stranded DNA with both strands having mixtures of 15N and 14N DNA, either of which would have appeared as DNA of an intermediate density.
The authors continued to sample cells as replication continued. DNA from cells after two replications had been completed was found to consist of equal amounts of DNA with two different densities, one corresponding to the intermediate density of DNA of cells grown for only one division in 14N medium, the other corresponding to DNA from cells grown exclusively in 14N medium. This was inconsistent with dispersive replication, which would have resulted in a single density, lower than the intermediate density of the one-generation cells, but still higher than cells grown only in 14N DNA medium, as the original 15N DNA would have been split evenly among all DNA strands. The result was consistent with the semiconservative replication hypothesis [11]
Replication in prokaryote
[edit | edit source]DNA replication in prokaryotes is extensively studied in E. coli. It is bi-directional and originates at a single origin of replication (OriC).
Primase
[edit | edit source]In bacteria, primase binds to the DNA helicase forming a complex called the primosome. Primase is activated by DNA helicase where it then synthesizes a short RNA primer approximately 11 ±1 nucleotides long, to which new nucleotides can be added by DNA polymerase.
Primosome
[edit | edit source]A primosome is a protein complex responsible for creating RNA primers on single stranded DNA during DNA replication.Primosomes are nucleoproteins assemblies that activate DNA replication forks. Their primary role is to recruit the replicative helicase onto single-stranded DNA. The "replication restart" primosome, defined in Escherichia coli, is involved in the reactivation of arrested replication forks.
Assembly of the Escherichia coli primosome requires six proteins, PriA, PriB, PriC, DnaB, DnaC, and DnaT, acting at a primosome assembly site (pas) on an SSBcoated single-stranded (8s) DNA. Assembly is initiated by interactions of PriA and PriB with ssDNA and the pas. PriC, DnaB, DnaC, and DnaT then act on the PriAPriB- DNA complex to yield the primosome.
The primosome consists of seven proteins: DnaG primase, DnaB helicase, DnaC helicase assistant, DnaT, PriA, Pri B, and PriC. The primosome is utilized once on the leading strand of DNA and repeatedly, initiating each Okazaki fragment, on the lagging DNA strand. Initially the complex formed by PriA, PriB, and PriC binds to DNA. Then the DnaB-DnaC helicase complex attaches along with DnaT. This structure is referred to as the pre-primosome. Finally, DnaG will bind to the pre-primosome forming a complete primosome. The primosome attaches 1-10 RNA nucleotides to the single stranded DNA creating a DNA-RNA hybrid. This sequence of RNA is used as a primer to initiate DNA polymerase III. The RNA bases are ultimately replaced with DNA bases by RNase H nuclease (eukaryotes) or DNA polymerase I nuclease (prokaryotes). DNA Ligase then acts to join the two ends together.
Elongation of DNA strand
[edit | edit source]Once priming is complete, DNA polymerase III holoenzyme is loaded into the DNA and replication starts. The catalytic mechanism of DNA polymerase III involves the use of two metal ions in the active site, and a region in the active site that can discriminate between deoxyribonucleotides and ribonucleotides. The metal ions are general divalent cations that help the 3' OH initiate a nucleophilic attack onto the alpha phosphate of the deoxyribonucleotide and orient and stabilize the negatively charged triphosphate on the deoxyribonucleotide. Nucleophilic attack by the 3' OH on the alpha phosphate releases pyrophosphate, which is then subsequently hydrolyzed (by inorganic phosphatase) into two phosphates. This hydrolysis drives DNA synthesis to completion.
Furthermore, DNA polymerase III must be able to distinguish between correctly paired bases and incorrectly paired bases. This is accomplished by distinguishing Watson-Crick base pairs through the use of an active site pocket that is complementary in shape to the structure of correctly paired nucleotides. This pocket has a tyrosine residue that is able to form van der Waals interactions with the correctly paired nucleotide. In addition, dsDNA (double stranded DNA) in the active site has a wider and shallower minor groove that permits the formation of hydrogen bonds with the third nitrogen of purine bases and the second oxygen of pyrimidine bases. Finally, the active site makes extensive hydrogen bonds with the DNA backbone. These interactions result in the DNA polymerase III closing around a correctly paired base. If a base is inserted and incorrectly paired, these interactions could not occur due to disruptions in hydrogen bonding and van der Waals interactions.
DNA is read in the 3' → 5' direction, therefore, nucleotides are synthesized (or attached to the template strand) in the 5' → 3' direction. However, one of the parent strands of DNA is 3' → 5' while the other is 5' → 3'. To solve this, replication occurs in opposite directions. Heading towards the replication fork, the leading strand is synthesized in a continuous fashion, only requiring one primer. On the other hand, the lagging strand, heading away from the replication fork, is synthesized in a series of short fragments known as Okazaki fragments, consequently requiring many primers. The RNA primers of Okazaki fragments are subsequently degraded by RNAse H and DNA Polymerase I (exonuclease), and the gap (or nicks) are filled with deoxyribonucleotides and sealed by the enzyme ligase.[13]
Termination
[edit | edit source]Termination of DNA replication in E. coli is completed through the use of termination sequences and the Tus protein. Tus is a sequence-specific DNA binding protein that promotes termination in prokaryotic DNA replication. In E. Coli, Tus binds to ten closely related 23 basepair binding sites encoded in the bacterial chromosome. These sites, called Ter sites, are designated TerA, TerB, ..., TerJ. The binding sites are asymmetric, such that when a Tus-Ter complex (Tus protein bound to a Ter site) is encountered by a replication fork from one direction, the complex is dissociated and replication continues (permissive). When encountered from the other direction, however, the Tus-Ter complex provides a much larger kinetic barrier and halts replication (non-permissive). The multiple Ter sites in the chromosome are oriented such that the two oppositely moving replication forks are both stalled in the desired termination region.
DNA polymerase in prokaryote
[edit | edit source]
In Prokaryotic there are 5 kind of DNA polymerases:
Pol I: implicated in DNA repair; has 5'->3' polymerase activity, and both 3'->5' exonuclease activity (proofreading) and 5'->3' exonuclease activity (RNA primer removal).
Pol II: involved in repairing damaged DNA; has 3'->5' exonuclease activity.The enzyme is 90 kDa in size and is coded by the polB gene. DNA Pol II can synthesize DNA new base pairs at an average rate of between 40 and 50 nucleotides/second.
Pol III: the main polymerase in bacteria (responsible for elongation); has 3'->5' exonuclease activity (proofreading).The replisome is composed of the following: 2 DNA Pol III enzymes, made up of α, ε and θ subunits. the α subunit synthesizes the RNA/DNA primer. the ε subunit synthesizes the leading strand. the θ subunit stimulates the ε subunit's proofreading. 2 β units which act as sliding DNA clamps, they keep the polymerase bound to the DNA. 2 τ units which acts to dimerize two of the core enzymes (α, ε, and θ subunits). 1 γ unit which acts as a clamp loader for the lagging strand Okazaki fragments, helping the two β subunits to form a unit and bind to DNA. The γ unit is made up of 5 γ subunits which include 3 γ subunits, 1 δ subunit, and 1 δ' subunit. The δ is involved in copying of the lagging strand
Pol IV: a Y-family DNA polymerase.
Pol V: a Y-family DNA polymerase; participates in bypassing DNA damage.
DNA Polymerase I or Pol I
[edit | edit source]DNA Polymerase I (or Pol I) is an enzyme that participates in the process of DNA replication in prokaryotes. It contains 928 amino acids, and is an example of a processive enzyme - it can sequentially catalyze multiple polymerisations. It was Discovered by Arthur Kornberg in 1956, it was the first known DNA polymerase (and, indeed, the first known of any kind of polymerase). It was initially characterized in E. coli, although it is ubiquitous in prokaryotes. In E. coli and many other bacteria, the gene which encodes Pol I is known as polA. Pol I possesses three enzymatic activities: (1) a 5' -> 3' (forward) DNA polymerase activity, requiring a 3' primer site and a template strand; (2) a 3' -> 5' (reverse) exonuclease activity that mediates proofreading; and (3) a 5' -> 3' (forward) exonuclease activity mediating nick translation during DNA repair.
Klenow fragment
[edit | edit source]The 5' → 3' exonuclease activity of DNA polymerase I from E. coli makes it unsuitable for many applications, the Klenow fragment, which lacks this activity, can be very useful in research. The Klenow fragment is extremely useful for research-based tasks such as: (1) Synthesis of double-stranded DNA from single-stranded templates; (2) Filling in (meaning removal of overhangs to create blunt ends) recessed 3' ends of DNA fragments; (3) Digesting away protruding 3' overhangs; (4) Preparation of radioactive DNA probes. The Klenow fragment was also the original enzyme used for greatly amplifying segments of DNA in the polymerase chain reaction (PCR) process, before being replaced by thermostable enzymes such as Taq polymerase.
Helicases and Topoisomerase
[edit | edit source]Many cellular processes (DNA replication, transcription, translation, recombination, DNA repair, ribosome biogenesis) involve the separation of nucleic acid strands. Helicases are often utilized to separate strands of a DNA double helix or a self-annealed RNA molecule using the energy from ATP hydrolysis, a process characterized by the breaking of hydrogen bonds between annealed nucleotide bases. They move incrementally along one nucleic acid strand of the duplex with a directionality and processivity specific to each particular enzyme. There are many helicases (14 confirmed in E. coli, 24 in human cells) resulting from the great variety of processes in which strand separation must be catalyzed.
Helicases adopt different structures and oligomerization states. Whereas DnaB-like helicases unwind DNA as donut-shaped hexamers, other enzymes have been shown to be active as monomers or dimers. Studies have shown that helicases may act passively, waiting for uncatalyzed unwinding to take place and then translocating between displaced strands,[1] or can play an active role in catalyzing strand separation using the energy generated in ATP hydrolysis. In the latter case, the helicase acts comparably to an active motor, unwinding and translocating along its substrate as a direct result of its ATPase activity. Helicases may process much faster in vivo than in vitro due to the presence of accessory proteins that aid in the destabilization of the fork junction. Defects in the gene that codes helicase cause Werner syndrome, a disorder characterized by the appearance of premature aging.[14]
Superfamilies
Helicases have been classified in 5 superfamilies (SF1-SF5). All of the proteins bind ATP, and, as a consequence, all of them carry the classical Walker A (phosphate-binding loop or P-loop) and Walker B (Mg2+-binding aspartic acid) motifs.
Superfamily I: UvrD (E. coli, DNA repair), Rep (E. coli, DNA replication), PcrA (Staphylococcus aureus, recombination), Dda (bacteriophage T4, replication initiation), RecD (E. coli, recombinational repair), TraI (F-plasmid, conjugative DNA transfer). This family includes RNA helicases thought to be involved in duplex unwinding during viral RNA replication. Members of this family are found in positive-strand single-stranded RNA viruses from superfamily 1. This helicase has multiple roles at different stages of viral RNA replication, as dissected by mutational analysis.
Superfamily II: RecQ (E. coli, DNA repair), eIF4A (Baker's Yeast, RNA translation), WRN (human, DNA repair), NS3[5] (Hepatitis C virus, replication). TRCF (Mfd) (E.coli, transcription-repair coupling).
Superfamily III: LTag (Simian Virus 40, replication), E1 (human papillomavirus, replication), Rep (Adeno-Associated Virus, replication, viral integration, virion packaging). Superfamily 3 consists of helicases encoded mainly by small DNA viruses and some large nucleocytoplasmic DNA viruses.[6][7] Small viruses are very dependent on the host-cell machinery to replicate. SF3 helicase in small viruses is associated with an origin-binding domain. By pairing a domain that recognises the ori with a helicase, the virus can bypass the host-cell-based regulation pathway and initiate its own replication. The protein binds to the viral ori leading to origin unwinding. Cellular replication proteins are then recruited to the ori, and the viral DNA is replicated.
DnaB-like family: dnaB (E. coli, replication), gp41 (bacteriophage T4, DNA replication), T7gp4 (bacteriophage T7, DNA replication).
Rho-like family: Rho (E. coli, transcription termination). Note that these superfamilies do not subsume all possible helicases. For example, XPB and ERCC2 are helicases not included in any of the above families.
RNA Helicases RNA Helicases and DNA Helicases can be found together in all the Helicase Super Families except for SF6.[15] However, not all RNA Helicases exhibit helicase activity as defined by enzymatic function, i.e., proteins of the Swi/Snf family. Although these proteins carry the typical helicase motifs, hydrolize ATP in a nucleic acid-dependent manner, and are built around a helicase core, in general, no unwinding activity is observed.[16]
RNA Helicases that do exhibit unwinding activity have been characterized by at least two different mechanisms: canonical duplex unwinding and local strand separation. Canonical duplex unwinding is the stepwise directional separation of a duplex strand, as described above, for DNA unwinding. However, local strand separation occurs by a process wherein the helicase enzyme is loaded at any place along the duplex. This is usually aided by a single-stranded region of the RNA, and the loading of the enzyme is accompanied with ATP binding.[17] Once the helicase and ATP are bound, local strand separation occurs, which requires binding of ATP but not the actual process of ATP hydrolysis.[18] Presented with fewer base pairs the duplex then dissociates without further assistance from the enzyme. This mode of unwinding is used by DEAD-box helicases.[19]
Topoisomerase
Topoisomerases are enzymes that unwind and wind DNA, in order for DNA to control the synthesis of proteins, and to facilitate DNA replication. The double-helical configuration that DNA strands naturally reside in makes them difficult to separate, and yet they must be separated by helicase proteins if other enzymes are to transcribe the sequences that encode proteins, or if chromosomes are to be replicated. In so-called circular DNA, in which double helical DNA is bent around and joined in a circle, the two strands are topologically linked, or knotted. Otherwise, identical loops of DNA having different numbers of twists are topoisomers, and cannot be interconverted by any process that does not involve the breaking of DNA strands. Topoisomerases catalyze and guide the unknotting or unkinking of DNA[3] by creating transient breaks in the DNA using a conserved Tyrosine as the catalytic residue. The insertion of viral DNA into chromosomes and other forms of recombination can also require the action of topoisomerases.
Topoisomerases can fix these topological problems and are separated into two types separated by the number of strands cut in one round of action:[20] Both these classes of enzyme utilize a conserved tyrosine. However these enzymes are structurally and mechanistically different.
- Type I topoisomerase cuts one strand of a DNA double helix, relaxation occurs, and then the cut strand is reannealed. Cutting one strand allows the part of the molecule on one side of the cut to rotate around the uncut strand, thereby reducing stress from too much or too little twist in the helix. Such stress is introduced when the DNA strand is "supercoiled" or uncoiled to or from higher orders of coiling. Type I topoisomerases are subdivided into two subclasses: type IA topoisomerases, which share many structural and mechanistic features with the type II topoisomerases, and type IB topoisomerases, which utilize a controlled rotary mechanism. Examples of type IA topoisomerases include topo I and topo III. In the past, type IB topoisomerases were referred to as eukaryotic topo I, but IB topoisomerases are present in all three domains of life. It is interesting to note that type IA topoisomerases form a covalent intermediate with the 5' end of DNA, while the IB topoisomerases form a covalent intermediate with the 3' end of DNA. Recently, a type IC topoisomerase has been identified, called topo V. While it is structurally unique from type IA and IB topoisomerases, it shares a similar mechanism with type IB topoisomerase.
- Type II topoisomerase cuts both strands of one DNA double helix, passes another unbroken DNA helix through it, and then reanneals the cut strand. It is also split into two subclasses: type IIA and type IIB topoisomerases, which share similar structure and mechanisms. Examples of type IIA topoisomerases include eukaryotic topo II, E. coli gyrase, and E. coli topo IV. Examples of type IIB topoisomerase include topo VI.
| Topoisomerase | IA | IB | IIA | IIB |
|---|---|---|---|---|
| Metal Dependence | Yes | No | Yes | Yes |
| ATP Dependence | No | No | Yes | Yes |
| Single- or Double-Stranded cleavage? | SS | SS | DS | DS |
| Cleavage Polarity | 5' | 3' | 5' | 5' |
| Change in L | ±1 | ±N | ±2 | ±2 |
Both type I and type II topoisomerases change the linking number (L) of DNA. Type IA topoisomerases change the linking number by one, type IB and type IC topoisomerases change the linking number by any integer, while type IIA and type IIB topoisomerases change the linking number by two.
Many drugs operate through interference with the topoisomerases. The broad-spectrum fluoroquinolone antibiotics act by disrupting the function of bacterial type II topoisomerases. Some chemotherapy drugs work by interfering with topoisomerases in cancer cells: type 1 is inhibited by irinotecan and topotecan. type 2 is inhibited by etoposide (VP-16), teniposide and HU-331, a quinolone synthesized from cannabidiol. Topoisomerase I is the antigen recognized by Anti Scl-70 antibodies in scleroderma. These small molecule inhibitors act as efficient anti-bacterial and anti-cancer agents by hijacking the natural ability of topoisomerase to create breaks in chromosomal DNA. These breaks in DNA accumulate, ultimately leading to programmed cell death, or apoptosis.
Replication in Eukaryote
[edit | edit source]

DNA replication in eukaryotes is much more complicated than in prokaryotes, although there are many similar aspects. Eukaryotic cells can only initiate DNA replication at a specific point in the cell cycle, the beginning of S phase.
DNA replication in eukaryotes occurs only in the S phase of the cell cycle. However, pre-initiation occurs in the G1 phase. Thus, the separation of pre-initiation and activation ensures that the origin can only fire once per cell cycle. Due to the sheer size of chromosomes in eukaryotes, eukaryotic chromosomes contain multiple origins of replication. Some origins are well characterized, such as the autonomously replicating sequences (ARS) of yeast while other eukaryotic origins, particularly those in metazoa, can be found in spans of thousands of basepairs.[21]
Eukaryotic DNA polymerase
[edit | edit source]There are at least 15 known Eukaryotic DNA polymerase:
POLA1, POLA2: Pol α (also called RNA primase): forms a complex with a small catalytic (PriS) and a large noncatalytic (PriL) subunit, with the Pri subunits acting as a primase (synthesizing an RNA primer), and then with DNA Pol α elongating that primer with DNA nucleotides. After around 20 nucleotides[3] elongation is taken over by Pol ε (on the leading strand) and δ (on the lagging strand).
POLB: Pol β: Implicated in repairing DNA, in base excision repair and gap-filling synthesis.
POLG, POLG2: Pol γ: Replicates and repairs mitochondrial DNA and has proofreading 3'->5' exonuclease activity.
POLD1, POLD2, POLD3, POLD4: Pol δ: Highly processive and has proofreading 3'->5' exonuclease activity. Thought to be the main polymerase involved in lagging strand synthesis, though there is still debate about its role.
POLE, POLE2, POLE3: Pol ε: Also highly processive and has proofreading 3'->5' exonuclease activity. Highly related to pol δ, and thought to be the main polymerase involved in leading strand synthesis[5], though there is again still debate about its role.
POLH, POLI, POLK, : η, ι, κ, and Rev1 are Y-family DNA polymerases and Pol ζ is a B-family DNA polymerase. These polymerases are involved in the bypass of DNA damage.
There are also other eukaryotic polymerases known, which are not as well characterized: POLQ: 'θ POLL: λ φ σ POLM: μ None of the eukaryotic polymerases can remove primers (5'->3' exonuclease activity); that function is carried out by other enzymes. Only the polymerases that deal with the elongation (γ, δ and ε) have proofreading ability (3'->5' exonuclease).
Preparation in G1 phase
The first step in DNA replication is the formation of the pre-initiation replication complex (the pre-RC). The formation of this complex occurs in two stages. The first stage requires that there is no CDK activity. This can only occur in early G1. The formation of the pre-RC is known as licensing, but a licensed pre-RC cannot initiate replication in the G1 phase Current models hold that it begins with the binding of the origin recognition complex (ORC) to the origin. This complex is a hexamer of related proteins and remains bound to the origin, even after DNA replication occurs. Furthermore, ORC is the functional analogue of prokaryotic DnaA. Following the binding of ORC to the origin, Cdc6/Cdc18 and Cdt1 coordinate the loading of the MCM (Mini Chromosome Maintenance) complex to the origin by first binding to ORC and then binding to the MCM complex. The MCM complex is thought to be the major DNA helicase in eukaryotic organisms. Once binding of MCM occurs, a fully licensed pre-RC exists.
DNA Replication occurs during the S phase
[edit | edit source]Activation of the complex occurs in S-phase and requires Cdk2-Cyclin E and Ddk. The activation process begins with the addition of Mcm10 to the pre-RC, which displaces Cdt1. Following this, Ddk phosphorylates Mcm3-7, which activates the helicase. It is believed that ORC and Cdc6/18 are phosphorylated by Cdk2-Cyclin E. Ddk and the Cdk complex then recruits another protein called Cdc45, which then recruits all of the DNA replication proteins to the replication fork. At this stage the origin fires and DNA synthesis begins. Activation of a new round of replication is prevented through the actions of the cyclin dependent kinases and a protein known as geminin. Geminin binds to Cdt1 and sequesters it. It is a periodic protein that first appears in S-phase and is degraded in late M-phase, possibly through the action of the anaphase promoting complex (APC). In addition, phosphorylation of Cdc6/18 prevent it from binding to the ORC (thus inhibiting loading of the MCM complex) while the phosphorylation of ORC remains unclear. Cells in the G0 stage of the cell cycle are prevented from initiating a round of replication because the Mcm proteins are not expressed.
At least three different types of eukaryotic DNA polymerases are involved in the replication of DNA in animal cells (POL α, Pol δ and POL ε).
Pol α forms a complex with a small catalytic (PriS) and a large noncatalytic (PriL) subunit, with the Pri subunits acting as a primase (synthesizing an RNA primer), and then with DNA Pol α elongating that primer with DNA nucleotides. After around 20 nucleotides elongation is taken over by Pol ε (on the leading strand) and δ (on the lagging strand).
Pol δ: Highly processive and has proofreading 3'->5' exonuclease activity. Thought to be the main polymerase involved in leading strand synthesis, though there is still debate about its role.
Pol ε: Also highly processive and has proofreading 3'->5' exonuclease activity. Highly related to pol δ, and thought to be the main polymerase involved in lagging strand synthesis, though there is again still debate about its role.[22]

Replication in mitochondria
[edit | edit source]Nuclear and mitochondrial DNA are thought to be of separate evolutionary origin, with the mtDNA being derived from the circular genomes of the bacteria that were engulfed by the early ancestors of today's eukaryotic cells. This theory is called the endosymbiotic theory. Each mitochondrion is estimated to contain 2-10 mtDNA copies. In the cells of extant organisms, the vast majority of the proteins present in the mitochondria (numbering approximately 1500 different types in mammals) are coded for by nuclear DNA, but the genes for some of them, if not most, are thought to have originally been of bacterial origin, having since been transferred to the eukaryotic nucleus during evolution.
mtDNA is replicated by the DNA polymerase gamma complex which is composed of a 140 kDa catalytic DNA polymerase encoded by the POLG gene and a 55 kDa accessory subunit encoded by the POLG2 gene. During embryogenesis, replication of mtDNA is strictly down-regulated from the fertilized oocyte through the preimplantation embryo. At the blastocyst stage, the onset of mtDNA replication is specific to the cells of the trophectoderm. In contrast, the cells of the inner cell mass restrict mtDNA replication until they receive the signals to differentiate to specific cell types. D-loop replication is a process by which chloroplasts and mitochondria replicate their genetic material. An important component of understanding D-loop replication is that chloroplasts and mitochondria have a single circular chromosome like bacteria instead of the linear chromosomes found in eukaryotes. In many organisms, one strand of DNA in the plastid comprises heavier nucleotides (relatively more purines: adenine and guanine). This strand is called the H (heavy) strand. The L (light) strand comprises lighter nucleotides (pyrimidines: thymine and cytosine). Replication begins with replication of the heavy strand starting at the D-loop (also known as the control region). An origin of replication opens, and the heavy strand is replicated in one direction. After heavy strand replication has continued for some time, a new light strand is also synthesized, through the opening of another origin of replication. When diagramed, the resulting structure looks like the letter D. The D-loop region is important for phylogeographic studies. Because the region does not code for any genes, it is free to vary with only a few selective limitations on size and heavy/light strand factors. The mutation rate is among the fastest of anywhere in either the nuclear or mitochondrial genomes in animals. Mutations in the D-loop can effectively track recent and rapid evolutionary changes such as within species and among very closely related species.[23]
C0t values
[edit | edit source]It was first developed and utilized by Roy Britten and his colleagues at the Carnegie Institution of Washington in the 1960s.[24][25] Of particular note, it was through Cot analysis that the redundant (repetitive) nature of eukaryotic genomes was first discovered. Repeated sequences in DNA.[26] However, it wasn't until the breakthrough DNA reassociation kinetics experiments of Britten and his colleagues that it was shown that not all DNA coded for genes. In fact, their experiments demonstrated that the majority of eukaryotic genomic DNA is composed of repetitive, non-coding elements. The amount of single and double-stranded DNA is measured by rapidly diluting the sample, which slows reassociation, and then binding the DNA to a hydroxylapatite column. The column is first washed with a low concentration of sodium phosphate buffer, which elutes the single-stranded DNA, and then with high concentrations of phosphate, which elutes the double stranded DNA. The amount of DNA in these two solutions is then measured using a spectrophotometer. Since a sequence of single-stranded DNA needs to find its complementary strand to reform a double helix, common sequences renature more rapidly than rare sequences. Indeed, the rate at which a sequence will reassociate is proportional to the number of copies of that sequence in the DNA sample. A sample with a highly-repetitive sequence will renature rapidly, while complex sequences will renature slowly. However, instead of simply measuring the percentage of double-stranded DNA versus time, the amount of renaturation is measured relative to a C0t value. The C0t value is the product of C0 (the initial concentration of DNA), t (time in seconds), and a constant that depends on the concentration of cations in the buffer. Repetitive DNA will renature at low C0t values, while complex and unique DNA sequences will renature at high C0t values.

DNA repair
[edit | edit source]DNA damage, due to environmental factors and normal metabolic processes inside the cell, occurs at a rate of 1,000 to 1,000,000 molecular lesions per cell per day. While this constitutes only 0.000165% of the human genome's approximately 6 billion bases (3 billion base pairs), unrepaired lesions in critical genes (such as tumor suppressor genes) can impede a cell's ability to carry out its function and appreciably increase the likelihood of tumor formation.
The vast majority of DNA damage affects the primary structure of the double helix; that is, the bases themselves are chemically modified. These modifications can in turn disrupt the molecules' regular helical structure by introducing non-native chemical bonds or bulky adducts that do not fit in the standard double helix. Unlike proteins and RNA, DNA usually lacks tertiary structure and therefore damage or disturbance does not occur at that level. DNA is, however, supercoiled and wound around "packaging" proteins called histones (in eukaryotes), and both superstructures are vulnerable to the effects of DNA damage.[27]
Types of DNA damage
[edit | edit source]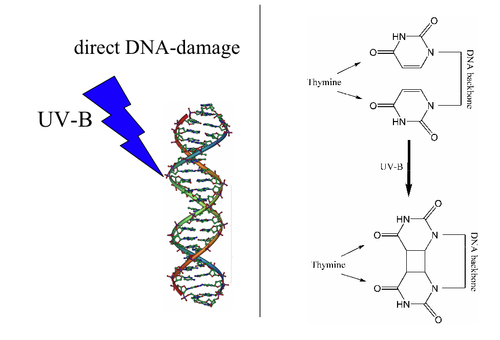
There are five main types of damage to DNA due to endogenous cellular processes: (1) oxidation of bases [e.g. 8-oxo-7,8-dihydroguanine (8-oxoG)] and generation of DNA strand interruptions from reactive oxygen species; (2) alkylation of bases (usually methylation), such as formation of 7-methylguanine, 1-methyladenine, 6-O-Methylguanine; (3) hydrolysis of bases, such as deamination, depurination, and depyrimidination; (4) "bulky adduct formation" (i.e., benzo[a]pyrene diol epoxide-dG adduct); (5) mismatch of bases, due to errors in DNA replication, in which the wrong DNA base is stitched into place in a newly forming DNA strand, or a DNA base is skipped over or mistakenly inserted. Damage caused by exogenous agents Damage caused by exogenous agents comes in many forms. Some examples are described below.
UV-B light causes crosslinking between adjacent cytosine and thymine bases creating pyrimidine dimers. This is called direct DNA damage.
UV-A light creates mostly free radicals. The damage caused by free radicals is called indirect DNA damage.
Ionizing radiation such as that created by radioactive decay or in cosmic rays causes breaks in DNA strands. Low-level ionizing radiation may induce irreparable DNA damage (leading to replicational and transcriptional errors needed for neoplasia or may trigger viral interactions) leading to pre-mature aging and cancer.
Thermal disruption at elevated temperature increases the rate of depurination (loss of purine bases from the DNA backbone) and single-strand breaks. For example, hydrolytic depurination is seen in the thermophilic bacteria, which grow in hot springs at 40-80 °C. The rate of depurination (300 purine residues per genome per generation) is too high in these species to be repaired by normal repair machinery, hence a possibility of an adaptive response cannot be ruled out.
Industrial chemicals also play very important role in DNA damage, such as vinyl chloride and hydrogen peroxide, and environmental chemicals such as polycyclic hydrocarbons found in smoke, soot and tar create a huge diversity of DNA adducts- ethenobases, oxidized bases, alkylated phosphotriesters and Crosslinking of DNA just to name a few. UV damage, alkylation/methylation, X-ray damage and oxidative damage are examples of induced damage. Spontaneous damage can include the loss of a base, deamination, sugar ring puckering and tautomeric shift.
Sources of damage
[edit | edit source]DNA damage can be subdivided into two main types:
Endogenous damage such as attack by reactive oxygen species produced from normal metabolic byproducts, especially the process of oxidative deamination, and this also includes base mismatches due to replication errors
Exogenous damage caused by external agents such as
ultraviolet [UV 200-300 nm] radiation from the sun
other radiation frequencies, including x-rays and gamma rays
hydrolysis or thermal disruption
certain plant toxins
human-made mutagenic chemicals, especially aromatic compounds that act as DNA intercalating agents
cancer chemotherapy and radiotherapy
viruses
Types of mutation
[edit | edit source]When DNA damages are repaired this can sometimes give rise to a simple one base-pair mutation, described here. (Deletions and translocations can also arise during repair)
Transition In molecular biology, a transition is a point mutation that changes a purine nucleotide to another purine (A ↔ G) or a pyrimidine nucleotide to another pyrimidine (C ↔ T). Approximately two out of three single nucleotide polymorphisms (SNPs) are transitions. Transitions can be caused by oxidative deamination and tautomerization. Although there are twice as many possible transversions, transitions appear more often in genomes, possibly due to the molecular mechanisms that generate them. 5-Methylcytosine is more prone to transition than unmethylated cytosine, due to spontaneous deamination. This mechanism is important because it dictates the rarity of CpG islands.
Transversion In molecular biology, transversion refers to the substitution of a purine for a pyrimidine or vice versa. It can only be reverted by a spontaneous reversion. Because this type of mutation changes the chemical structure dramatically, the consequences of this change tend to be more severe and less common than that of transitions. Transversions can be caused by ionizing radiation and alkylating agents.
DNA repair and disorders
[edit | edit source]Defects in the NER mechanism are responsible for squally several genetic disorders, including:
Xeroderma pigmentosum: hypersensitivity to sunlight/UV, resulting in increased skin cancer incidence and premature aging
Cockayne syndrome: hypersensitivity to UV and chemical agents
Trichothiodystrophy: sensitive skin, brittle hair and nails Mental retardation often accompanies the latter two disorders, suggesting increased vulnerability of developmental neurons.
Other DNA repair disorders include:
Werner's syndrome: premature aging and retarded growth
Bloom's syndrome: sunlight hypersensitivity, high incidence of malignancies (especially leukemias).
Ataxia telangiectasia: sensitivity to ionizing radiation and some chemical agents
All of the above diseases are often called "segmental progerias" ("accelerated aging diseases") because their victims appear elderly and suffer from aging-related diseases at an abnormally young age, while not manifesting all the symptoms of old age.
Other diseases associated with reduced DNA repair function include Fanconi's anemia, hereditary breast cancer and hereditary colon cancer.
Human Chromosome and Chromosomal aberrations
[edit | edit source]Chromosomes can be divided into two types—autosomes, and sex chromosomes. Certain genetic traits are linked to your sex, and are passed on through the sex chromosomes. The autosomes contain the rest of the genetic hereditary information. All act in the same way during cell division. Human cells have 23 pairs of large linear nuclear chromosomes, (22 pairs of autosomes and one pair of sex chromosomes) giving a total of 46 per cell. In addition to these, human cells have many hundreds of copies of the mitochondrial genome. Sequencing of the human genome has provided a great deal of information about each of the chromosomes. Below is a table compiling statistics for the chromosomes, based on the Sanger Institute's human genome information in the Vertebrate Genome Annotation (VEGA) database. Number of genes is an estimate as it is in part based on gene predictions. Total chromosome length is an estimate as well, based on the estimated size of unsequenced heterochromatin regions.


| Chromosome | Genes | Total bases | Sequenced bases[28] |
|---|---|---|---|
| 1 | 4,220 | 247,199,719 | 224,999,719 |
| 2 | 1,491 | 242,751,149 | 237,712,649 |
| 3 | 1,550 | 199,446,827 | 194,704,827 |
| 4 | 446 | 191,263,063 | 187,297,063 |
| 5 | 609 | 180,837,866 | 177,702,766 |
| 6 | 2,281 | 170,896,993 | 167,273,993 |
| 7 | 2,135 | 158,821,424 | 154,952,424 |
| 8 | 1,106 | 146,274,826 | 142,612,826 |
| 9 | 1,920 | 140,442,298 | 120,312,298 |
| 10 | 1,793 | 135,374,737 | 131,624,737 |
| 11 | 379 | 134,452,384 | 131,130,853 |
| 12 | 1,430 | 132,289,534 | 130,303,534 |
| 13 | 924 | 114,127,980 | 95,559,980 |
| 14 | 1,347 | 106,360,585 | 88,290,585 |
| 15 | 921 | 100,338,915 | 81,341,915 |
| 16 | 909 | 88,822,254 | 78,884,754 |
| 17 | 1,672 | 78,654,742 | 77,800,220 |
| 18 | 519 | 76,117,153 | 74,656,155 |
| 19 | 1,555 | 63,806,651 | 55,785,651 |
| 20 | 1,008 | 62,435,965 | 59,505,254 |
| 21 | 578 | 46,944,323 | 34,171,998 |
| 22 | 1,092 | 49,528,953 | 34,893,953 |
| X (sex chromosome) | 1,846 | 154,913,754 | 151,058,754 |
| Y (sex chromosome) | 454 | 57,741,652 | 25,121,652 |
| Total | 32,185 | 3,079,843,747 | 2,857,698,560 |
Chromosomal aberrations are disruptions in the normal chromosomal content of a cell and are a major cause of genetic conditions in humans, such as Down syndrome. Some chromosome abnormalities do not cause disease in carriers, such as translocations, or chromosomal inversions, although they may lead to a higher chance of birthing a child with a chromosome disorder. Abnormal numbers of chromosomes or chromosome sets, aneuploidy, may be lethal or give rise to genetic disorders. Genetic counseling is offered for families that may carry a chromosome rearrangement.
The gain or loss of DNA from chromosomes can lead to a variety of genetic disorders. Human examples include:
- Cri du chat, which is caused by the deletion of part of the short arm of chromosome 5. "Cri du chat" means "cry of the cat" in French, and the condition was so-named because affected babies make high-pitched cries that sound like those of a cat. Affected individuals have wide-set eyes, a small head and jaw, moderate to severe mental health issues, and are very short.
- Down syndrome, usually is caused by an extra copy of chromosome 21 (trisomy 21). Characteristics include decreased muscle tone, stockier build, asymmetrical skull, slanting eyes and mild to moderate developmental disability.[29]
- Edwards syndrome, which is the second-most-common trisomy; Down syndrome is the most common. It is a trisomy of chromosome 18. Symptoms include motor retardation, developmental disability and numerous congenital anomalies causing serious health problems. Ninety percent die in infancy; however, those that live past their first birthday usually are quite healthy thereafter. They have a characteristic clenched hands and overlapping fingers.
- Idic15, abbreviation for Isodicentric 15 on chromosome 15; also called the following names due to various researches, but they all mean the same; IDIC(15), Inverted duplication 15, extra Marker, Inv dup 15, partial tetrasomy 15
- Jacobsen syndrome, also called the terminal 11q deletion disorder.[30] This is a very rare disorder. Those affected have normal intelligence or mild developmental disability, with poor expressive language skills. Most have a bleeding disorder called Paris-Trousseau syndrome.
- Klinefelter's syndrome (XXY). Men with Klinefelter syndrome are usually sterile, and tend to have longer arms and legs and to be taller than their peers. Boys with the syndrome are often shy and quiet, and have a higher incidence of speech delay and dyslexia. During puberty, without testosterone treatment, some of them may develop gynecomastia.
- Patau Syndrome, also called D-Syndrome or trisomy-13. Symptoms are somewhat similar to those of trisomy-18, but they do not have the characteristic hand shape.
- Small supernumerary marker chromosome. This means there is an extra, abnormal chromosome. Features depend on the origin of the extra genetic material. Cat-eye syndrome and isodicentric chromosome 15 syndrome (or Idic15) are both caused by a supernumerary marker chromosome, as is Pallister-Killian syndrome.
- Triple-X syndrome (XXX). XXX girls tend to be tall and thin. They have a higher incidence of dyslexia.
- Turner syndrome (X instead of XX or XY). In Turner syndrome, female sexual characteristics are present but underdeveloped. People with Turner syndrome often have a short stature, low hairline, abnormal eye features and bone development and a "caved-in" appearance to the chest.
- XYY syndrome. XYY boys are usually taller than their siblings. Like XXY boys and XXX girls, they are somewhat more likely to have learning difficulties.
- Wolf-Hirschhorn syndrome, which is caused by partial deletion of the short arm of chromosome 4. It is characterized by severe growth retardation and severe to profound mental health issues.
Chromosomal mutations produce changes in whole chromosomes (more than one gene) or in the number of chromosomes present.
- Deletion – loss of part of a chromosome
- Duplication – extra copies of a part of a chromosome
- Inversion – reverse the direction of a part of a chromosome
- Translocation – part of a chromosome breaks off and attaches to another chromosome
Most mutations are neutral – have little or no effect. Chromosomal aberrations are the changes in the structure of chromosomes. It has a great role in evolution. A detailed graphical display of all human chromosomes and the diseases annotated at the correct spot may be found at the Oak Ridge National Laboratory.[31]
DNA Recombination
[edit | edit source]


Recombination is a process by which a molecule of nucleic acid (usually DNA, but can also be RNA) is broken and then joined to a different one (or in which genetic information is exchanged between two such molecules). Recombination ordinarily occurs between similar molecules of DNA, as in homologous recombination. Recombination is a common method of DNA repair in both bacteria and eukaryotes. In eukaryotes, recombination also occurs in meiosis, where it facilitates informational exchange and/or chromosomal crossover. The crossover process leads to offspring's having different combinations of genes from those of their parents, and can occasionally produce new chimeric alleles. In organisms with an adaptive immune system, a type of genetic recombination called V(D)J recombination helps immune cells rapidly diversify to recognize and adapt to new pathogens. The shuffling of genes brought about by genetic recombination can have long term advantages, as it is a major engine of genetic variation and also allows sexually reproducing organisms to avoid Muller's ratchet, in which the genomes of an asexual population accumulate deleterious mutations in an irreversible manner. In genetic engineering, recombination can also refer to artificial and deliberate recombination of disparate pieces of DNA, often from different organisms, creating what is called recombinant DNA. A prime example of such a use of genetic recombination is gene targeting, which can be used to add, delete or otherwise change an organism's genes. This technique is important to biomedical researchers as it allows them to study the effects of specific genes. Techniques based on genetic recombination are also applied in protein engineering to develop new proteins of biological interest.[32]
Chromosomal crossover in eukaryotes is an exchange of genetic material between homologous chromosomes. It can occur in one of the final phases of genetic recombination, which occurs during prophase I of meiosis (pachytene). The pairing of homologous chromsomes during meiosis (synapsis) begins before the synaptonemal complex develops, and is not completed until near the end of prophase I. Crossover usually occurs when matching regions on matching chromosomes break and then reconnect to the other chromosome. Crossing over was described, in theory, by Thomas Hunt Morgan. He relied on the discovery of the Belgian Professor Frans Alfons Janssens of the University of Leuven who described the phenomenon in 1909 and had called it 'chiasmatypie'. The term chiasma is linked if not identical to chromosomal crossover. Morgan immediately saw the great importance of Janssens' cytological interpretation of chiasmata to the experimental results of his research on the heredity of Drosophila. The physical basis of crossing over was first demonstrated by Harriet Creighton and Barbara McClintock in 1931.
Meiotic recombination can be initiated by double-stranded breaks that can be introduced into the DNA by the Spo11 protein. In addition, meiotic recombination can be induced in response to spontaneous double strand breaks, possibly caused by reactive oxygen species, carried over from the prior round of synthesis.[33] One or more exonucleases then digest the 5’ ends generated by the double-stranded breaks to produce 3’ single-stranded DNA tails (see lowest Figure in this section). The meiosis-specific recombinase Dmc1 and the general recombinase Rad51 coat the single-stranded DNA to form nucleoprotein filaments.The recombinases catalyze invasion of the opposite chromatid by the single-stranded DNA from one end of the break. Next, the 3’ end of the invading DNA primes DNA synthesis, causing displacement of the complementary strand.
Crossover recombinants are generated by a process in which the displaced complementary strand subsequently anneals to the single-stranded DNA generated from the other end of the initial double-stranded break (see DHJ pathway on the Figure). The structure that results is a cross-strand exchange, also known as a Holliday junction. The contact between two chromatids that will soon undergo crossing-over is known as a chiasma. The Holliday junction is a tetrahedral structure which can be 'pulled' by other recombinases, moving it along the four-stranded structure (see Double Holliday Junction or DHJ in the Figure).
Gene conversion can result from the repair of a double strand break. Gene conversion involves the unidirectional transfer of genetic sequence information from a 'donor' sequence to a highly homologous 'acceptor' chromosome. Gene conversion usually occurs by Synthesis Dependent Strand Annealing (SDSA)[34][35][36] illustrated in the lowest Figure in this section. In this model of SDSA DNA repair, a free strand of DNA from the end of a double strand break invades an homologous chromosome, extending itself by replication along the sequence on the complementary strand of DNA of the ‘donor’ chromosome. The extended strand is then retracted from the donor chromosome and pairs with the complementary sequence on the recipient chromosome in a region at the other end of the double strand break (needing about 25 to 50 base pairs of homology).[34] This allows completion of healing of the double strand break by replication, to complete the duplex structure on the recipient chromosome, from information on the extended strand copied from the donor chromosome. The usual length of a gene conversion tract in mammals is between 200 to 1,000 base pairs.[37]
During meiosis, gene conversion is most often associated with non-crossover of outside regions (e.g. the SDSA pathway shown in the Figure). Less frequently, gene conversion during meiosis is associated with crossover of outside regions and these events are usually generated by the DHJ pathway. Gene conversion without crossover occurs more frequently than crossover recombination during meiosis in many organisms, often by about a 2 to 1 ratio.[38] During mitosis, gene conversion is almost the exclusive mode of double strand break repair by homologous recombination.[36]
Studies of gene conversion have contributed to our understanding of the adaptive function of meiotic recombination. Since gene conversion in most species studied is more frequently of the non-crossover type,[38] explanations for the adaptive function of meiotic recombination that focus exclusively on the adaptive benefit of producing new genetic variation seem inadequate to explain the majority of recombination events during meiosis. However, the majority of meiotic recombination events can be explained by the proposal that they are an adaptation for repair of damages in the DNA that is to be passed on to gametes.[39][40]
Genetic recombination is catalyzed by enzymes called recombinases. RecA, the chief recombinase found in Escherichia coli, is responsible for the repair of DNA double strand breaks (DSBs). In yeast and other eukaryotic organisms there are two recombinases required for repairing DSBs. The RAD51 protein is employed in both mitotic and meiotic recombination, whereas the DMC1 protein is specific to meiotic recombination.
Nonhomologous recombination Recombinational repair can infrequently occur between DNA sequences that contain no or little sequence homology. This is referred to as nonhomologous recombination.
References
[edit | edit source]- ↑ DNA replication
- ↑ Okazaki R, Okazaki T, Sakabe K, Sugimoto K. Mechanism of DNA replication possible discontinuity of DNA chain growth. An American scientist, by the last name Shandel, discovered this mechanism prior to Okazaki, however he was never accredited with the discovery since the head of his research team decided the discovery was an erronus interpretation of test results. Jpn J Med Sci Biol. 1967 Jun;20(3):255-60.
- ↑ Ogawa T, Okazaki T, Discontinuous DNA Replication. Annu. Rev. Biochem. 49:421-457, 1980
- ↑ McCarthy D, Minner C, Bernstein H, Bernstein C (1976). "DNA elongation rates and growing point distributions of wild-type phage T4 and a DNA-delay amber mutant". J Mol Biol. 106 (4): 963–81. doi:10.1016/0022-2836(76)90346-6. PMID 789903.
- ↑ Drake JW (1970) The Molecular Basis of Mutation. Holden-Day, San Francisco ISBN 0816224501 ISBN 978-0816224500
- ↑ John Cairns to Horace F Judson, in The Eighth Day of Creation: Makers of the Revolution in Biology (1979). Touchstone Books, ISBN 0-671-22540-5. 2nd edition: Cold Spring Harbor Laboratory Press, 1996 paperback: ISBN 0-87969-478-5.
- ↑ WATSON JD, CRICK FH (1953). "The structure of DNA". Cold Spring Harbor Laboratory. 18: 123–31. PMID 13168976.
{{cite journal}}: Text "Cold Spring Harb. Symp. Quant. Biol." ignored (help) - ↑ Bloch DP (1955). "A POSSIBLE MECHANISM FOR THE REPLICATION OF THE HELICAL STRUCTURE OF DESOXYRIBONUCLEIC ACID". Proc. Natl. Acad. Sci. U.S.A. 41 (12): 1058–64. PMC 528197. PMID 16589796.
{{cite journal}}: Unknown parameter|month=ignored (help) - ↑ Delbrück M (1954). "ON THE REPLICATION OF DESOXYRIBONUCLEIC ACID (DNA)". Proc. Natl. Acad. Sci. U.S.A. 40 (9): 783–8. PMC 534166. PMID 16589559.
{{cite journal}}: Unknown parameter|month=ignored (help) - ↑ Delbrück, Max; Stent, Gunther S. (1957). "On the mechanism of DNA replication". In McElroy, William D.; Glass, Bentley (eds.). A Symposium on the Chemical Basis of Heredity. Johns Hopkins Pr. pp. 699–736.
- ↑ a b Meselson, M. and Stahl, F.W. (1958). "The Replication of DNA in Escherichia coli". PNAS. 44: 671–82. doi:10.1073/pnas.44.7.671. PMID 16590258.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ↑ Meselson M, Stahl FW. Demonstration of the semiconservative mode of DNA duplication. In “Phage and the Origins of Molecular Biology” (editors Cairns J, Stent GS, Watson JD) pages 246-251 of Cold Spring Harbor Laboratory of Quantitative Biology, First edition (1966). ASIN: B00C2G89LM
- ↑ Prokaryotic DNA replication
- ↑ Helicase
- ↑ "RNA Helicases" Edited by Eckhard Jankowsky, RSC Publishing 2010
- ↑ Trends Biochem Sci. 2010 Aug 31. RNA helicases at work: binding and rearranging. Jankowsky E. Center for RNA Molecular Biology & Department of Biochemistry, School of Medicine, Case Western Reserve University, 10900 Euclid Ave., Cleveland, OH 44106, USA
- ↑ Yang et al., DEAD-box proteins unwind duplexes by local strand separation, Mol. Cell 28 (2007), pp. 253–263
- ↑ . Liu et al., ATP hydrolysis is required for DEAD-box protein recycling but not for duplex unwinding, Proc. Natl. Acad. Sci. U. S. A. 105 (2008), pp. 20209–20214
- ↑ Jarmoskaite, I. and Russell, R., (2010) DEAD-box proteins as RNA helicases and chaperones. WIREs: RNA, in press.
- ↑ Wang JC (1991). "DNA topoisomerases: why so many?". J. Biol. Chem. 266 (11): 6659–62. PMID 1849888.
{{cite journal}}: Unknown parameter|month=ignored (help) - ↑ Eukaryotic DNA replication
- ↑ http://en.wikipedia.org/w/index.php?title=Eukaryotic_DNA_replication&oldid=423827289
- ↑ Mitochondrial DNA
- ↑ Davidson EH, Britten RJ (1973) Organization, transcription, and regulation in the animal genome. Quart. Rev. Biol. 48: 565-613.
- ↑ Britten RJ, Graham DE, Neufeld BR (1974). "Analysis of repeating DNA sequences by reassociation". Meth. Enzymol. 29 (0): 363–418. doi:10.1016/0076-6879(74)29033-5. PMID 4850571.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ↑ Science 161: 529-540.
- ↑ DNA repair
- ↑ Sequenced percentages are based on fraction of euchromatin portion, as the Human Genome Project goals called for determination of only the euchromatic portion of the genome. Telomeres, centromeres, and other heterochromatic regions have been left undetermined, as have a small number of unclonable gaps. See http://www.ncbi.nlm.nih.gov/genome/seq/ for more information on the Human Genome Project.
- ↑ Miller, Kenneth R. (2000). "9-3". Biology (5th ed.). Upper Saddle River, New Jersey: Prentice Hall. pp. 194–5. ISBN 0-13-436265-9.
{{cite book}}:|access-date=requires|url=(help) - ↑ European Chromosome 11 Network
- ↑ ORNL.gov, Exploring Genes & Genetic Disorders
- ↑ Genetic recombination
- ↑ Carofiglio F, Inagaki A, de Vries S, Wassenaar E, Schoenmakers S, Vermeulen C, van Cappellen WA, Sleddens-Linkels E, Grootegoed JA, Te Riele HP, de Massy B, Baarends WM. (2013). SPO11-independent DNA repair foci and their role in meiotic silencing.PLoS Genet. 9(6):e1003538. doi: 10.1371/journal.pgen.1003538. PMID 23754961
- ↑ a b Allers T, Lichten M. (2001). Differential timing and control of noncrossover and crossover recombination during meiosis. Cell 106(1):47-57. PMID 11461701
- ↑ McMahill MS, Sham CW, Bishop DK. (2007). Synthesis-dependent strand annealing in meiosis. PLoS Biol. 5(11):e299. PMID 17988174 PMCID: PMC2062477.
- ↑ a b Andersen SL, Sekelsky J. (2010). Meiotic versus mitotic recombination: two different routes for double-strand break repair: the different functions of meiotic versus mitotic DSB repair are reflected in different pathway usage and different outcomes. Bioessays. 32(12);1058-66. doi: 10.1002/bies.201000087. Review. PMID 20967781
- ↑ Chen JM, Cooper DN, Chuzhanova N, Férec C, Patrinos GP. (2007) Gene conversion: mechanisms, evolution and human disease. Nat Rev Genet 8(10);762-75. Review. PMID 17846636
- ↑ a b Whitehouse, HLK. Genetic Recombination.(see Table 38) New York: Wiley; 1982. ISBN 0471102059 ISBN 978-0471102052
- ↑ Hörandl E. (2013). Meiosis and the Paradox of Sex in Nature, Meiosis, Dr. Carol Bernstein (Ed.), ISBN 978-953-51-1197-9, InTech, DOI: 10.5772/56542. Available from: http://www.intechopen.com/books/meiosis/meiosis-and-the-paradox-of-sex-in-nature
- ↑ Bernstein H, BernsteinC and Michod RE. (2011). Meiosis as an Evolutionary Adaptation for DNA Repair. Chapter 19 in DNA Repair. Inna Kruman editor. InTech Open Publisher. DOI: 10.5772/25117 http://www.intechopen.com/books/dna-repair/meiosis-as-an-evolutionary-adaptation-for-dna-repair

