Structural Biochemistry/Proteins are modified by fatty acids
Proteins are modified by fatty acid on the cysteine residue which induces a hydrophobic interactions. This is a very important post translation modification of proteins that diversifies proteins by enabling protein protein interactions, signal transduction and many other protein functions. One of the widely recognized lipid linked protein is G-protein1. The pathway that proteins follow is now being clearly understood. The discovery of multi-drug resistance (MDR) genes has changed the ways cells can target proteins. Another significant discovery is that of small lipopeptied that have been found on the surface of some proteins. Lipid modifications of proteins are important in eukaryotic cells, by increasing flexibility and increasing protein specificity. There are three covalent lipid modification of eukaryotic cells: Myristoylation, Palmitoylation and Isoprenylation
Myristoylation[edit | edit source]
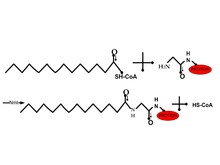
Many cellular peptides are covalently linked to a rare 14-carbon chain (acylation) on a amid bond of glycine. Since this modification was mostly observed in the cytosol, the hydrophobic function was not the only function in this modification. The N-myristoyl transferase was purified from yeast and mammals. Due to the solubility and catalysis after the removal of the interior Met can be achieved using myristoyl-coenzyme A. The acyl chain plays a more important role in the utility by an enzyme compared to the hydrophobic region 2. In vivo the substitution of the methylene group with S or O therefore making it more hydrophilic. Therefore this changes the distribution of proteins in the membrane. This can be used to target drug delivery 2.
Palmitoylation[edit | edit source]
This involves a thioester bond on the cysteine residue and primary palmitric acid. However, the mechanism that regulates this process is poorly understood due to the difficulty of purifying labile (membrane-bound palmitoyl transferases). Another challenge is that Palmitoyl-CoA is an excellent acyl donor, but no protein sequence specifications need to be fulfilied. Yet, it has been observed that sites of interest are located close to the transmembrane region on the cytoplasm side. Many of the proteins that are located near the cytoplasm region are palmitoylated 2.
Isoprenylation[edit | edit source]
This is a triptlet modifications at the C terminal, depending on the primary sequence of motif CAAX (C, cysteine; A, aliphatic aminoacid; X, any amino acid). The first step is to add to isoprenoid lipid farnesol to cysteine residue which will add a 15th cardon. The second step is to protolytycally remove the AAX and carboxyl-methylation of the a-carboxyl 2.
GPI-anchored proteins[edit | edit source]
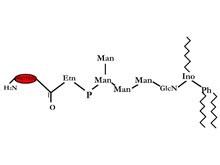
GPI-anchored proteins were discovered from Trypanosoma Variant Surface Glycoprotein (VSG) and mammalian Thy-1 antigen. It has now been seen on the surface of many eukaryotic cells including humans and yeast. A test for this bond was discovered by low using bacterial phosphatidylinositol-specific phospholipases C (PI-PLC), which releases the GPI-anchored. The amid c-terminal is linked to ethanolamine which is further linked by a phosphodiester bond to tri-mannosyl glucosaminyl core glycan. On the 6th position of the inositol of the tri-mannosyl glucosaminyl core glycan a phosphatidylinositol (PI) is linked on the outer layer of the bi-layer. There are many possible variations in the glycerol group: alkyl or acyl and may also differ in chain length. Another structural difference is that the glycan can have extra sugars some examples are aGal, /JGalNAc or aMan, and/or ethanolamine phosphate. Due to these many structural differences therefore we have many different functions that these proteins can fulfill 2.
Conclusion[edit | edit source]
Only four methods have been identified to modify eukaryotic proteins. However, this doesn't mean that there are no other lipid modifications of proteins. Using metabolic labeling and SDS-PAGE to identify proteins that are regulated by lipids it was discovered that about 10-50 proteins were observed. This may lead to a conclusion that about 10-50% are modified by lipids. One of the major functions of lipid modification of protein is localization. Analyzing lipid modification of proteins we may better understand cell function 2.
Reference[edit | edit source]
1. Jui-Yun Lu and Sandra L. Hofmann, Lysosomal Metabolism of Lipid-Modified Proteins- Journal of Lipid Research
2.ANTHONY I. MAGEE,Lipid Modification of proteins and its relevance to protein targeting. COMMENTARY