Data Mining Algorithms In R/Clustering/Partitioning Around Medoids (PAM)
Introduction
[edit | edit source]Clustering is an unsupervised machine learning algorithm that groups entities, from a dataset, that have high degree of similarity in the same cluster. Nowadays lots of areas are using these kinds of algorithms to separate datasets into groups in an automated way, and still have a good quality result.
The clustering process is not a universal process because there are many groups of datasets, for some of them the kind of metric used is relevant, for others the entities that represent each cluster are more interesting. Like dataset groups there are many clustering algorithms each one tries to take advantage of data type, so this way each one of them is more suited for a specific kind of data.
This section will explain a little more about the Partitioning Around Medoids (PAM) Algorithm, showing how the algorithm works, what are its parameters and what they mean, an example of a dataset, how to execute the algorithm, and the result of that execution with the dataset as input.
The Partitioning Around Medoids (PAM) Algorithm
[edit | edit source]Algorithm
[edit | edit source]The PAM algorithm was developed by Leonard Kaufman and Peter J. Rousseeuw, and this algorithm is very similar to K-means, mostly because both are partitional algorithms, in other words, both break the dataset into groups (clusters), and both work by trying to minimize the error, but PAM works with Medoids, that are an entity of the dataset that represent the group in which it is inserted, and K-means works with Centroids, that are artificially created entity that represent its cluster.
The PAM algorithm partitions the dataset of n objects into k clusters, where both the dataset and the number k is an input of the algorithm. This algorithm works with a matrix of dissimilarity, whose goal is to minimize the overall dissimilarity between the representants of each cluster and its members. The algorithm uses the following model to solve the problem:
Subject to:
Where F(x) is the main function to minimize, d(i,j) is the dissimilarity measurement between the entities i and j, and zij is a variable that ensures that only the dissimilarity between entities from the same cluster will be computed in the main function. The others expressions are constraints that have the following functions: (1.) ensures that every single entity is assigned to one cluster and only one cluster, (2.) ensures that the entity is assigned to its medoid that represents the cluster, (3.) ensures that there are exactly k clusters and (4.) lets the decision variables assume just the values of 0 or 1.
The PAM algorithm can work over two kinds of input, the first is the matrix representing every entity and the values of its variables, and the second is the dissimilarity matrix directly, in the latter the user can provide the dissimilarity directly as an input to the algorithm, instead of the data matrix representing the entities. Either way the algorithm reaches a solution to the problem, in a general analysis the algorithm proceeds this way:
Build phase:
- 1. Choose k entities to become the medoids, or in case these entities were provided use them as the medoids;
- 2. Calculate the dissimilarity matrix if it was not informed;
- 3. Assign every entity to its closest medoid;
Swap phase:
- 4. For each cluster search if any of the entities of the cluster lower the average dissimilarity coefficient, if it does select the entity that lowers this coefficient the most as the medoid for this cluster;
- 5. If at least one medoid has changed go to (3), else end the algorithm.
Implementation
[edit | edit source]THIS IS NOT DESCRIBING THE "PAM" ALGORITHM. This is a k-means variant using medoids, but not PAM with its charateristic BUILD and SWAP phases.
The pseudocode of PAM algorithm is shown below:
Algorithm 1: PAM Algorithm Input: E = {e1,e2,...en} (dataset to be clustered or matrix of dissimilarity)
- k (number of clusters)
- metric (kind of metric to use on dissimilarity matrix)
- diss (flag indicating that E is the matrix of dissimilarity or not)
Output: M = {m1,m2,...,mk} (vector of clusters medoids)
- L = {l(e) | e = 1,2,...,n} (set of cluster labels of E)
foreach mi € M do
- mi ← ej € E; (e.g. random selection)
end if diss ≠ true
- Dissimilarity ← CalculateDissimilarityMatrix(E, metric);
else
- Dissimilarity ← E;
end repeat
- foreach ei € E do
- l(ei) ← argminDissimilarity(ei, Dissimilarity, M);
- end
- changed ← false;
- foreach mi € M do
- Mtmp ← SelectBestClusterMedoids(E, Dissimilarity, L);
- end
- if Mtmp ≠ M
- M ← Mtmp;
- changed ← true;
- end
until changed = true;
In the R programming language, the PAM algorithm is available in the cluster package and can be called by the following command:
pam(x, k, diss, metric, medoids, stand, cluster.only, do.swap, keep.diss, keep.data, trace.lev)
Where the parameters are:
x: numerical data matrix representing the dataset entities, or can be the dissimilarity matrix, it depends on the value of the diss parameter. In case x is a data matrix each row is an entity and each column is an variable, and in this case missing values are allowed as long as every pair of entities has at least one case not missing. In case x is a dissimilarity matrix it is not allowed to have missing values.
k: number of clusters that the dataset will be partitioned where 0 < k < n, where n is the number of entities.
diss: logical flag, if it is TRUE x is used as the dissimilarity matrix, if it is FALSE, then x will be considered as a data matrix.
metric: an string specifying each of the two metrics will be used to calculate the dissimilarity matrix, the metric variable can be “euclidean” to use the Euclidean distance, or can be “manhattan” to use the Manhattan distance.
stand: logical flag, if it is TRUE then the measurements in x will be standardized before calculating the dissimilarities. Measurements are standardized for each column, by subtracting the column's mean value and dividing by the variable's mean absolute deviation. If x is a dissimilarity matrix then this parameter is ignored.
cluster.only: logical flag, if it is TRUE, only the clustering will be computed and returned.
do.swap: logical flag, indicates if the swap phase should happen (TRUE) or not (FALSE).
keep.diss: logical flag indicating if the dissimilarities should (TRUE) or not (FALSE) be kept in the result.
keep.data: logical flag indicating if the input data x should (TRUE) or not (FALSE) be kept in the result.
trace.lev: an numeric parameters specifying a trace level for printing diagnostics during the build and swap phase of the algorithm. Default 0 does not print anything.
The PAM algorithm return a pam object that contains the information about the result of the execution of the algorithm.
Visualization
[edit | edit source]In R there are two ways of seeing the result of the PAM algorithm, the first one is to print the object that the algorithm returns, and the second one is to plot the data from the object creating a graphic of the result. The first way of visualizing the information is a bit more complicated to understand but it gives a more complete and accurate information, but the second way is a lot more easy to understand and lets the user have a better view of the information and to add information that would be relevant for him.
To view the data of the result of the execution of PAM algorithm in a textual way there are two ways one more simple that gives a more summarized information about the object, and another one that gives you a more complete information about it. In the two commands listed below the first one prints the information in a summarized way, while the second one prints it in a more complete way.
print (result) summary (result)
The other way of visualizing the data from the result of the execution of the algorithm is using graphics and that can be done by using the following command:
plot (result)
Example: To show an example of use of the algorithm and a result from its execution a simple dataset with few entities and few dimension was used, as it is shown in the following table:
Table 1: Simple dataset
| Object | Attribute x | Attribute y |
|---|---|---|
| 1 | 1 | 1 |
| 2 | 2 | 3 |
| 3 | 1 | 2 |
| 4 | 2 | 2 |
| 5 | 10 | 4 |
| 6 | 11 | 5 |
| 7 | 10 | 6 |
| 8 | 12 | 5 |
| 9 | 11 | 6 |
As we can see the data is separated in two clusters, so we will use an k = 2. The PAM algorithm can be executed as follows:
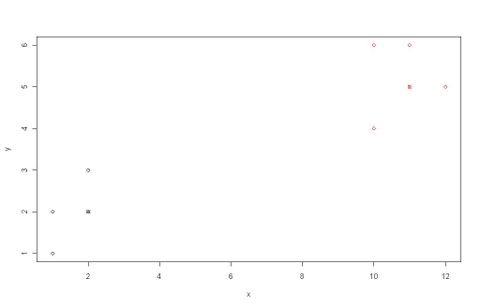
#load the table from a file x <- read.table(“table.txt”) #execute the pam algorithm with the dataset created for the example result <- pam(x, 2, FALSE, "euclidean") #print the results data in the screen summary(result) #plot a graphic showing the clusters and the medoids of each cluster plot(result$data, col = result$clustering) points(result$medoids, col = 1:2, pch = 4)
Printing the result form the execution gives you:
Medoids:
ID x y
4 4 2 2
6 6 11 5
Clustering vector:
1 2 3 4 5 6 7 8 9
1 1 1 1 2 2 2 2 2
Objective function:
build swap
1.255618 0.915849
Numerical information per cluster:
size max_diss av_diss diameter separation
[1,] 4 1.414214 0.8535534 2.236068 8.062258
[2,] 5 1.414214 0.9656854 2.236068 8.062258
Isolated clusters:
L-clusters: character(0)
L*-clusters: [1] 1 2
Silhouette plot information:
cluster neighbor sil_width
3 1 2 0.8898942
4 1 2 0.8788422
1 1 2 0.8549629
2 1 2 0.8297000
6 2 1 0.8790384
9 2 1 0.8631441
8 2 1 0.8425790
7 2 1 0.8232848
5 2 1 0.7747713
Average silhouette width per cluster:
[1] 0.8633498 0.8365635
Average silhouette width of total data set:
[1] 0.8484685
36 dissimilarities, summarized :
Min. 1st Qu. Median Mean 3rd Qu. Max.
1.0000 1.4142 8.3951 6.1559 9.9362 11.7050
Metric : euclidean
Number of objects : 9
Available components:
[1] "medoids" "id.med" "clustering" "objective" "isolation" "clusinfo"
"silinfo" "diss" "call"
[10] "data"
While plotting gives you:
Case Study
[edit | edit source]In this section we will see a case study using PAM.
Scenario
[edit | edit source]In this case study a part of the database iris available in the R package datasets was used. This famous (Fisher’s or Anderson’s) iris data set gives the measurements in centimeters of the variables sepal length and width and petal length and width, respectively, for 50 flowers from each of 3 species of iris. The species are Iris setosa, versicolor, and virginica. Using the data that this dataset provides us, it is natural to think of verifying if the flowers of each one of three species of iris are really similar to the others from the same species, so in this case study length and width of both petal and sepal will be used to cluster the dataset into 3 groups and then verify if the clusters really match with the flowers species.
The dataset that was used into this case study consist of the following columns:
- Flower: An id of the flower;
- Sepal.Length: A numeric value of the length of the sepal in centimeters;
- Sepal.Width: A numeric value of the width of the sepal in centimeters;
- Petal.Length: A numeric value of the length of the petal in centimeters;
- Petal.Width: A numeric value of the width of the petal in centimeters;
- Species: A text identifying the species of the flower.
Input Data
[edit | edit source]The input data is a table consisting of 50% (75 entities) of the original iris dataset that have 150 flowers and 5 attributes each. So the dataset used in this case study is represented by the following table:
Table 2: Sample from iris dataset
| Flower | Sepal.Length | Sepal.Width | Petal.Length | Petal.Width | Species |
|---|---|---|---|---|---|
| 1 | 5.1 | 3.5 | 1.4 | 0.2 | setosa |
| 2 | 4.9 | 3.0 | 1.4 | 0.2 | setosa |
| 3 | 4.7 | 3.2 | 1.3 | 0.2 | setosa |
| 4 | 4.6 | 3.1 | 1.5 | 0.2 | setosa |
| 5 | 5.0 | 3.6 | 1.4 | 0.2 | setosa |
| 6 | 5.4 | 3.9 | 1.7 | 0.4 | setosa |
| 7 | 4.6 | 3.4 | 1.4 | 0.3 | setosa |
| 8 | 5.0 | 3.4 | 1.5 | 0.2 | setosa |
| 9 | 4.4 | 2.9 | 1.4 | 0.2 | setosa |
| 10 | 4.9 | 3.1 | 1.5 | 0.1 | setosa |
| 11 | 5.4 | 3.7 | 1.5 | 0.2 | setosa |
| 12 | 4.8 | 3.4 | 1.6 | 0.2 | setosa |
| 13 | 4.8 | 3.0 | 1.4 | 0.1 | setosa |
| 14 | 4.3 | 3.0 | 1.1 | 0.1 | setosa |
| 15 | 5.8 | 4.0 | 1.2 | 0.2 | setosa |
| 16 | 5.7 | 4.4 | 1.5 | 0.4 | setosa |
| 17 | 5.4 | 3.9 | 1.3 | 0.4 | setosa |
| 18 | 5.1 | 3.5 | 1.4 | 0.3 | setosa |
| 19 | 5.7 | 3.8 | 1.7 | 0.3 | setosa |
| 20 | 5.1 | 3.8 | 1.5 | 0.3 | setosa |
| 21 | 5.4 | 3.4 | 1.7 | 0.2 | setosa |
| 22 | 5.1 | 3.7 | 1.5 | 0.4 | setosa |
| 23 | 4.6 | 3.6 | 1.0 | 0.2 | setosa |
| 24 | 5.1 | 3.3 | 1.7 | 0.5 | setosa |
| 25 | 4.8 | 3.4 | 1.9 | 0.2 | setosa |
| 51 | 7.0 | 3.2 | 4.7 | 1.4 | versicolor |
| 52 | 6.4 | 3.2 | 4.5 | 1.5 | versicolor |
| 53 | 6.9 | 3.1 | 4.9 | 1.5 | versicolor |
| 54 | 5.5 | 2.3 | 4.0 | 1.3 | versicolor |
| 55 | 6.5 | 2.8 | 4.6 | 1.5 | versicolor |
| 56 | 5.7 | 2.8 | 4.5 | 1.3 | versicolor |
| 57 | 6.3 | 3.3 | 4.7 | 1.6 | versicolor |
| 58 | 4.9 | 2.4 | 3.3 | 1.0 | versicolor |
| 59 | 6.6 | 2.9 | 4.6 | 1.3 | versicolor |
| 60 | 5.2 | 2.7 | 3.9 | 1.4 | versicolor |
| 61 | 5.0 | 2.0 | 3.5 | 1.0 | versicolor |
| 62 | 5.9 | 3.0 | 4.2 | 1.5 | versicolor |
| 63 | 6.0 | 2.2 | 4.0 | 1.0 | versicolor |
| 64 | 6.1 | 2.9 | 4.7 | 1.4 | versicolor |
| 65 | 5.6 | 2.9 | 3.6 | 1.3 | versicolor |
| 66 | 6.7 | 3.1 | 4.4 | 1.4 | versicolor |
| 67 | 5.6 | 3.0 | 4.5 | 1.5 | versicolor |
| 68 | 5.8 | 2.7 | 4.1 | 1.0 | versicolor |
| 69 | 6.2 | 2.2 | 4.5 | 1.5 | versicolor |
| 70 | 5.6 | 2.5 | 3.9 | 1.1 | versicolor |
| 71 | 5.9 | 3.2 | 4.8 | 1.8 | versicolor |
| 72 | 6.1 | 2.8 | 4.0 | 1.3 | versicolor |
| 73 | 6.3 | 2.5 | 4.9 | 1.5 | versicolor |
| 74 | 6.1 | 2.8 | 4.7 | 1.2 | versicolor |
| 75 | 6.4 | 2.9 | 4.3 | 1.3 | versicolor |
| 101 | 6.3 | 3.3 | 6.0 | 2.5 | virginica |
| 102 | 5.8 | 2.7 | 5.1 | 1.9 | virginica |
| 103 | 7.1 | 3.0 | 5.9 | 2.1 | virginica |
| 104 | 6.3 | 2.9 | 5.6 | 1.8 | virginica |
| 105 | 6.5 | 3.0 | 5.8 | 2.2 | virginica |
| 106 | 7.6 | 3.0 | 6.6 | 2.1 | virginica |
| 107 | 4.9 | 2.5 | 4.5 | 1.7 | virginica |
| 108 | 7.3 | 2.9 | 6.3 | 1.8 | virginica |
| 109 | 6.7 | 2.5 | 5.8 | 1.8 | virginica |
| 110 | 7.2 | 3.6 | 6.1 | 2.5 | virginica |
| 111 | 6.5 | 3.2 | 5.1 | 2.0 | virginica |
| 112 | 6.4 | 2.7 | 5.3 | 1.9 | virginica |
| 113 | 6.8 | 3.0 | 5.5 | 2.1 | virginica |
| 114 | 5.7 | 2.5 | 5.0 | 2.0 | virginica |
| 115 | 5.8 | 2.8 | 5.1 | 2.4 | virginica |
| 116 | 6.4 | 3.2 | 5.3 | 2.3 | virginica |
| 117 | 6.5 | 3.0 | 5.5 | 1.8 | virginica |
| 118 | 7.7 | 3.8 | 6.7 | 2.2 | virginica |
| 119 | 7.7 | 2.6 | 6.9 | 2.3 | virginica |
| 120 | 6.0 | 2.2 | 5.0 | 1.5 | virginica |
| 121 | 6.9 | 3.2 | 5.7 | 2.3 | virginica |
| 122 | 5.6 | 2.8 | 4.9 | 2.0 | virginica |
| 123 | 7.7 | 2.8 | 6.7 | 2.0 | virginica |
| 124 | 6.3 | 2.7 | 4.9 | 1.8 | virginica |
| 125 | 6.7 | 3.3 | 5.7 | 2.1 | virginica |
Execution
[edit | edit source]The process was done as follows:
#import data data <- read.table(“sampleiris.txt”) #execution result <- pam(data[1:4], 3, FALSE, “euclidean”) #print results summary(result) #plot clusters plot (data, col = result$clustering) #add the medoids to the plot points(result$medoids, col = 1:3, pch = 4)
Output
[edit | edit source]The following data was printed as result of the execution:
Medoids:
ID Sepal.Length Sepal.Width Petal.Length Petal.Width
8 8 5.0 3.4 1.5 0.2
64 39 6.1 2.9 4.7 1.4
103 53 7.1 3.0 5.9 2.1
Clustering vector:
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22
23 24 25 51 52 53 54 55 56
1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
1 1 1 2 2 2 2 2 2
57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 101 102 103
104 105 106 107 108 109 110 111 112
2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 3 2 3
3 3 3 2 3 3 3 2 2
113 114 115 116 117 118 119 120 121 122 123 124 125
3 2 2 3 3 3 3 2 3 2 3 2 3
Objective function:
build swap
0.7148339 0.6990539
Numerical information per cluster:
size max_diss av_diss diameter separation
[1,] 25 1.236932 0.5137400 2.042058 1.9000000
[2,] 34 1.951922 0.8085343 2.727636 0.3741657
[3,] 16 1.284523 0.7559609 2.147091 0.3741657
Isolated clusters:
L-clusters: [1] 1
L*-clusters: character(0)
Silhouette plot information:
cluster neighbor sil_width
1 1 2 0.84941732
5 1 2 0.84830238
8 1 2 0.84812593
18 1 2 0.84784555
12 1 2 0.83221128
22 1 2 0.82890349
20 1 2 0.82456328
3 1 2 0.82337894
7 1 2 0.81910409
10 1 2 0.81662688
11 1 2 0.80769429
2 1 2 0.80592613
13 1 2 0.80278163
4 1 2 0.79810574
23 1 2 0.79482977
24 1 2 0.78999596
17 1 2 0.78539723
21 1 2 0.78454015
25 1 2 0.77452963
6 1 2 0.75995941
9 1 2 0.74605493
14 1 2 0.74277337
19 1 2 0.72082914
15 1 2 0.71581750
16 1 2 0.66155611
68 2 3 0.60036142
56 2 3 0.59753885
62 2 3 0.59698924
72 2 3 0.59691421
70 2 3 0.59514179
54 2 3 0.58507022
67 2 3 0.56989428
60 2 1 0.56350914
63 2 3 0.55592514
75 2 3 0.54720666
74 2 3 0.53971473
64 2 3 0.53757677
69 2 3 0.51098390
65 2 1 0.50762488
107 2 3 0.48295375
55 2 3 0.46851074
52 2 3 0.46827948
59 2 3 0.44164146
66 2 3 0.42147865
71 2 3 0.41421605
73 2 3 0.41282512
122 2 3 0.40891392
120 2 3 0.40207904
57 2 3 0.39510378
114 2 3 0.37176468
124 2 3 0.34854822
102 2 3 0.33532624
61 2 1 0.32662688
58 2 1 0.20142024
51 2 3 0.19024422
115 2 3 0.16320750
53 2 3 0.11554863
112 2 3 -0.07433144
111 2 3 -0.07748205
103 3 2 0.59622203
106 3 2 0.59241159
108 3 2 0.58027197
110 3 2 0.56716967
123 3 2 0.56182697
121 3 2 0.55568135
119 3 2 0.53242285
118 3 2 0.52551154
125 3 2 0.51206488
105 3 2 0.49243542
101 3 2 0.45749953
113 3 2 0.44409513
109 3 2 0.37181492
117 3 2 0.26375026
116 3 2 0.21777715
104 3 2 0.21412781
Average silhouette width per cluster:
[1] 0.7931708 0.4153331 0.4678177
Average silhouette width of total data set:
[1] 0.5524757
2775 dissimilarities, summarized :
Min. 1st Qu. Median Mean 3rd Qu. Max.
0.1000 1.1136 2.5080 2.6329 3.9006 7.0852
Metric : euclidean
Number of objects : 75
Available components:
[1] "medoids" "id.med" "clustering" "objective" "isolation" "clusinfo"
"silinfo" "diss" "call"
[10] "data"
And the following graphic was generated as well:
Analysis
[edit | edit source]After the execution of the algorithm and the analysis of the data it was possible to tell that the clusters were well grouped and correlated with the species of each flower. In the data there was a total of 75 elements, 25 from Setosa species, 25 from Versicolor species and 25 from Virginica species, and the algorithm clustered the elements from Setosa as cluster 1, the ones from Versicolor as cluster 2 and the ones from Virginica as cluster 3. After verifying the results we find that out of 75 elements, 66 were correctly clustered, giving an error margin of 12%—which is a very good result.
References
[edit | edit source]- The R Development Core Team, R: A Language and Environment for Statistical Computing.
- Kaufman, L., Rousseeuw, P. J., Clustering by Means of Medoids.